-検索条件
-検索結果
検索 (著者・登録者: oh & s)の結果9,909件中、1から50件目までを表示しています
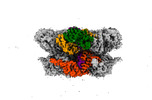
EMDB-42603: 
Human p97/VCP structure with a triazole inhibitor (NSC799462/hexamer)
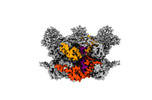
EMDB-42625: 
Human p97/VCP R155H mutant structure with a triazole inhibitor (NSC804515)
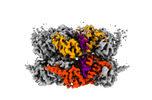
EMDB-42626: 
Human p97/VCP R155H mutant structure with a triazole inhibitor (NSC819701/up)
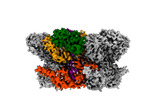
EMDB-42627: 
Human p97/VCP R155H mutant structure with a triazole inhibitor (NSC819701/down)
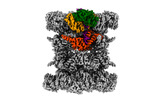
EMDB-44748: 
Human p97/VCP structure with a triazole inhibitor (NSC799462/dodecamer)

EMDB-41766: 
CryoEM structure of D2 dopamine receptor in complex with GoA KE mutant, scFv16, and dopamine

EMDB-41776: 
CryoEM structure of D2 dopamine receptor in complex with GoA KE mutant and dopamine

EMDB-18162: 
N5-methyl-H4MPT:CoM methyltransferase -coenzyme M complex + CoM

EMDB-19209: 
TadA/CpaF with ADP

EMDB-19275: 
TadA/CpaF with AMPPNP

EMDB-19279: 
TadA/CpaF nucleotide free

PDB-8rjf: 
TadA/CpaF with ADP

PDB-8rkd: 
TadA/CpaF with AMPPNP

PDB-8rkl: 
TadA/CpaF nucleotide free

EMDB-45623: 
MicroED structure of the C11 cysteine protease clostripain

PDB-9cip: 
MicroED structure of the C11 cysteine protease clostripain

EMDB-38845: 
Icosahedrally averaged cryo-EM reconstruction of PhiKZ capsid before applying the "block-based" reconstruction method

EMDB-38846: 
Block 1 of PhiKZ capsid

EMDB-38848: 
Block 2 of PhiKZ capsid

EMDB-39002: 
Composite cryo-EM map of PhiKZ capsid after applying the "block-based" reconstruction method

PDB-8y6v: 
Near-atomic structure of icosahedrally averaged jumbo bacteriophage PhiKZ capsid

EMDB-37571: 
cryo-EM structure of alligator haemoglobin in carbonmonoxy form

EMDB-37572: 
cryo-EM structure of alligator haemoglobin in oxy form

EMDB-37573: 
cryo-EM structure of alligator haemoglobin in deoxy form

EMDB-37574: 
cryo-EM structure of human haemoglobin in carbonmonoxy form

EMDB-37575: 
cryo-EM structure of human haemoglobin in oxy form

EMDB-37576: 
cryo-EM structure of human haemoglobin in deoxy form

PDB-8wix: 
cryo-EM structure of alligator haemoglobin in carbonmonoxy form

PDB-8wiy: 
cryo-EM structure of alligator haemoglobin in oxy form

PDB-8wiz: 
cryo-EM structure of alligator haemoglobin in deoxy form

PDB-8wj0: 
cryo-EM structure of human haemoglobin in carbonmonoxy form

PDB-8wj1: 
cryo-EM structure of human haemoglobin in oxy form

PDB-8wj2: 
cryo-EM structure of human haemoglobin in deoxy form
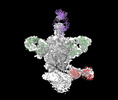
EMDB-44246: 
Cryo-EM structure of HIV-1 JRFL v6 Env in complex with vaccine-elicited, Membrane Proximal External Region (MPER) directed antibody DH1317.4.

EMDB-43257: 
AP-6 bound human TMEM175
EMDB-43259: 
2-PPA bound human TMEM175

PDB-8vic: 
AP-6 bound human TMEM175

PDB-8vie: 
2-PPA bound human TMEM175

EMDB-18128: 
Mycobacterium smegmatis RNA polymerase in complex with HelD, SigA and RbpA in State I

PDB-8q3i: 
Mycobacterium smegmatis RNA polymerase in complex with HelD, SigA and RbpA in State I

EMDB-41452: 
Cryo-EM structure of monomeric alpha-Klotho

PDB-8toh: 
Cryo-EM structure of monomeric alpha-Klotho

EMDB-50222: 
Human DNA polymerase epsilon bound to DNA and PCNA (open conformation)

EMDB-50223: 
Human DNA polymerase epsilon bound to DNA and PCNA (ajar conformation)

EMDB-50224: 
Human DNA polymerase epsilon bound to DNA and PCNA (closed conformation)

EMDB-50225: 
Human DNA Polymerase epsilon bound to T-C mismatched DNA (Post-Insertion state)

EMDB-50226: 
Human DNA Polymerase epsilon bound to T-C mismatched DNA (Polymerase Arrest state)

EMDB-50227: 
Human DNA Polymerase epsilon bound to T-C mismatched DNA (Frayed Substrate state)

EMDB-50228: 
Human DNA Polymerase epsilon bound to T-C mismatched DNA (Mismatch Excision state)
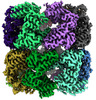
EMDB-45078: 
E.coli GroEL + PBZ1587 inhibitor
ページ:
 ムービー
ムービー コントローラー
コントローラー 構造ビューア
構造ビューア EMN検索について
EMN検索について



 wwPDBはEMDBデータモデルのバージョン3へ移行します
wwPDBはEMDBデータモデルのバージョン3へ移行します
