[English] 日本語
 Yorodumi
Yorodumi- EMDB-35063: The complex structure of Omicron BA.4 RBD with BD604, S309, and S304 -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | The complex structure of Omicron BA.4 RBD with BD604, S309, and S304 | ||||||||||||
 Map data Map data | |||||||||||||
 Sample Sample |
| ||||||||||||
 Keywords Keywords | antibody viral protein complex / IMMUNE SYSTEM/VIRAL PROTEIN / IMMUNE SYSTEM-VIRAL PROTEIN complex | ||||||||||||
| Function / homology |  Function and homology information Function and homology informationsymbiont-mediated disruption of host tissue / Maturation of spike protein / Translation of Structural Proteins / Virion Assembly and Release / host cell surface / host extracellular region / symbiont-mediated-mediated suppression of host tetherin activity / Induction of Cell-Cell Fusion / structural constituent of virion / positive regulation of viral entry into host cell ...symbiont-mediated disruption of host tissue / Maturation of spike protein / Translation of Structural Proteins / Virion Assembly and Release / host cell surface / host extracellular region / symbiont-mediated-mediated suppression of host tetherin activity / Induction of Cell-Cell Fusion / structural constituent of virion / positive regulation of viral entry into host cell / membrane fusion / host cell endoplasmic reticulum-Golgi intermediate compartment membrane / Attachment and Entry / entry receptor-mediated virion attachment to host cell / receptor-mediated virion attachment to host cell / host cell surface receptor binding / symbiont-mediated suppression of host innate immune response / endocytosis involved in viral entry into host cell / receptor ligand activity / fusion of virus membrane with host plasma membrane / fusion of virus membrane with host endosome membrane / viral envelope / symbiont entry into host cell / virion attachment to host cell / host cell plasma membrane / SARS-CoV-2 activates/modulates innate and adaptive immune responses / virion membrane / membrane / identical protein binding / plasma membrane Similarity search - Function | ||||||||||||
| Biological species |  Homo sapiens (human) / Homo sapiens (human) /  | ||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 2.36 Å | ||||||||||||
 Authors Authors | He QW / Xu ZP / Xie YF | ||||||||||||
| Funding support |  China, 3 items China, 3 items
| ||||||||||||
 Citation Citation |  Journal: Cell Rep Med / Year: 2023 Journal: Cell Rep Med / Year: 2023Title: An updated atlas of antibody evasion by SARS-CoV-2 Omicron sub-variants including BQ.1.1 and XBB. Authors: Qingwen He / Lili Wu / Zepeng Xu / Xiaoyun Wang / Yufeng Xie / Yan Chai / Anqi Zheng / Jianjie Zhou / Shitong Qiao / Min Huang / Guijun Shang / Xin Zhao / Youjun Feng / Jianxun Qi / George Fu Gao / Qihui Wang /  Abstract: Emerging Omicron sub-variants are causing global concerns, and their immune evasion should be monitored continuously. We previously evaluated the escape of Omicron BA.1, BA.1.1, BA.2, and BA.3 from ...Emerging Omicron sub-variants are causing global concerns, and their immune evasion should be monitored continuously. We previously evaluated the escape of Omicron BA.1, BA.1.1, BA.2, and BA.3 from an atlas of 50 monoclonal antibodies (mAbs), covering seven epitope classes of the severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) receptor-binding domain (RBD). Here, we update the atlas of totally 77 mAbs against emerging sub-variants including BQ.1.1 and XBB and find that BA.4/5, BQ.1.1, and XBB display further evasion. Besides, investigation into the correlation