+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-22301 | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | SARS-CoV-2 Spike D614G variant, minus RBD | ||||||||||||
 Map data Map data | |||||||||||||
 Sample Sample |
| ||||||||||||
 Keywords Keywords | SARS-CoV-2 / Spike / D614G / VIRAL PROTEIN | ||||||||||||
| Function / homology |  Function and homology information Function and homology informationsymbiont-mediated disruption of host tissue / Maturation of spike protein / Translation of Structural Proteins / Virion Assembly and Release / host cell surface / host extracellular region / symbiont-mediated-mediated suppression of host tetherin activity / Induction of Cell-Cell Fusion / structural constituent of virion / positive regulation of viral entry into host cell ...symbiont-mediated disruption of host tissue / Maturation of spike protein / Translation of Structural Proteins / Virion Assembly and Release / host cell surface / host extracellular region / symbiont-mediated-mediated suppression of host tetherin activity / Induction of Cell-Cell Fusion / structural constituent of virion / positive regulation of viral entry into host cell / membrane fusion / host cell endoplasmic reticulum-Golgi intermediate compartment membrane / Attachment and Entry / entry receptor-mediated virion attachment to host cell / receptor-mediated virion attachment to host cell / host cell surface receptor binding / symbiont-mediated suppression of host innate immune response / endocytosis involved in viral entry into host cell / receptor ligand activity / fusion of virus membrane with host plasma membrane / fusion of virus membrane with host endosome membrane / viral envelope / symbiont entry into host cell / virion attachment to host cell / host cell plasma membrane / SARS-CoV-2 activates/modulates innate and adaptive immune responses / virion membrane / membrane / identical protein binding / plasma membrane Similarity search - Function | ||||||||||||
| Biological species |  | ||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.7 Å | ||||||||||||
 Authors Authors | Wang X / Egri SB | ||||||||||||
| Funding support |  United States, 3 items United States, 3 items
| ||||||||||||
 Citation Citation |  Journal: Cell / Year: 2020 Journal: Cell / Year: 2020Title: Structural and Functional Analysis of the D614G SARS-CoV-2 Spike Protein Variant. Authors: Leonid Yurkovetskiy / Xue Wang / Kristen E Pascal / Christopher Tomkins-Tinch / Thomas P Nyalile / Yetao Wang / Alina Baum / William E Diehl / Ann Dauphin / Claudia Carbone / Kristen ...Authors: Leonid Yurkovetskiy / Xue Wang / Kristen E Pascal / Christopher Tomkins-Tinch / Thomas P Nyalile / Yetao Wang / Alina Baum / William E Diehl / Ann Dauphin / Claudia Carbone / Kristen Veinotte / Shawn B Egri / Stephen F Schaffner / Jacob E Lemieux / James B Munro / Ashique Rafique / Abhi Barve / Pardis C Sabeti / Christos A Kyratsous / Natalya V Dudkina / Kuang Shen / Jeremy Luban /   Abstract: The SARS-CoV-2 spike (S) protein variant D614G supplanted the ancestral virus worldwide, reaching near fixation in a matter of months. Here we show that D614G was more infectious than the ancestral ...The SARS-CoV-2 spike (S) protein variant D614G supplanted the ancestral virus worldwide, reaching near fixation in a matter of months. Here we show that D614G was more infectious than the ancestral form on human lung cells, colon cells, and on cells rendered permissive by ectopic expression of human ACE2 or of ACE2 orthologs from various mammals, including Chinese rufous horseshoe bat and Malayan pangolin. D614G did not alter S protein synthesis, processing, or incorporation into SARS-CoV-2 particles, but D614G affinity for ACE2 was reduced due to a faster dissociation rate. Assessment of the S protein trimer by cryo-electron microscopy showed that D614G disrupts an interprotomer contact and that the conformation is shifted toward an ACE2 binding-competent state, which is modeled to be on pathway for virion membrane fusion with target cells. Consistent with this more open conformation, neutralization potency of antibodies targeting the S protein receptor-binding domain was not attenuated. #1:  