-Search query
-Search result
Showing 1 - 50 of 72 items for (author: ries & ab)

EMDB-19163: 
Trimeric HSV-1F gB ectodomain in postfusion conformation with three bound HDIT101 Fab molecules.
Method: single particle / : Kalbermatter D, Seyfizadeh N, Imhof T, Ries M, Mueller C, Jenner L, Blumenschein E, Yendrzheyevskiy A, Moog K, Eckert D, Engel R, Diebolder P, Chami M, Krauss J, Schaller T, Arndt M

EMDB-19164: 
Trimeric HSV-1F gB ectodomain in postfusion conformation with three bound HDIT102 Fab molecules.
Method: single particle / : Kalbermatter D, Seyfizadeh N, Imhof T, Ries M, Mueller C, Jenner L, Blumenschein E, Yendrzheyevskiy A, Moog K, Eckert D, Engel R, Diebolder P, Chami M, Krauss J, Schaller T, Arndt M

EMDB-19165: 
Trimeric HSV-2F gB ectodomain in postfusion conformation with three bound HDIT101 Fab molecules.
Method: single particle / : Kalbermatter D, Seyfizadeh N, Imhof T, Ries M, Mueller C, Jenner L, Blumenschein E, Yendrzheyevskiy A, Moog K, Eckert D, Engel R, Diebolder P, Chami M, Krauss J, Schaller T, Arndt M

EMDB-19166: 
Trimeric HSV-2G gB ectodomain in postfusion conformation with three bound HDIT102 Fab molecules.
Method: single particle / : Kalbermatter D, Seyfizadeh N, Imhof T, Ries M, Mueller C, Jenner L, Blumenschein E, Yendrzheyevskiy A, Moog K, Eckert D, Engel R, Diebolder P, Chami M, Krauss J, Schaller T, Arndt M

PDB-8rgz: 
Trimeric HSV-1F gB ectodomain in postfusion conformation with three bound HDIT101 Fab molecules.
Method: single particle / : Kalbermatter D, Seyfizadeh N, Imhof T, Ries M, Mueller C, Jenner L, Blumenschein E, Yendrzheyevskiy A, Moog K, Eckert D, Engel R, Diebolder P, Chami M, Krauss J, Schaller T, Arndt M

PDB-8rh0: 
Trimeric HSV-1F gB ectodomain in postfusion conformation with three bound HDIT102 Fab molecules.
Method: single particle / : Kalbermatter D, Seyfizadeh N, Imhof T, Ries M, Mueller C, Jenner L, Blumenschein E, Yendrzheyevskiy A, Moog K, Eckert D, Engel R, Diebolder P, Chami M, Krauss J, Schaller T, Arndt M

PDB-8rh1: 
Trimeric HSV-2F gB ectodomain in postfusion conformation with three bound HDIT101 Fab molecules.
Method: single particle / : Kalbermatter D, Seyfizadeh N, Imhof T, Ries M, Mueller C, Jenner L, Blumenschein E, Yendrzheyevskiy A, Moog K, Eckert D, Engel R, Diebolder P, Chami M, Krauss J, Schaller T, Arndt M

PDB-8rh2: 
Trimeric HSV-2G gB ectodomain in postfusion conformation with three bound HDIT102 Fab molecules.
Method: single particle / : Kalbermatter D, Seyfizadeh N, Imhof T, Ries M, Mueller C, Jenner L, Blumenschein E, Yendrzheyevskiy A, Moog K, Eckert D, Engel R, Diebolder P, Chami M, Krauss J, Schaller T, Arndt M

EMDB-41265: 
Cryo-EM structure of the Tripartite ATP-independent Periplasmic (TRAP) transporter SiaQM from Haemophilus influenzae (parallel dimer)
Method: single particle / : Davies JS, Currie MC, Dobson RCJ, North RA

EMDB-41266: 
Cryo-EM structure of the Tripartite ATP-independent Periplasmic (TRAP) transporter SiaQM from Haemophilus influenzae (antiparallel dimer)
Method: single particle / : Davies JS, Currie MC, Dobson RCJ, North RA

