-Search query
-Search result
Showing 1 - 50 of 386 items for (author: meier & b)

EMDB-17461: 
Cytochrome bc1 complex (Bos taurus)

PDB-8p65: 
Cytochrome bc1 complex (Bos taurus)

EMDB-17863: 
2.7 A cryo-EM structure of in vitro assembled type 1 pilus rod

EMDB-17878: 
2.5 A cryo-EM structure of the in vitro FimD-catalyzed assembly of type 1 pilus rod

PDB-8psv: 
2.7 A cryo-EM structure of in vitro assembled type 1 pilus rod

PDB-8ptu: 
2.5 A cryo-EM structure of the in vitro FimD-catalyzed assembly of type 1 pilus rod

EMDB-18207: 
Ubiquitin ligation to substrate by a cullin-RING E3 ligase & Cdc34: NEDD8-CUL2-RBX1-ELOB/C-FEM1C with trapped UBE2R2~donor UB-Sil1 peptide, Glacios map

EMDB-18230: 
Ubiquitin ligation to substrate by a cullin-RING E3 ligase & Cdc34: NEDD8-CUL2-RBX1-ELOB/C-FEM1C with trapped UBE2R2~donor UB-Sil1 peptide
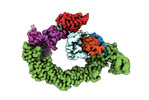
EMDB-18915: 
Ubiquitin ligation to neosubstrate by a cullin-RING E3 ligase & Cdc34: NEDD8-CUL2-RBX1-ELOB/C-VHL-MZ1 with trapped UBE2R2~donor UB-BRD4 BD2

PDB-8q7r: 
Ubiquitin ligation to substrate by a cullin-RING E3 ligase & Cdc34: NEDD8-CUL2-RBX1-ELOB/C-FEM1C with trapped UBE2R2~donor UB-Sil1 peptide

PDB-8r5h: 
Ubiquitin ligation to neosubstrate by a cullin-RING E3 ligase & Cdc34: NEDD8-CUL2-RBX1-ELOB/C-VHL-MZ1 with trapped UBE2R2~donor UB-BRD4 BD2

EMDB-16319: 
Structure of the K+/H+ exchanger KefC with GSH

PDB-8by2: 
Structure of the K+/H+ exchanger KefC with GSH.

EMDB-16318: 
Structure of the K/H exchanger KefC

PDB-8bxg: 
Structure of the K/H exchanger KefC.

EMDB-16052: 
Cryo-EM structure of stalled rabbit 80S ribosomes in complex with human CCR4-NOT and CNOT4

PDB-8bhf: 
Cryo-EM structure of stalled rabbit 80S ribosomes in complex with human CCR4-NOT and CNOT4

EMDB-27920: 
3H03 Fab in complex with influenza virus neuraminidase from A/Brevig Mission/1/1918 (H1N1)

EMDB-27921: 
2H08 Fab in complex with influenza virus neuraminidase from A/Brevig Mission/1/1918 (H1N1)

PDB-8e6j: 
3H03 Fab in complex with influenza virus neuraminidase from A/Brevig Mission/1/1918 (H1N1)

PDB-8e6k: 
2H08 Fab in complex with influenza virus neuraminidase from A/Brevig Mission/1/1918 (H1N1)

EMDB-16155: 
Structure of rabbit 80S ribosome translating beta-tubulin in complex with tetratricopeptide protein 5 (TTC5) and S-phase Cyclin A Associated Protein residing in the ER (SCAPER)

PDB-8bpo: 
Structure of rabbit 80S ribosome translating beta-tubulin in complex with tetratricopeptide protein 5 (TTC5) and S-phase Cyclin A Associated Protein residing in the ER (SCAPER)

EMDB-15118: 
13pf undecorated microtubule from recombinant human tubulin (alpha1B, beta3) lacking the C-terminal tail

EMDB-15119: 
13pf undecorated microtubule from recombinant human tubulin (alpha1B, beta3) with spliced unmodified C-terminal tail on alpha1B.

EMDB-15120: 
13pf undecorated microtubule from recombinant human tubulin (alpha1B, beta3) with spliced C-terminal tail containing 10E branch on alpha1B.

EMDB-16036: 
human MDM2-5S RNP

EMDB-16037: 
hexameric ct-sc 5S RNP monomer

EMDB-16038: 
hexameric ct-sc 5S RNP dimer

EMDB-16040: 
pre-60S with human 5S RNP

PDB-8bgu: 
human MDM2-5S RNP

EMDB-13477: 
Small subunit of the Chlamydomonas reinhardtii mitoribosome

EMDB-15295: 
African cichlid nackednavirus capsid at pH 7.5

EMDB-16371: 
African cichlid nackednavirus capsid at pH 5.5

PDB-8aac: 
African cichlid nackednavirus capsid at pH 7.5

PDB-8c0o: 
African cichlid nackednavirus capsid at pH 5.5

EMDB-15274: 
Single Particle cryo-EM of the lipid binding protein P116 (MPN213) from Mycoplasma pneumoniae at 3.3 Angstrom resolution.

EMDB-15275: 
Single Particle cryo-EM of the empty lipid binding protein P116 (MPN213) from Mycoplasma pneumoniae at 4 Angstrom resolution

EMDB-15276: 
Single Particle cryo-EM of the lipid binding protein P116 (MPN213) refilled with FBS from Mycoplasma pneumoniae at 3.5 Angstrom resolution.

EMDB-15277: 
Single Particle cryo-EM of the lipid binding protein P116 (MPN213) from Mycoplasma pneumoniae bound to HDL.

PDB-8a9a: 
Single Particle cryo-EM of the lipid binding protein P116 (MPN213) from Mycoplasma pneumoniae at 3.3 Angstrom resolution.

PDB-8a9b: 
Single Particle cryo-EM of the empty lipid binding protein P116 (MPN213) from Mycoplasma pneumoniae at 4 Angstrom resolution

EMDB-27243: 
Cryo-EM map of MORF-WHs in complex with 197bp nucleosome aided by scFv
EMDB-14144: 
Influenza A/H7N9 polymerase elongation complex
EMDB-14222: 
Early transcription elongation state of influenza A/H7N9 polymerase backtracked due to double incoproation of nucleotide analogue T1106 and with singly incoporated T1106 at the +1 position

EMDB-14240: 
Early transcription elongation state of influenza B polymerase backtracked due to double incoproation of nucleotide analogue T1106

EMDB-14857: 
Symmetric dimer of influenza A/H7N9 polymerase bound to 5' vRNA hook

EMDB-14858: 
Influenza A/H7N9 polymerase apo-protein dimer complex
EMDB-15984: 
Early transcription elongation state of influenza B/Mem polymerase backtracked due to double incoproation of nucleotide analogue T1106 and with singly incoporated T1106 at the U +1 position
EMDB-15996: 
Early transcription elongation state of influenza B/Mem polymerase backtracked due to double incoproation of nucleotide analogue T1106 and with singly incoporated T1106 at the C +1 position
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
