-Search query
-Search result
Showing 1 - 50 of 68 items for (author: nickel & l)

EMDB-53511: 
SpCas9 with computationally designed SpCas9_b10 binder
Method: single particle / : Pacesa M, Nickel L, Correia BE

EMDB-53510: 
SpCas9 with computationally designed SpCas9_b3 binder
Method: single particle / : Pacesa M, Nickel L, Correia BE

EMDB-19716: 
Cryo-EM structure of apo human SLC19A3 in outward-open state
Method: single particle / : Gabriel F, Loew C

EMDB-19750: 
Cryo-EM structure of thiamine-bound human SLC19A3 in outward-open state
Method: single particle / : Gabriel F, Loew C

EMDB-19752: 
Cryo-EM structure of fedratinib-bound human SLC19A3 in inward-open state
Method: single particle / : Gabriel F, Loew C

EMDB-19753: 
Cryo-EM structure of hydroxychloroquine-bound human SLC19A3 in inward-open state
Method: single particle / : Gabriel F, Loew C

EMDB-19754: 
Cryo-EM structure of thiamine-bound human SLC19A3 in inward-open state
Method: single particle / : Gabriel F, Loew C

EMDB-19755: 
Cryo-EM structure of amprolium-bound human SLC19A3 in inward-open state
Method: single particle / : Gabriel F, Loew C

EMDB-51088: 
Cryo-EM structure of apo human SLC19A3 in inward-open state
Method: single particle / : Gabriel F, Loew C

PDB-8s4u: 
Cryo-EM structure of apo human SLC19A3 in outward-open state
Method: single particle / : Gabriel F, Loew C

PDB-8s5u: 
Cryo-EM structure of thiamine-bound human SLC19A3 in outward-open state
Method: single particle / : Gabriel F, Loew C

PDB-8s5w: 
Cryo-EM structure of fedratinib-bound human SLC19A3 in inward-open state
Method: single particle / : Gabriel F, Loew C

PDB-8s5z: 
Cryo-EM structure of hydroxychloroquine-bound human SLC19A3 in inward-open state
Method: single particle / : Gabriel F, Loew C

PDB-8s61: 
Cryo-EM structure of thiamine-bound human SLC19A3 in inward-open state
Method: single particle / : Gabriel F, Loew C

PDB-8s62: 
Cryo-EM structure of amprolium-bound human SLC19A3 in inward-open state
Method: single particle / : Gabriel F, Loew C

PDB-9g5k: 
Cryo-EM structure of apo human SLC19A3 in inward-open state
Method: single particle / : Gabriel F, Loew C

EMDB-19005: 
structure of the GLMP/MFSD1 complex
Method: single particle / : Jungnickel KEJ, Loew C

EMDB-19006: 
Lysosomal peptide transporter
Method: single particle / : Jungnickel KEJ, Loew C

PDB-8r8q: 
Lysosomal peptide transporter
Method: single particle / : Jungnickel KEJ, Loew C

EMDB-15244: 
Tomogram of an Ebola VLP composed of GP, VP40, NP, VP24 and VP35 at pH 7.4 (Figure 1A-D)
Method: electron tomography / : Winter SL, Chlanda P

EMDB-15268: 
Tomogram of an Ebola VLP composed of VP40 at pH 4.5 (Figure 1J)
Method: electron tomography / : Winter SL, Chlanda P
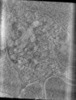
EMDB-15951: 
Tomogram of an EBOV-infected Huh7 cell showing a late endosome with internalized EBOV particles
Method: electron tomography / : Winter SL, Chlanda P

EMDB-15956: 
Tomogram of an extracellular EBOV particle adjacent to an EBOV-infected Huh7 cell
Method: electron tomography / : Winter SL, Chlanda P

EMDB-16128: 
Tomogram of a late endosome of A549 cell infected with influenza A virus.
Method: electron tomography / : Klein S, Chlanda P

EMDB-16129: 
Tomogram of a late endosome of A549 cell infected with influenza A virus (Figure 6C).
Method: electron tomography / : Klein S, Chlanda P

EMDB-16130: 
Tomogram of a late endosome of A549 cell infected with influenza A virus (Figure 6E)
Method: electron tomography / : Klein S, Chlanda P

EMDB-16131: 
Tomogram of a late endosome of A549 cell infected with influenza A virus (Figure 6G)
Method: electron tomography / : Klein S, Chlanda P

EMDB-16132: 
Tomogram of a late endosome of A549 cell infected with influenza A virus (Figure 6I,K,M)
Method: electron tomography / : Klein S, Chlanda P