PDB-8thi: 
Cryo-EM structure of the Tripartite ATP-independent Periplasmic (TRAP) transporter SiaQM from Haemophilus influenzae (parallel dimer)
Method: single particle / : Davies JS, Currie MC, Dobson RCJ, North RA

PDB-8thj: 
Cryo-EM structure of the Tripartite ATP-independent Periplasmic (TRAP) transporter SiaQM from Haemophilus influenzae (antiparallel dimer)
Method: single particle / : Davies JS, Currie MC, Dobson RCJ, North RA
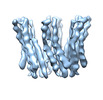
EMDB-28960: 
Cryo-electron microscopy structure of the octadecameric collagen-like (ABC-Ala)6 peptide triple helical assembly
Method: single particle / : Kreutzberger MA, Yu LT, Hancu MC, Egelman EH, Hartgerink JD

EMDB-12736: 
Ribosomal methyltransferase KsgA bound to small ribosomal subunit
Method: single particle / : Stephan NC, Ries AB

EMDB-23156: 
SARS-CoV 2 Spike Protein bound to LY-CoV555
Method: single particle / : Goldsmith JA, McLellan JS

PDB-7l3n: 
SARS-CoV 2 Spike Protein bound to LY-CoV555
Method: single particle / : Goldsmith JA, McLellan JS

EMDB-11080: 
RC-LH1(16) complex from Rhodopseudomonas palustris
Method: single particle / : Swainsbury DJK, Qian P, Hitchcock A, Hunter CN

EMDB-11081: 
RC-LH1(14)-W complex from Rhodopseudomonas palustris
Method: single particle / : Swainsbury DJK, Qian P, Hitchcock A, Hunter CN

PDB-6z5r: 
RC-LH1(16) complex from Rhodopseudomonas palustris
Method: single particle / : Swainsbury DJK, Qian P, Hitchcock A, Hunter CN

PDB-6z5s: 
RC-LH1(14)-W complex from Rhodopseudomonas palustris
Method: single particle / : Swainsbury DJK, Qian P, Hitchcock A, Hunter CN

EMDB-22943: 
Cryo-EM structure of single ACE2-bound SARS-CoV-2 trimer spike at pH 5.5
Method: single particle / : Gorman J, Rapp M, Kwong PD, Shapiro L

EMDB-22949: 
Cryo-EM Structure of Double ACE2-Bound SARS-CoV-2 Trimer Spike at pH 5.5
Method: single particle / : Gorman J, Rapp M, Kwong PD, Shapiro L

EMDB-22950: 
Cryo-EM structure of Triple ACE2-bound SARS-CoV-2 Trimer Spike at pH 5.5
Method: single particle / : Gorman J, Rapp M, Kwong PD, Shapiro L

PDB-7kne: 
Cryo-EM structure of single ACE2-bound SARS-CoV-2 trimer spike at pH 5.5
Method: single particle / : Gorman J, Rapp M, Kwong PD, Shapiro L

PDB-7knh: 
Cryo-EM Structure of Double ACE2-Bound SARS-CoV-2 Trimer Spike at pH 5.5
Method: single particle / : Gorman J, Rapp M, Kwong PD, Shapiro L

PDB-7kni: 
Cryo-EM structure of Triple ACE2-bound SARS-CoV-2 Trimer Spike at pH 5.5
Method: single particle / : Gorman J, Rapp M, Kwong PD, Shapiro L

EMDB-22922: 
ACE2-RBD Focused Refinement Using Symmetry Expansion of Applied C3 for Triple ACE2-bound SARS-CoV-2 Trimer Spike at pH 7.4
Method: single particle / : Gorman J, Kwong PD

EMDB-22927: 
Cryo-EM structure of triple ACE2-bound SARS-CoV-2 trimer spike at pH 7.4
Method: single particle / : Gorman J, Kwong PD, Shapiro L

EMDB-22932: 
Cryo-EM structure of double ACE2-bound SARS-CoV-2 trimer Spike at pH 7.4
Method: single particle / : Gorman J, Kwong PD, Shapiro L

EMDB-22941: 
Cryo-EM structure of single ACE2-bound SARS-CoV-2 trimer spike at pH 7.4
Method: single particle / : Gorman J, Kwong PD, Shapiro L

