[English] 日本語
 Yorodumi
Yorodumi- EMDB-42055: Structure of eastern equine encephalitis virus VLP unliganded qua... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of eastern equine encephalitis virus VLP unliganded quasi-threefold spike protein | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Eastern Equine Encephalitis Virus / VIRUS LIKE PARTICLE | |||||||||
| Function / homology |  Function and homology information Function and homology informationtogavirin / T=4 icosahedral viral capsid / host cell endoplasmic reticulum / channel activity / monoatomic ion transmembrane transport / symbiont-mediated suppression of host toll-like receptor signaling pathway / host cell Golgi apparatus / entry receptor-mediated virion attachment to host cell / symbiont-mediated suppression of host gene expression / serine-type endopeptidase activity ...togavirin / T=4 icosahedral viral capsid / host cell endoplasmic reticulum / channel activity / monoatomic ion transmembrane transport / symbiont-mediated suppression of host toll-like receptor signaling pathway / host cell Golgi apparatus / entry receptor-mediated virion attachment to host cell / symbiont-mediated suppression of host gene expression / serine-type endopeptidase activity / fusion of virus membrane with host endosome membrane / symbiont entry into host cell / host cell plasma membrane / host cell nucleus / virion membrane / structural molecule activity / proteolysis / RNA binding Similarity search - Function | |||||||||
| Biological species |   Eastern equine encephalitis virus Eastern equine encephalitis virus | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.0 Å | |||||||||
 Authors Authors | Abraham J / Yang P / Li W / Fan X / Pan J | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2024 Journal: Nat Commun / Year: 2024Title: Structural basis for VLDLR recognition by eastern equine encephalitis virus. Authors: Pan Yang / Wanyu Li / Xiaoyi Fan / Junhua Pan / Colin J Mann / Haley Varnum / Lars E Clark / Sarah A Clark / Adrian Coscia / Himanish Basu / Katherine Nabel Smith / Vesna Brusic / Jonathan Abraham /   Abstract: Eastern equine encephalitis virus (EEEV) is the most virulent alphavirus that infects humans, and many survivors develop neurological sequelae, including paralysis and intellectual disability. ...Eastern equine encephalitis virus (EEEV) is the most virulent alphavirus that infects humans, and many survivors develop neurological sequelae, including paralysis and intellectual disability. Alphavirus spike proteins comprise trimers of heterodimers of glycoproteins E2 and E1 that mediate binding to cellular receptors and fusion of virus and host cell membranes during entry. We recently identified very-low density lipoprotein receptor (VLDLR) and apolipoprotein E receptor 2 (ApoER2) as cellular receptors for EEEV and a distantly related alphavirus, Semliki Forest virus (SFV). Here, we use single-particle cryo-electron microscopy (cryo-EM) to determine structures of the EEEV and SFV spike glycoproteins bound to the VLDLR ligand-binding domain and found that EEEV and SFV interact with the same cellular receptor through divergent binding modes. Our studies suggest that the ability of LDLR-related proteins to interact with viral spike proteins through very small footprints with flexible binding modes results in a low evolutionary barrier to the acquisition of LDLR-related proteins as cellular receptors for diverse sets of viruses. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_42055.map.gz emd_42055.map.gz | 104.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-42055-v30.xml emd-42055-v30.xml emd-42055.xml emd-42055.xml | 21.1 KB 21.1 KB | Display Display |  EMDB header EMDB header |
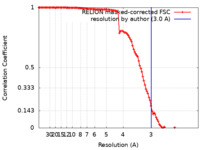
| FSC (resolution estimation) |  emd_42055_fsc.xml emd_42055_fsc.xml | 11.1 KB | Display |  FSC data file FSC data file |
| Images |  emd_42055.png emd_42055.png | 62.7 KB | ||
| Filedesc metadata |  emd-42055.cif.gz emd-42055.cif.gz | 7 KB | ||
| Others |  emd_42055_half_map_1.map.gz emd_42055_half_map_1.map.gz emd_42055_half_map_2.map.gz emd_42055_half_map_2.map.gz | 90.6 MB 90.6 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-42055 http://ftp.pdbj.org/pub/emdb/structures/EMD-42055 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-42055 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-42055 | HTTPS FTP |
-Related structure data
| Related structure data |  8ua9MC  8ua4C  8ua8C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_42055.map.gz / Format: CCP4 / Size: 115.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_42055.map.gz / Format: CCP4 / Size: 115.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.06 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_42055_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_42055_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Eastern equine encephalitis virus
