-Search query
-Search result
Showing 1 - 50 of 9,249 items for (author: lin & k)
EMDB-42603: 
Human p97/VCP structure with a triazole inhibitor (NSC799462/hexamer)
EMDB-42625: 
Human p97/VCP R155H mutant structure with a triazole inhibitor (NSC804515)
EMDB-42626: 
Human p97/VCP R155H mutant structure with a triazole inhibitor (NSC819701/up)
EMDB-42627: 
Human p97/VCP R155H mutant structure with a triazole inhibitor (NSC819701/down)
EMDB-44748: 
Human p97/VCP structure with a triazole inhibitor (NSC799462/dodecamer)

EMDB-44551: 
Map of eastern equine encephalitis virus q3 spike protein in complex with VLDLR without masked refinement
EMDB-60628: 
Carazolol-activated human beta3 adrenergic receptor
EMDB-60629: 
Epinephrine-activated human beta3 adrenergic receptor

EMDB-42291: 
Structure of the human INTS9-INTS11-BRAT1 complex

EMDB-42292: 
Structure of the Drosophila IntS11-CG7044(dBRAT1) complex

EMDB-50124: 
Mammalian ternary complex of a translating 80S ribosome, NAC and NatA/E

EMDB-50125: 
Mammalian quaternary complex of a translating 80S ribosome, NAC, MetAP1 and NatA/E

EMDB-50126: 
Mammalian quaternary complex of a translating 80S ribosome, NAC, MetAP1 and NatA/E-HYPK

EMDB-18158: 
Sharpened map of the Gallus gallus 80S rotated ribosome in complex with eEF2 and SERBP1 (cryoSPARC)

EMDB-18159: 
Focused map of the 40S ribosomal subunit head of the Gallus gallus (rotated)

EMDB-18160: 
Focused map of the 40S ribosomal subunit body of the Gallus gallus (rotated)

EMDB-18161: 
Focused map of the 60S ribosomal subunit ot the Gallus gallus

EMDB-18165: 
Sharpened map of the Gallus gallus 80S non-rotated ribosome (cryoSPARC)

EMDB-18166: 
Focused map of the 40S ribosomal subunit head of the Gallus gallus (non-rotated)

EMDB-18167: 
Focused map of the 40S ribosomal subunit body of the Gallus gallus (non-rotated)

EMDB-18168: 
Composite map of the Gallus gallus 80S non-rotated ribosome

EMDB-18169: 
Composite map of the Gallus gallus 80S rotated ribosome in complex with eEF2 and SERBP1
EMDB-18621: 
ASCT2 trimer in lipid nanodiscs with bound glutamine and Na+ ions in the outward-facing state (OFS)
EMDB-18622: 
ASCT2 protomer in lipid nanodiscs with bound glutamine and Na+ ions in the outward-facing state (OFS.1)
EMDB-18623: 
ASCT2 protomer in lipid nanodiscs with bound glutamine and Na+ ions in the outward-facing state (OFS.2)
EMDB-18624: 
ASCT2 protomer in lipid nanodiscs with bound glutamine and Na+ ions in the outward-facing state (OFS.3)
EMDB-18625: 
ASCT2 protomer in lipid nanodiscs with bound glutamine and Na+ ions in the intermediate outward-facing state (iOFS-up)
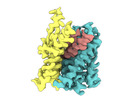
EMDB-18626: 
ASCT2 protomer in lipid nanodiscs with bound glutamine and Na+ ions in the intermediate outward-facing state (iOFS-down)
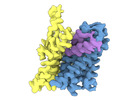
EMDB-18627: 
ASCT2 protomer in lipid nanodiscs under low Na+ concentration in the outward-facing state (OFS)
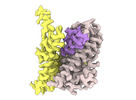
EMDB-18628: 
ASCT2 protomer in lipid nanodiscs under low Na+ concentration in the intermediate outward-facing state (iOFS-up)

PDB-8qro: 
ASCT2 trimer in lipid nanodiscs with bound glutamine and Na+ ions in the outward-facing state (OFS)

PDB-8qrp: 
ASCT2 protomer in lipid nanodiscs with bound glutamine and Na+ ions in the outward-facing state (OFS.1)

PDB-8qrq: 
ASCT2 protomer in lipid nanodiscs with bound glutamine and Na+ ions in the outward-facing state (OFS.2)

PDB-8qrr: 
ASCT2 protomer in lipid nanodiscs with bound glutamine and Na+ ions in the outward-facing state (OFS.3)

PDB-8qrs: 
ASCT2 protomer in lipid nanodiscs with bound glutamine and Na+ ions in the intermediate outward-facing state (iOFS-up)

PDB-8qru: 
ASCT2 protomer in lipid nanodiscs with bound glutamine and Na+ ions in the intermediate outward-facing state (iOFS-down)

PDB-8qrv: 
ASCT2 protomer in lipid nanodiscs under low Na+ concentration in the outward-facing state (OFS)

PDB-8qrw: 
ASCT2 protomer in lipid nanodiscs under low Na+ concentration in the intermediate outward-facing state (iOFS-up)

EMDB-42050: 
Structure of eastern equine encephalitis virus VLP in complex with VLDLR LA1

EMDB-42055: 
Structure of eastern equine encephalitis virus VLP unliganded quasi-threefold spike protein

EMDB-38845: 
Icosahedrally averaged cryo-EM reconstruction of PhiKZ capsid before applying the "block-based" reconstruction method

EMDB-38846: 
Block 1 of PhiKZ capsid

EMDB-38848: 
Block 2 of PhiKZ capsid

EMDB-39002: 
Composite cryo-EM map of PhiKZ capsid after applying the "block-based" reconstruction method

PDB-8y6v: 
Near-atomic structure of icosahedrally averaged jumbo bacteriophage PhiKZ capsid

EMDB-41770: 
Apo form of human ATE1

EMDB-42071: 
human ATE1 in complex with Arg-tRNA and a peptide substrate

PDB-8tzv: 
Apo form of human ATE1

PDB-8uau: 
human ATE1 in complex with Arg-tRNA and a peptide substrate
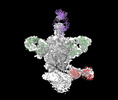
EMDB-44246: 
Cryo-EM structure of HIV-1 JRFL v6 Env in complex with vaccine-elicited, Membrane Proximal External Region (MPER) directed antibody DH1317.4.
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