EMDB-16133: 
Tomogram of a late endosome of A549 cell infected with influenza A virus (Figure 6O)
Method: electron tomography / : Klein S, Chlanda P

EMDB-15705: 
Tomogram of a late endosome of A549 cell (Figure 1R)
Method: electron tomography / : Klein S, Chlanda P

EMDB-15707: 
Tomogram of a late endosome of A549 cell treated with IFN-beta (Figure 1T)
Method: electron tomography / : Klein S, Chlanda P
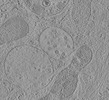
EMDB-15708: 
Tomogram of a late endosome of A549-IFITM3 cell (Figure 1V)
Method: electron tomography / : Klein S, Chlanda P

EMDB-26438: 
Transcription antitermination complex: "pre-engaged" Qlambda-loading complex
Method: single particle / : Yin Z, Ebright RH

EMDB-26439: 
Transcription antitermination complex: NusA-containing "engaged" Qlambda-loading complex
Method: single particle / : Yin Z, Ebright RH

PDB-7ubm: 
Transcription antitermination complex: "pre-engaged" Qlambda-loading complex
Method: single particle / : Yin Z, Ebright RH

PDB-7ubn: 
Transcription antitermination complex: NusA-containing "engaged" Qlambda-loading complex
Method: single particle / : Yin Z, Ebright RH

EMDB-15131: 
Tomogram of a late endosome of A549-IFITM3 cells infected with influenza A virus (Figure S7)
Method: electron tomography / : Klein S, Chlanda P

EMDB-15130: 
Tomogram of a late endosome of A549-IFITM3 cells infected with influenza A virus (Figure 4E)
Method: electron tomography / : Klein S, Chlanda P
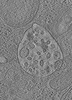
EMDB-15132: 
Tomogram of a late endosome of A549-IFITM3 cells infected with influenza A virus (Figure S8)
Method: electron tomography / : Klein S, Chlanda P

EMDB-15133: 
Tomogram of a late endosome of A549-IFITM3 cells infected with influenza A virus (Figure S9)
Method: electron tomography / : Klein S, Chlanda P

EMDB-24148: 
Escherichia coli sigma 70-dependent paused transcription elongation complex
Method: single particle / : Molodtsov V, Su M

PDB-7n4e: 
Escherichia coli sigma 70-dependent paused transcription elongation complex
Method: single particle / : Molodtsov V, Su M, Ebright RH

EMDB-24424: 
Cryo-EM structure of Thermus thermophilus reiterative transcription complex with 11nt oligo-G RNA
Method: single particle / : Liu Y, Ebright RH

PDB-7rdq: 
Cryo-EM structure of Thermus thermophilus reiterative transcription complex with 11nt oligo-G RNA
Method: single particle / : Liu Y, Ebright RH

EMDB-12324: 
In situ cryo-electron tomogram of the Synechocystis wild-type VIPP1 strain grown in high light
Method: electron tomography / : Wietrzynski W, Klumpe S, Heinz S, Spaniol B, Schaffer M, Rast A, Nickelsen J, Schroda M, Engel BD

EMDB-12325: 
In situ cryo-electron tomogram of the Synechocystis F4E VIPP1 mutant grown in high light
Method: electron tomography / : Wietrzynski W, Klumpe S, Heinz S, Spaniol B, Schaffer M, Rast A, Nickelsen J, Schroda M, Engel BD

EMDB-12326: 
In situ cryo-electron tomogram of the Synechocystis V11E VIPP1 mutant grown in high light
Method: electron tomography / : Wietrzynski W, Klumpe S, Heinz S, Spaniol B, Schaffer M, Rast A, Nickelsen J, Schroda M, Engel BD

EMDB-12327: 
In situ cryo-electron tomogram of a cluster of VIPP1-mCherry structures inside the Chlamydomonas chloroplast
Method: electron tomography / : Wietrzynski W, Klumpe S, Heinz S, Spaniol B, Schaffer M, Rast A, Nickelsen J, Schroda M, Engel BD

EMDB-12328: 
In situ cryo-electron tomogram of a cluster of VIPP1-mCherry structures inside the Chlamydomonas chloroplast
Method: electron tomography / : Wietrzynski W, Klumpe S, Heinz S, Spaniol B, Schaffer M, Rast A, Nickelsen J, Schroda M, Engel BD

EMDB-12329: 
In situ cryo-electron tomogram of a cluster of VIPP1-mCherry structures inside the Chlamydomonas chloroplast
Method: electron tomography / : Wietrzynski W, Klumpe S, Heinz S, Spaniol B, Schaffer M, Rast A, Nickelsen J, Schroda M, Engel BD
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