| Entire | Name:   Eastern equine encephalitis virus Eastern equine encephalitis virus |
|---|---|
| Components |
|
-Supramolecule #1: Eastern equine encephalitis virus
| Supramolecule | Name: Eastern equine encephalitis virus / type: virus / ID: 1 / Parent: 0 / Macromolecule list: #4 / NCBI-ID: 11021 / Sci species name: Eastern equine encephalitis virus / Virus type: VIRUS-LIKE PARTICLE / Virus isolate: STRAIN / Virus enveloped: Yes / Virus empty: No |
|---|
-Macromolecule #1: Envelope glycoprotein E1
| Macromolecule | Name: Envelope glycoprotein E1 / type: protein_or_peptide / ID: 1 / Details: Spike glycoprotein E1 / Number of copies: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Eastern equine encephalitis virus Eastern equine encephalitis virus |
| Molecular weight | Theoretical: 47.984246 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: YEHTAVMPNK VGIPYKALVE RPGYAPVHLQ IQLVNTRIIP STNLEYITCK YKTKVPSPVV KCCGATQCTS KPHPDYQCQV FSGVYPFMW GGAYCFCDTE NTQMSEAYVE RSEECSIDHA KAYKVHTGTV QAMVNITYGS VSWRSADVYV NGETPAKIGD A KLIIGPLS ...String: YEHTAVMPNK VGIPYKALVE RPGYAPVHLQ IQLVNTRIIP STNLEYITCK YKTKVPSPVV KCCGATQCTS KPHPDYQCQV FSGVYPFMW GGAYCFCDTE NTQMSEAYVE RSEECSIDHA KAYKVHTGTV QAMVNITYGS VSWRSADVYV NGETPAKIGD A KLIIGPLS SAWSPFDNKV VVYGHEVYNY DFPEYGTGKA GSFGDLQSRT STSNDLYANT NLKLQRPQAG IVHTPFTQVP SG FERWKKD KGAPLNDVAP FGCSIALEPL RAENCAVGSI PISIDIPDAA FTRISETPTV SDLECKITEC TYAFDFGGIA TVA YKSSKA GNCPIHSPSG VAVIKENDVT LAESGSFTFH FSTANIHPAF KLQVCTSAVT CKGDCKPPKD HIVDYPAQHT ESFT SAISA TAWSWIKVLV GGTSAFIVLG LIATAVVALV LFFHRH UniProtKB: Structural polyprotein |
-Macromolecule #2: Structural polyprotein
| Macromolecule | Name: Structural polyprotein / type: protein_or_peptide / ID: 2 / Number of copies: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Eastern equine encephalitis virus Eastern equine encephalitis virus |
| Molecular weight | Theoretical: 46.818742 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: DLDTHFTQYK LARPYIADCP NCGHSRCDSP IAIEEVRGDA HAGVIRIQTS AMFGLKTDGV DLAYMSFMNG KTQKSIKIDN LHVRTSAPC SLVSHHGYYI LAQCPPGDTV TVGFHDGPNR HTCTVAHKVE FRPVGREKYR HPPEHGVELP CNRYTHKRAD Q GHYVEMHQ ...String: DLDTHFTQYK LARPYIADCP NCGHSRCDSP IAIEEVRGDA HAGVIRIQTS AMFGLKTDGV DLAYMSFMNG KTQKSIKIDN LHVRTSAPC SLVSHHGYYI LAQCPPGDTV TVGFHDGPNR HTCTVAHKVE FRPVGREKYR HPPEHGVELP CNRYTHKRAD Q GHYVEMHQ PGLVADHSLL SIHSAKVKIT VPSGAQVKYY CKCPDVRKGI TSSDHTTTCT DVKQCRAYLI DNKKWVYNSG RL PRGEGDT FKGKLHVPFV PVKAKCIATL APEPLVEHKH RTLILHLHPD HPTLLTTRSL GSDANPTRQW IERPTTVNFT VTG EGLEYT WGNHPPKRVW AQESGEGNPH GWPHEVVVYY YNRYPLTTII GLCTCVAIIM VSCVTSVWLL CRTRNLCITP YKLA PNAQV PILLALLCCI KPT UniProtKB: Structural polyprotein |
-Macromolecule #3: Structural polyprotein
| Macromolecule | Name: Structural polyprotein / type: protein_or_peptide / ID: 3 / Number of copies: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Eastern equine encephalitis virus Eastern equine encephalitis virus |
| Molecular weight | Theoretical: 6.147061 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: TVMCVLANIT FPCDQPPCMP CCYEKNPHET LTMLEQNYDS RAYDQLLDAA VKCN UniProtKB: Structural polyprotein |
-Macromolecule #4: Capsid protein
| Macromolecule | Name: Capsid protein / type: protein_or_peptide / ID: 4 / Number of copies: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Eastern equine encephalitis virus Eastern equine encephalitis virus |
| Molecular weight | Theoretical: 29.21293 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MFPYPTLNYP PMAPINPMAY RDPNPPRRRW RPFRPPLAAQ IEDLRRSIAN LTLKQRAPNP PAGPPAKRKK PAPSLSLRRK KKRPPPPAK KQKRKPKPGK RQRMCMKLES DKTFPIMLNG QVNGYACVVG GRVFKPLHVE GRIDNEQLAA IKLKKASIYD L EYGDVPQC ...String: MFPYPTLNYP PMAPINPMAY RDPNPPRRRW RPFRPPLAAQ IEDLRRSIAN LTLKQRAPNP PAGPPAKRKK PAPSLSLRRK KKRPPPPAK KQKRKPKPGK RQRMCMKLES DKTFPIMLNG QVNGYACVVG GRVFKPLHVE GRIDNEQLAA IKLKKASIYD L EYGDVPQC MKSDTLQYTS DKPPGFYNWH HGAVQYENNR FTVPRGVGGK GDSGRPILDN KGRVVAIVLG GVNEGSRTAL SV VTWNQKG VTVKDTPEGS EPW UniProtKB: Structural polyprotein |
-Macromolecule #5: 2-acetamido-2-deoxy-beta-D-glucopyranose
| Macromolecule | Name: 2-acetamido-2-deoxy-beta-D-glucopyranose / type: ligand / ID: 5 / Number of copies: 9 / Formula: NAG |
|---|---|
| Molecular weight | Theoretical: 221.208 Da |
| Chemical component information |  ChemComp-NAG: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.2 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 54.1 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.5 µm / Nominal defocus min: 1.0 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller









 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)