-検索条件
-検索結果
検索 (著者・登録者: qiao & z)の結果419件中、1から50件目までを表示しています

EMDB-38142: 
Structure of CCT6-HR-ATP-AlFx

EMDB-38143: 
Structure of apoferritin

EMDB-38145: 
Consensus map of TBCA-apoferritin

EMDB-38147: 
Structure of CCT6-HR

EMDB-39651: 
Structure of the focused refined TBCA-apoferritin

EMDB-43736: 
Umb1 umbrella toxin particle

EMDB-43737: 
Umb1 umbrella toxin particle (local refinement of UmbB1 bound ALF of UmbC1 and UmbA1)

PDB-8w20: 
Umb1 umbrella toxin particle

PDB-8w22: 
Umb1 umbrella toxin particle (local refinement of UmbB1 bound ALF of UmbC1 and UmbA1)
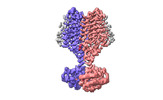
EMDB-37444: 
Cryo-EM structure of PAO1-ImcA with GMPCPP

PDB-8wcn: 
Cryo-EM structure of PAO1-ImcA with GMPCPP

EMDB-35678: 
Apo state of Arabidopsis AZG1 at pH 7.4

EMDB-35679: 
Endogenous substrate adenine bound state of Arabidopsis AZG1 at pH 5.5

EMDB-35680: 
6-BAP bound state of Arabidopsis AZG1

EMDB-35681: 
trans-Zeatin bound state of Arabidopsis AZG1 at pH7.4

EMDB-35682: 
kinetin bound state of Arabidopsis AZG1

EMDB-37658: 
trans-Zeatin bound state of Arabidopsis AZG1 at pH5.5

PDB-8irl: 
Apo state of Arabidopsis AZG1 at pH 7.4

PDB-8irm: 
Endogenous substrate adenine bound state of Arabidopsis AZG1 at pH 5.5

PDB-8irn: 
6-BAP bound state of Arabidopsis AZG1

PDB-8iro: 
trans-Zeatin bound state of Arabidopsis AZG1 at pH7.4

PDB-8irp: 
kinetin bound state of Arabidopsis AZG1

PDB-8wmq: 
trans-Zeatin bound state of Arabidopsis AZG1 at pH5.5

EMDB-37681: 
Apo state of Arabidopsis AZG1 T440Y

PDB-8wo7: 
Apo state of Arabidopsis AZG1 T440Y

EMDB-35416: 
Cryo-EM structure of the Clr6S (Clr6-HDAC) complex from S. pombe

PDB-8ifg: 
Cryo-EM structure of the Clr6S (Clr6-HDAC) complex from S. pombe

EMDB-35417: 
Clr6 HDAC (Clr6S)-nucleosome complex
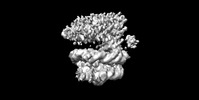
EMDB-36278: 
Rpd3S-nucleosome (H3K36me3 modified)

EMDB-35063: 
The complex structure of Omicron BA.4 RBD with BD604, S309, and S304

EMDB-35064: 
SARS-CoV-2 Omicron BA.2 RBD complexed with BD-604 and S304 Fab

PDB-8hws: 
The complex structure of Omicron BA.4 RBD with BD604, S309, and S304

PDB-8hwt: 
SARS-CoV-2 Omicron BA.2 RBD complexed with BD-604 and S304 Fab
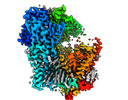
EMDB-36748: 
Structure of a fungal 1,3-beta-glucan synthase

PDB-8jzn: 
Structure of a fungal 1,3-beta-glucan synthase

EMDB-34954: 
Cryo-EM structure of SSX1 bound to the H2AK119Ub nucleosome at a resolution of 3.05 angstrom

EMDB-36747: 
Cryo-EM structure of 2 SSX1 bound to H2AK119Ub nucleosome at a resolution of 3.6 angstroms (2:1 complex)

PDB-8hqy: 
Cryo-EM structure of SSX1 bound to the H2AK119Ub nucleosome at a resolution of 3.05 angstrom
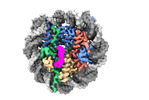
EMDB-34956: 
Cryo-EM structure of SSX1 bound to the unmodified nucleosome at a resolution of 3.02 angstrom

EMDB-34957: 
Cryo-EM structure of H2AK119Ub nucleosome at a resolution of 6.11 angstrom

PDB-8hr1: 
Cryo-EM structure of SSX1 bound to the unmodified nucleosome at a resolution of 3.02 angstrom

EMDB-34522: 
Cryo-EM Structure of SARS-CoV-2 BA.2 Spike protein in complex with BA7535

EMDB-34526: 
Cryo-EM structure of SARS-CoV-2 BA.2 RBD in complex with BA7535 fab (local refinement)

PDB-8h7l: 
Cryo-EM Structure of SARS-CoV-2 BA.2 Spike protein in complex with BA7535

PDB-8h7z: 
Cryo-EM structure of SARS-CoV-2 BA.2 RBD in complex with BA7535 fab (local refinement)

EMDB-35790: 
GMPCPP-Alpha1A/Beta2A-microtubule decorated with kinesin

EMDB-35791: 
GMPCPP-Alpha1C/Beta2A-microtubule decorated with kinesin

EMDB-35792: 
GMPCPP-Alpha4A/Beta2A-microtubule decorated with kinesin

PDB-8ixa: 
GMPCPP-Alpha1A/Beta2A-microtubule decorated with kinesin non-seam region

PDB-8ixb: 
GMPCPP-Alpha1A/Beta2A-microtubule decorated with kinesin seam region
ページ:
 ムービー
ムービー コントローラー
コントローラー 構造ビューア
構造ビューア EMN検索について
EMN検索について



 wwPDBはEMDBデータモデルのバージョン3へ移行します
wwPDBはEMDBデータモデルのバージョン3へ移行します