of binding and neutralization of mAbs reveals the important role of antigenic conformation in mAb functioning. Moreover, the complex structures of BA.2 RBD/BD-604/S304 and BA.4/5 RBD/BD-604/S304/S309 further elucidate the molecular mechanism of antibody evasion by these sub-variants. By focusing on the identified broadly potent mAbs, we find a general hotspot epitope on the RBD, which could guide the design of vaccines and calls for new broad-spectrum countermeasures against COVID-19. | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_35063.map.gz emd_35063.map.gz | 158.3 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-35063-v30.xml emd-35063-v30.xml emd-35063.xml emd-35063.xml | 23.3 KB 23.3 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_35063.png emd_35063.png | 55.1 KB | ||
| Filedesc metadata |  emd-35063.cif.gz emd-35063.cif.gz | 6.8 KB | ||
| Others |  emd_35063_half_map_1.map.gz emd_35063_half_map_1.map.gz emd_35063_half_map_2.map.gz emd_35063_half_map_2.map.gz | 165 MB 165 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-35063 http://ftp.pdbj.org/pub/emdb/structures/EMD-35063 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-35063 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-35063 | HTTPS FTP |
-Related structure data
| Related structure data |  8hwsMC  8hwtC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_35063.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_35063.map.gz / Format: CCP4 / Size: 178 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.85 Å | ||||||||||||||||||||||||||||||||||||
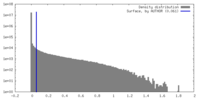
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #1
| File | emd_35063_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_35063_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Omicron BA.4/5 RBD in complex with Fabs of BD-604, S304 and S309
| Entire | Name: Omicron BA.4/5 RBD in complex with Fabs of BD-604, S304 and S309 |
|---|---|
| Components |
|
-Supramolecule #1: Omicron BA.4/5 RBD in complex with Fabs of BD-604, S304 and S309
| Supramolecule | Name: Omicron BA.4/5 RBD in complex with Fabs of BD-604, S304 and S309 type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Spike protein S2'
| Macromolecule | Name: Spike protein S2' / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 22.13598 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: TNLCPFDEVF NATRFASVYA WNRKRISNCV ADYSVLYNFA PFFAFKCYGV SPTKLNDLCF TNVYADSFVI RGNEVSQIAP GQTGNIADY NYKLPDDFTG CVIAWNSNKL DSKVGGNYNY RYRLFRKSNL KPFERDISTE IYQAGNKPCN GVAGVNCYFP L QSYGFRPT ...String: TNLCPFDEVF NATRFASVYA WNRKRISNCV ADYSVLYNFA PFFAFKCYGV SPTKLNDLCF TNVYADSFVI RGNEVSQIAP GQTGNIADY NYKLPDDFTG CVIAWNSNKL DSKVGGNYNY RYRLFRKSNL KPFERDISTE IYQAGNKPCN GVAGVNCYFP L QSYGFRPT YGVGHQPYRV VVLSFELLHA PATVCGPK UniProtKB: Spike glycoprotein |
-Macromolecule #2: BD-604 Fab Heavy chain
| Macromolecule | Name: BD-604 Fab Heavy chain / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 24.121109 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: EVQLVESGGG LIQPGGSLRL SCAASGIIVS SNYMTWVRQA PGKGLEWVSV IYSGGSTFYA DSVKGRFTIS RDNSKNTLYL QMSSLRAED TAVYYCARDL GPYGMDVWGQ GTTVTVSSAS TKGPSVFPLA PSSKSTSGGT AALGCLVKDY FPEPVTVSWN S GALTSGVH ...String: EVQLVESGGG LIQPGGSLRL SCAASGIIVS SNYMTWVRQA PGKGLEWVSV IYSGGSTFYA DSVKGRFTIS RDNSKNTLYL QMSSLRAED TAVYYCARDL GPYGMDVWGQ GTTVTVSSAS TKGPSVFPLA PSSKSTSGGT AALGCLVKDY FPEPVTVSWN S GALTSGVH TFPAVLQSSG LYSLSSVVTV PSSSLGTQTY ICNVNHKPSN TKVDKRVEPK SCDKTHTCPP C |
-Macromolecule #3: BD-604 Fab Light chain
