-Search query
-Search result
Showing 1 - 50 of 72 items for (author: pascal & ke)

EMDB-41617: 
CryoEM structure of PI3Kalpha

PDB-8tu6: 
CryoEM structure of PI3Kalpha

EMDB-29930: 
T. cruzi topoisomerase II alpha bound to dsDNA and the covalent inhibitor CT1

PDB-8gcc: 
T. cruzi topoisomerase II alpha bound to dsDNA and the covalent inhibitor CT1

EMDB-14421: 
Structure of yeast RNA Polymerase III-Ty1 integrase complex at 2.6 A (focus subunit AC40).

EMDB-14468: 
Structure of yeast RNA Polymerase III-DNA-Ty1 integrase complex (Pol III-DNA-IN1) at 3.1 A

EMDB-14469: 
Structure of yeast RNA Polymerase III-Ty1 integrase complex at 2.9 A (focus subunit C11 terminal Zn-ribbon in the funnel pore).

EMDB-14470: 
Structure of yeast RNA Polymerase III-Ty1 integrase complex at 2.7 A (focus subunit C11, no C11 C-terminal Zn-ribbon in the funnel pore).

EMDB-16299: 
Structure of yeast RNA Polymerase III elongation complex at 3.3 A

PDB-7z0h: 
Structure of yeast RNA Polymerase III-Ty1 integrase complex at 2.6 A (focus subunit AC40).

PDB-7z2z: 
Structure of yeast RNA Polymerase III-DNA-Ty1 integrase complex (Pol III-DNA-IN1) at 3.1 A

PDB-7z30: 
Structure of yeast RNA Polymerase III-Ty1 integrase complex at 2.9 A (focus subunit C11 terminal Zn-ribbon in the funnel pore).

PDB-7z31: 
Structure of yeast RNA Polymerase III-Ty1 integrase complex at 2.7 A (focus subunit C11, no C11 C-terminal Zn-ribbon in the funnel pore).

PDB-8bws: 
Structure of yeast RNA Polymerase III elongation complex at 3.3 A

EMDB-26005: 
Structure of the Inmazeb cocktail and resistance to escape against Ebola virus

PDB-7tn9: 
Structure of the Inmazeb cocktail and resistance to escape against Ebola virus

EMDB-25136: 
Structure of a partially disrupted IgE high affinity receptor complex bound to an omalizumab variant

PDB-7sht: 
Structure of a partially disrupted IgE high affinity receptor complex bound to an omalizumab variant

EMDB-23662: 
Complex of SARS-CoV-2 receptor binding domain with the Fab fragments of neutralizing antibodies REGN10985 and REGN10989

PDB-7m42: 
Complex of SARS-CoV-2 receptor binding domain with the Fab fragments of neutralizing antibodies REGN10985 and REGN10989

EMDB-11639: 
Cryo-EM structure of a prefusion stabilized SARS-CoV-2 Spike (D614N, R682S, R685G, A892P, A942P and V987P)(S-closed trimer)

EMDB-11719: 
Cryo-EM structure of a prefusion stabilized SARS-CoV-2 Spike (D614N, R682S, R685G, A892P, A942P and V987P)(One up trimer)

PDB-7a4n: 
Cryo-EM structure of a prefusion stabilized SARS-CoV-2 Spike (D614N, R682S, R685G, A892P, A942P and V987P)(S-closed trimer)

PDB-7ad1: 
Cryo-EM structure of a prefusion stabilized SARS-CoV-2 Spike (D614N, R682S, R685G, A892P, A942P and V987P)(One up trimer)

EMDB-11066: 
CryoEM structure of horse sodium/proton exchanger NHE9 in an inward-facing conformation

EMDB-11067: 
CryoEM structure of horse sodium/proton exchanger NHE9 without C-terminal regulatory domain in an inward-facing conformation

PDB-6z3y: 
CryoEM structure of horse sodium/proton exchanger NHE9 in an inward-facing conformation

PDB-6z3z: 
CryoEM structure of horse sodium/proton exchanger NHE9 without C-terminal regulatory domain in an inward-facing conformation

EMDB-22301: 
SARS-CoV-2 Spike D614G variant, minus RBD

EMDB-22137: 
Complex of SARS-CoV-2 receptor binding domain with the Fab fragments of two neutralizing antibodies

PDB-6xdg: 
Complex of SARS-CoV-2 receptor binding domain with the Fab fragments of two neutralizing antibodies

EMDB-10090: 
Hen egg-white lysozyme by serial electron diffraction

EMDB-10091: 
Granulovirus occlusion bodies by serial electron diffraction

PDB-6s2n: 
Hen egg-white lysozyme by serial electron diffraction

PDB-6s2o: 
Granulovirus occlusion bodies by serial electron diffraction

EMDB-10319: 
Cryo-EM structure of the full-length ABC-transporter IrtAB

EMDB-4645: 
CryoEM structure of the human ClC-1 chloride channel, membrane domain

EMDB-4646: 
CryoEM structure of the human ClC-1 chloride channel, CBS state 1

EMDB-4647: 
CryoEM structure of the human ClC-1 chloride channel, CBS state 1

EMDB-4649: 
CryoEM structure of the human ClC-1 chloride channel, CBS state 2

EMDB-4657: 
CryoEM structure of the human ClC-1 chloride channel, low pH

PDB-6qv6: 
CryoEM structure of the human ClC-1 chloride channel, membrane domain

PDB-6qvb: 
CryoEM structure of the human ClC-1 chloride channel, CBS state 3

PDB-6qvc: 
CryoEM structure of the human ClC-1 chloride channel, CBS state 1

PDB-6qvd: 
CryoEM structure of the human ClC-1 chloride channel, CBS state 2

PDB-6qvu: 
CryoEM structure of the human ClC-1 chloride channel, low pH

EMDB-7900: 
REGN3479 antibody Fab in complex with Ebola virus GP

EMDB-7901: 
REGN3470 antibody Fab in complex with Ebola virus GP

EMDB-7902: 
REGN3471 antibody Fab in complex with Ebola virus GP
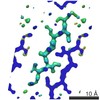
EMDB-8720: 
VGSNKGAIIGL from Amyloid Beta determined by MicroED
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