Journal: Biorxiv / Year: 2020 Journal: Biorxiv / Year: 2020Title: Structural and Functional Analysis of the D614G SARS-CoV-2 Spike Protein Variant Authors: Yurkovetskiy L / Wang X / Pascal KE / Tompkins-Tinch C / Nyalile T / Wang Y / Baum A / Diehl WE / Dauphin A / Carbone C / Veinotte K / Egri SB / Schaffner SF / Lemieux JE / Munro J / Rafique ...Authors: Yurkovetskiy L / Wang X / Pascal KE / Tompkins-Tinch C / Nyalile T / Wang Y / Baum A / Diehl WE / Dauphin A / Carbone C / Veinotte K / Egri SB / Schaffner SF / Lemieux JE / Munro J / Rafique A / Barve A / Sabeti PC / Kyratsous C / Dudkina N / Shen K / Luban J | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_22301.map.gz emd_22301.map.gz | 479.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-22301-v30.xml emd-22301-v30.xml emd-22301.xml emd-22301.xml | 17.1 KB 17.1 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_22301.png emd_22301.png | 172.9 KB | ||
| Filedesc metadata |  emd-22301.cif.gz emd-22301.cif.gz | 6.6 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-22301 http://ftp.pdbj.org/pub/emdb/structures/EMD-22301 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-22301 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-22301 | HTTPS FTP |
-Related structure data
| Related structure data |  6xs6MC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_22301.map.gz / Format: CCP4 / Size: 512 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_22301.map.gz / Format: CCP4 / Size: 512 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.52 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
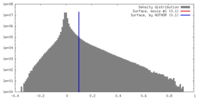
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : SARS2 Spike D614G
| Entire | Name: SARS2 Spike D614G |
|---|---|
| Components |
|
-Supramolecule #1: SARS2 Spike D614G
| Supramolecule | Name: SARS2 Spike D614G / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 417 kDa/nm |
-Macromolecule #1: Spike glycoprotein
| Macromolecule | Name: Spike glycoprotein / type: protein_or_peptide / ID: 1 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 139.258344 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MFVFLVLLPL VSSQCVNLTT RTQLPPAYTN SFTRGVYYPD KVFRSSVLHS TQDLFLPFFS NVTWFHAIHV SGTNGTKRFD NPVLPFNDG VYFASTEKSN IIRGWIFGTT LDSKTQSLLI VNNATNVVIK VCEFQFCNDP FLGVYYHKNN KSWMESEFRV Y SSANNCTF ...String: MFVFLVLLPL VSSQCVNLTT RTQLPPAYTN SFTRGVYYPD KVFRSSVLHS TQDLFLPFFS NVTWFHAIHV SGTNGTKRFD NPVLPFNDG VYFASTEKSN IIRGWIFGTT LDSKTQSLLI VNNATNVVIK VCEFQFCNDP FLGVYYHKNN KSWMESEFRV Y SSANNCTF EYVSQPFLMD LEGKQGNFKN LREFVFKNID GYFKIYSKHT PINLVRDLPQ GFSALEPLVD LPIGINITRF QT LLALHRS YLTPGDSSSG WTAGAAAYYV GYLQPRTFLL KYNENGTITD AVDCALDPLS ETKCTLKSFT VEKGIYQTSN FRV QPTESI VRFPNITNLC PFGEVFNATR FASVYAWNRK RISNCVADYS VLYNSASFST FKCYGVSPTK LNDLCFTNVY ADSF VIRGD EVRQIAPGQT GKIADYNYKL PDDFTGCVIA WNSNNLDSKV GGNYNYLYRL FRKSNLKPFE RDISTEIYQA GSTPC NGVE GFNCYFPLQS YGFQPTNGVG YQPYRVVVLS FELLHAPATV CGPKKSTNLV KNKCVNFNFN GLTGTGVLTE SNKKFL PFQ QFGRDIADTT DAVRDPQTLE ILDITPCSFG GVSVITPGTN TSNQVAVLYQ GVNCTEVPVA IHADQLTPTW RVYSTGS NV FQTRAGCLIG AEHVNNSYEC DIPIGAGICA SYQTQTNSPA SVASQSIIAY TMSLGAENSV AYSNNSIAIP TNFTISVT T EILPVSMTKT SVDCTMYICG DSTECSNLLL QYGSFCTQLN RALTGIAVEQ DKNTQEVFAQ VKQIYKTPPI KDFGGFNFS QILPDPSKPS KRSFIEDLLF NKVTLADAGF IKQYGDCLGD IAARDLICAQ KFNGLTVLPP LLTDEMIAQY TSALLAGTIT SGWTFGAGA ALQIPFAMQM AYRFNGIGVT QNVLYENQKL IANQFNSAIG KIQDSLSSTA SALGKLQDVV NQNAQALNTL V KQLSSNFG AISSVLNDIL SRLDPPEAEV QIDRLITGRL QSLQTYVTQQ LIRAAEIRAS ANLAATKMSE CVLGQSKRVD FC GKGYHLM SFPQSAPHGV VFLHVTYVPA QEKNFTTAPA ICHDGKAHFP REGVFVSNGT HWFVTQRNFY EPQIITTDNT FVS GNCDVV IGIVNNTVYD PLQPELDSFK EELDKYFKNH TSPDVDLGDI SGINASVVNI QKEIDRLNEV AKNLNESLID LQEL GKYEQ YIKWPSGRLV PRGSPGSGYI PEAPRDGQAY VRKDGEWVLL STFLGHHHHH H UniProtKB: Spike glycoprotein |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: OTHER / Average electron dose: 0.6 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: OTHER / Imaging mode: OTHER |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Protocol: AB INITIO MODEL |
|---|---|
| Output model |  PDB-6xs6: |
 Movie
Movie Controller
Controller












 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)