PDB-7kmb: 
ACE2-RBD Focused Refinement Using Symmetry Expansion of Applied C3 for Triple ACE2-bound SARS-CoV-2 Trimer Spike at pH 7.4
Method: single particle / : Gorman J, Kwong PD, Shapiro L

PDB-7kms: 
Cryo-EM structure of triple ACE2-bound SARS-CoV-2 trimer spike at pH 7.4
Method: single particle / : Gorman J, Kwong PD, Shapiro L

PDB-7kmz: 
Cryo-EM structure of double ACE2-bound SARS-CoV-2 trimer Spike at pH 7.4
Method: single particle / : Gorman J, Kwong PD, Shapiro L

PDB-7knb: 
Cryo-EM structure of single ACE2-bound SARS-CoV-2 trimer spike at pH 7.4
Method: single particle / : Gorman J, Kwong PD, Shapiro L

EMDB-22515: 
Structure of SARS-CoV-2 spike at pH 4.5
Method: single particle / : Tsybovsky Y, Zhou T, Kwong PD

PDB-7jwy: 
Structure of SARS-CoV-2 spike at pH 4.5
Method: single particle / : Zhou T, Tsybovsky Y, Kwong PD

EMDB-22251: 
Structure of SARS-CoV-2 spike at pH 4.0
Method: single particle / : Tsybovsky Y, Zhou T, Olia A, Kwong PD

EMDB-22253: 
Consensus structure of SARS-CoV-2 spike at pH 5.5
Method: single particle / : Tsybovsky Y, Zhou T, Olia A, Kwong PD

EMDB-22254: 
Structure of SARS-CoV-2 spike at pH 5.5, single RBD up, conformation 1
Method: single particle / : Tsybovsky Y, Zhou T, Olia A, Kwong PD

EMDB-22255: 
Structure of SARS-CoV-2 spike at pH 5.5, single RBD up, conformation 2
Method: single particle / : Tsybovsky Y, Zhou T, Olia A, Kwong PD

PDB-6xlu: 
Structure of SARS-CoV-2 spike at pH 4.0
Method: single particle / : Zhou T, Tsybovsky Y, Olia A, Kwong PD

PDB-6xm0: 
Consensus structure of SARS-CoV-2 spike at pH 5.5
Method: single particle / : Zhou T, Tsybovsky Y, Olia A, Kwong PD

PDB-6xm3: 
Structure of SARS-CoV-2 spike at pH 5.5, single RBD up, conformation 1
Method: single particle / : Zhou T, Tsybovsky Y, Olia A, Kwong PD

PDB-6xm4: 
Structure of SARS-CoV-2 spike at pH 5.5, single RBD up, conformation 2
Method: single particle / : Zhou T, Tsybovsky Y, Olia A, Kwong PD

EMDB-22256: 
Structure of SARS-CoV-2 spike at pH 5.5, all RBDs down
Method: single particle / : Tsybovsky Y, Zhou T, Olia A, Kwong PD

PDB-6xm5: 
Structure of SARS-CoV-2 spike at pH 5.5, all RBDs down
Method: single particle / : Zhou T, Tsybovsky Y, Olia A, Kwong PD

EMDB-10051: 
Cryo-EM structure of the 90S pre-ribosome (Kre33-Noc4) from Chaetomium thermophilum, state A
Method: single particle / : Cheng J, Kellner N

EMDB-10052: 
Cryo-EM structure of the 90S pre-ribosome (Kre33-Noc4) from Chaetomium thermophilum, state B1
Method: single particle / : Cheng J, Kellner N

EMDB-10053: 
Cryo-EM structure of the 90S pre-ribosome (Kre33-Noc4) from Chaetomium thermophilum, state B2
Method: single particle / : Cheng J, Kellner N, Griesel S, Berninghausen O, Beckmann R, Hurt E

EMDB-10054: 
Cryo-EM structure of the 90S pre-ribosome (Kre33-Noc4) from Chaetomium thermophilum, state C
Method: single particle / : Cheng J, Kellner N
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
