-Search query
-Search result
Showing 1 - 50 of 3,341 items for (author: liu & le)

EMDB-31367: 
Structure of mumps virus nucleoprotein without C-arm

EMDB-42456: 
Omicron-S-MERS-RBD

EMDB-42948: 
Microtubule protofilament reconstruction in CCP5:microtubule class#1 complex

EMDB-44544: 
Microtubule protofilament reconstruction in CCP5:microtubule class#2 complex

EMDB-44545: 
Microtubule Protofilament reconstruction in CCP5:microtubule class#3 complex

EMDB-60397: 
EcGK filament at APO state. (CASP target)

PDB-8zri: 
EcGK filament at APO state.

EMDB-37515: 
Cryo-EM structure of inward-open state human norepinephrine transporter NET bound with antidepressant desipramine in KCl condition.

EMDB-37520: 
Cryo-EM structure of inward-open state human norepinephrine transporter NET bound with norepinephrine in nanodisc.

EMDB-39729: 
Cryo-EM structure of human norepinephrine transporter NET in the presence of the antidepressant atomoxetine in an outward-open state at resolution of 3.4 angstrom.

PDB-8wgr: 
Cryo-EM structure of inward-open state human norepinephrine transporter NET bound with antidepressant desipramine in KCl condition.

PDB-8wgx: 
Cryo-EM structure of inward-open state human norepinephrine transporter NET bound with norepinephrine in nanodisc.

PDB-8z1l: 
Cryo-EM structure of human norepinephrine transporter NET in the presence of the antidepressant atomoxetine in an outward-open state at resolution of 3.4 angstrom.

EMDB-60346: 
EcGK bundle at APO state. (CASP target)

PDB-8zpj: 
EcGK bundle at APO state.

EMDB-43551: 
CCHFV GP38 bound with ADI-46143 and ADI-46158 Fabs

EMDB-43552: 
CCHFV GP38 bound with ADI-58062 and ADI-63530 Fabs

EMDB-43553: 
CCHFV GP38 bound with ADI-58026 and ADI-63547 Fabs

EMDB-43604: 
CCHFV GP38 bound to ADI-46152 and ADI-58048 Fabs

PDB-8vww: 
CCHFV GP38 bound to ADI-46152 and ADI-58048 Fabs

EMDB-50409: 
Subtomogram average of 80S ribosomes in native S. cerevisiae

EMDB-50415: 
Subtomogram average of 80S ribosomes in S. cerevisiae under acute glucose starvation

EMDB-41260: 
SARS-CoV-2 BA.1 S-6P-no-RBD
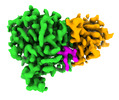
EMDB-42950: 
Structure of CCP5 class1
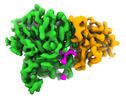
EMDB-42951: 
Structure of CCP5 class2
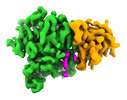
EMDB-42952: 
Structure of CCP5 class3
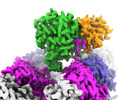
EMDB-42971: 
CCP5 in complex with microtubules class1
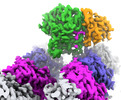
EMDB-42972: 
CCP5 in complex with microtubules class2
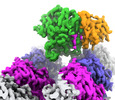
EMDB-42973: 
CCP5 in complex with microtubules class3

PDB-8v3q: 
Structure of CCP5 class1

PDB-8v3r: 
Structure of CCP5 class2

PDB-8v3s: 
Structure of CCP5 class3

PDB-8v4k: 
CCP5 in complex with microtubules class1

PDB-8v4l: 
CCP5 in complex with microtubules class2

PDB-8v4m: 
CCP5 in complex with microtubules class3

EMDB-44395: 
Full-length cross-linked Contactin 2 (CNTN2)

EMDB-44396: 
Cross-linked Contactin 2 Ig1-Ig6

EMDB-44397: 
Full-length cross-linked Contactin 2 (FN1 apart)

PDB-9ba4: 
Full-length cross-linked Contactin 2 (CNTN2)

PDB-9ba5: 
Cross-linked Contactin 2 Ig1-Ig6

EMDB-39025: 
Structure of HCoV-HKU1A spike in the functionally anchored-3up conformation with 3TMPRSS2

EMDB-39026: 
Local structure of HCoV-HKU1A spike in complex with TMPRSS2 and glycan

EMDB-39036: 
Structure of HCoV-HKU1C spike in the functionally anchored-1up conformation with 1TMPRSS2

EMDB-39037: 
Structure of HCoV-HKU1C spike in the functionally anchored-2up conformation with 2TMPRSS2

EMDB-39038: 
Structure of HCoV-HKU1C spike in the functionally anchored-3up conformation with 2TMPRSS2

EMDB-39039: 
Structure of HCoV-HKU1C spike in the functionally anchored-3up conformation with 3TMPRSS2

EMDB-39040: 
Local structure of HCoV-HKU1C spike in complex with TMPRSS2 and glycan

EMDB-39041: 
Structure of HCoV-HKU1C spike in the inactive-closed conformation

EMDB-39042: 
Structure of HCoV-HKU1C spike in the inactive-1up conformation

EMDB-39043: 
Structure of HCoV-HKU1C spike in the inactive-2up conformation
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