| Macromolecule | Name: BD-604 Fab Light chain / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 23.391863 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: DIQLTQSPSF LSASVGDRVT ITCRASQGIS SDLAWYQQKP GKAPNLLIYA ASTLQSGVPS RFSGSGSGTE FTLTISSLQP EDFATYYCQ QLNSDLYTFG QGTKLEIKRT VAAPSVFIFP PSDEQLKSGT ASVVCLLNNF YPREAKVQWK VDNALQSGNS Q ESVTEQDS ...String: DIQLTQSPSF LSASVGDRVT ITCRASQGIS SDLAWYQQKP GKAPNLLIYA ASTLQSGVPS RFSGSGSGTE FTLTISSLQP EDFATYYCQ QLNSDLYTFG QGTKLEIKRT VAAPSVFIFP PSDEQLKSGT ASVVCLLNNF YPREAKVQWK VDNALQSGNS Q ESVTEQDS KDSTYSLSST LTLSKADYEK HKVYACEVTH QGLSSPVTKS FNRGECS |
-Macromolecule #4: S304 Fab Heavy Chain
| Macromolecule | Name: S304 Fab Heavy Chain / type: protein_or_peptide / ID: 4 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 24.613428 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: VQLVESGGGL VQPGGSLRLS CAASGFTFSS YDMHWVRQTT GKGLEWVSTI GTAGDTYYPD SVKGRFTISR EDAKNSLYLQ MNSLRAGDT AVYYCARGDS SGYYYYFDYW GQGTLLTVSS ASTKGPSVFP LAPSSKSTSG GTAALGCLVK DYFPEPVTVS W NSGALTSG ...String: VQLVESGGGL VQPGGSLRLS CAASGFTFSS YDMHWVRQTT GKGLEWVSTI GTAGDTYYPD SVKGRFTISR EDAKNSLYLQ MNSLRAGDT AVYYCARGDS SGYYYYFDYW GQGTLLTVSS ASTKGPSVFP LAPSSKSTSG GTAALGCLVK DYFPEPVTVS W NSGALTSG VHTFPAVLQS SGLYSLSSVV TVPSSSLGTQ TYICNVNHKP SNTKVDKRVE PKSCDKTHTC PPC |
-Macromolecule #5: S304 Fab Light Chain
| Macromolecule | Name: S304 Fab Light Chain / type: protein_or_peptide / ID: 5 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 23.45801 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: DIEMTQSPSS LSAAVGDRVT ITCRASQSIG SYLNWYQQKP GKAPKLLIYA ASSLQSGVPS RFSGSGSGTD FTLTISSLQP EDFAIYYCQ QSYVSPTYTF GPGTKVDIKR TVAAPSVFIF PPSDEQLKSG TASVVCLLNN FYPREAKVQW KVDNALQSGN S QESVTEQD ...String: DIEMTQSPSS LSAAVGDRVT ITCRASQSIG SYLNWYQQKP GKAPKLLIYA ASSLQSGVPS RFSGSGSGTD FTLTISSLQP EDFAIYYCQ QSYVSPTYTF GPGTKVDIKR TVAAPSVFIF PPSDEQLKSG TASVVCLLNN FYPREAKVQW KVDNALQSGN S QESVTEQD SKDSTYSLSS TLTLSKADYE KHKVYACEVT HQGLSSPVTK SFNRGECS |
-Macromolecule #6: S309 Fab Heavy Chain
| Macromolecule | Name: S309 Fab Heavy Chain / type: protein_or_peptide / ID: 6 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 25.586625 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: QVQLVQSGAE VKKPGASVKV SCKASGYPFT SYGISWVRQA PGQGLEWMGW ISTYNGNTNY AQKFQGRVTM TTDTSTTTGY MELRRLRSD DTAVYYCARD YTRGAWFGES LIGGFDNWGQ GTLVTVSSAS TKGPSVFPLA PSSKSTSGGT AALGCLVKDY F PEPVTVSW ...String: QVQLVQSGAE VKKPGASVKV SCKASGYPFT SYGISWVRQA PGQGLEWMGW ISTYNGNTNY AQKFQGRVTM TTDTSTTTGY MELRRLRSD DTAVYYCARD YTRGAWFGES LIGGFDNWGQ GTLVTVSSAS TKGPSVFPLA PSSKSTSGGT AALGCLVKDY F PEPVTVSW NSGALTSGVH TFPAVLQSSG LYSLSSVVTV PSSSLGTQTY ICNVNHKPSN TKVDKRVEPK SCDKTHTCPP C |
-Macromolecule #7: S309 Fab Light Chain
| Macromolecule | Name: S309 Fab Light Chain / type: protein_or_peptide / ID: 7 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 23.291777 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: EIVLTQSPGT LSLSPGERAT LSCRASQTVS STSLAWYQQK PGQAPRLLIY GASSRATGIP DRFSGSGSGT DFTLTISRLE PEDFAVYYC QQHDTSLTFG GGTKVEIKRT VAAPSVFIFP PSDEQLKSGT ASVVCLLNNF YPREAKVQWK VDNALQSGNS Q ESVTEQDS ...String: EIVLTQSPGT LSLSPGERAT LSCRASQTVS STSLAWYQQK PGQAPRLLIY GASSRATGIP DRFSGSGSGT DFTLTISRLE PEDFAVYYC QQHDTSLTFG GGTKVEIKRT VAAPSVFIFP PSDEQLKSGT ASVVCLLNNF YPREAKVQWK VDNALQSGNS Q ESVTEQDS KDSTYSLSST LTLSKADYEK HKVYACEVTH QGLSSPVTKS FNRGECS |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.2 mg/mL |
|---|---|
| Buffer | pH: 8 |
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.0 µm / Nominal defocus min: 1.0 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller








 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)