-Search query
-Search result
Showing 1 - 50 of 2,960 items for (author: peng & s)

EMDB-43514: 
SPOT-RASTR - a cryo-EM specimen preparation technique that overcomes problems with preferred orientation and the air/water interface

EMDB-45962: 
Cryo-EM structure of Tulane virus 9-6-17 variant capsid protein VP1 5-12-18

EMDB-45963: 
Cryo-EM structure of Tulane virus 9-6-17 variant capsid protein VP1 9-14-18

EMDB-45964: 
Cryo-EM structure of Tulane virus 9-6-17 variant capsid protein VP1 9-14-18, DTT-treated
EMDB-37139: 
Structure of SARS-CoV Spike protein complexed with antibody PW5-5
EMDB-37143: 
The local refined map of SARS-CoV-2 XBB Variant Spike protein complexed with antibody PW5-535
EMDB-37144: 
Trimer state of SARS-CoV Spike protein complexed with antibody PW5-535
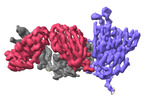

EMDB-37145: 
The local refined map of SARS-CoV Spike protein complexed with antibody PW5-5
EMDB-37160: 
Monomer state of SARS-CoV-2 XBB Variant Spike protein trimer complexed with antibody PW5-5
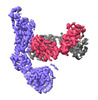
EMDB-37161: 
Monomer state of SARS-CoV Spike protein complexed with antibody PW5-535
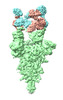
EMDB-37162: 
Structure of SARS-CoV-2 Omicron BA.1 Spike complexed with antibody PW5-570
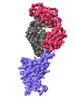
EMDB-37163: 
The local refined map of SARS-CoV-2 Omicron BA.1 Spike complexed with antibody PW5-570
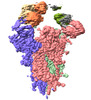
EMDB-37164: 
State 1 of SARS-CoV-2 XBB Variant Spike protein trimer complexed with antibody PW5-5
EMDB-37165: 
Structure of SARS-CoV-2 XBB Variant Spike protein complexed with broadly neutralizing antibody PW5-535

PDB-8kdm: 
Structure of SARS-CoV Spike protein complexed with antibody PW5-5

PDB-8kdr: 
The local refined map of SARS-CoV-2 XBB Variant Spike protein complexed with antibody PW5-535

PDB-8kds: 
Trimer state of SARS-CoV Spike protein complexed with antibody PW5-535

PDB-8kdt: 
The local refined map of SARS-CoV Spike protein complexed with antibody PW5-5

PDB-8kej: 
Monomer state of SARS-CoV-2 XBB Variant Spike protein trimer complexed with antibody PW5-5

PDB-8kek: 
Monomer state of SARS-CoV Spike protein complexed with antibody PW5-535

PDB-8keo: 
Structure of SARS-CoV-2 Omicron BA.1 Spike complexed with antibody PW5-570

PDB-8kep: 
The local refined map of SARS-CoV-2 Omicron BA.1 Spike complexed with antibody PW5-570

PDB-8keq: 
State 1 of SARS-CoV-2 XBB Variant Spike protein trimer complexed with antibody PW5-5

PDB-8ker: 
Structure of SARS-CoV-2 XBB Variant Spike protein complexed with broadly neutralizing antibody PW5-535

EMDB-17696: 
Structure of human 48S translation initiation complex in open codon scanning state (48S-1)

EMDB-17697: 
Structure of human 48S translation initiation complex in AUG recognition state after eIF5-induced GTP hydrolysis by eIF2 (48S-2)

EMDB-17698: 
Structure of human 48S translation initiation complex upon transfer of initiator tRNA to eIF5B (48S-3)

EMDB-17699: 
Structure of human 48S translation initiation complex after eIF5 release (48S-4)

EMDB-17700: 
Structure of human 48S translation initiation complex after eIF2 release prior 60S subunit joining (48S-5)

EMDB-17701: 
Structure of human 48S translation initiation complex with initiator tRNA, eIF1A and eIF3 (off-pathway)

EMDB-19128: 
Structure of human eIF3 core from closed 48S translation initiation complex

PDB-8pj1: 
Structure of human 48S translation initiation complex in open codon scanning state (48S-1)

PDB-8pj2: 
Structure of human 48S translation initiation complex in AUG recognition state after eIF5-induced GTP hydrolysis by eIF2 (48S-2)

PDB-8pj3: 
Structure of human 48S translation initiation complex upon transfer of initiator tRNA to eIF5B (48S-3)

PDB-8pj4: 
Structure of human 48S translation initiation complex after eIF5 release (48S-4)

PDB-8pj5: 
Structure of human 48S translation initiation complex after eIF2 release prior 60S subunit joining (48S-5)

PDB-8pj6: 
Structure of human 48S translation initiation complex with initiator tRNA, eIF1A and eIF3 (off-pathway)

PDB-8rg0: 
Structure of human eIF3 core from closed 48S translation initiation complex

EMDB-36914: 
Cryo-EM structure of Streptomyces coelicolor transcription initiation complex with the global transcription factor AfsR

EMDB-44482: 
Cryo-EM structure of the HIV-1 JR-FL IDL Env trimer in complex with PGT122 Fab
EMDB-44484: 
Cryo-EM structure of the HIV-1 BG505 IDL Env trimer in complex with 3BNC117 and 10-1074 Fabs
EMDB-44491: 
Cryo-EM structure of the HIV-1 WITO IDL Env trimer in complex with PGT122 Fab

PDB-9ber: 
Cryo-EM structure of the HIV-1 JR-FL IDL Env trimer in complex with PGT122 Fab

PDB-9bew: 
Cryo-EM structure of the HIV-1 BG505 IDL Env trimer in complex with 3BNC117 and 10-1074 Fabs

PDB-9bf6: 
Cryo-EM structure of the HIV-1 WITO IDL Env trimer in complex with PGT122 Fab

EMDB-39645: 
The structure of HKU1-B S protein with bsAb1

EMDB-39646: 
the complex structure of the H4B6 Fab with the RBD of Omicron BA.5 S protein

PDB-8yww: 
The structure of HKU1-B S protein with bsAb1

PDB-8ywx: 
the complex structure of the H4B6 Fab with the RBD of Omicron BA.5 S protein

EMDB-38873: 
cryo-EM structure of Staphylococcus aureus(ATCC 29213) 50S ribosome in complex with MCX-190.
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
