-Search query
-Search result
Showing all 41 items for (author: kinoshita & m)

EMDB-39761: 
Structure of the S-ring region of the Vibrio flagellar MS-ring protein FliF with 34-fold symmetry applied

EMDB-39763: 
Structure of the S-ring region of the Vibrio flagellar MS-ring protein FliF with 35-fold symmetry applied

EMDB-39764: 
Homomeric 34mer of the Vibrio flagellar MS-ring protein FliF without symmetry imposition

EMDB-39765: 
Homomeric 35mer of the Vibrio flagellar MS-ring protein FliF without symmetry imposition

EMDB-60959: 
Salmonella enterica serovar Typhimurium FliC(G426A)delta(204-292) forming the L-type straight filament

PDB-9iwq: 
Salmonella enterica serovar Typhimurium FliC(G426A)delta(204-292) forming the L-type straight filament

EMDB-36048: 
Cryo-EM structure of hZnT7-Fab complex in zinc-unbound state, determined in outward-facing conformation

EMDB-36049: 
Cryo-EM structure of hZnT7-Fab complex in zinc-bound state, determined in outward-facing conformation

EMDB-36050: 
Cryo-EM structure of hZnT7-Fab complex in zinc-unbound state, determined in heterogeneous conformations- one subunit in an inward-facing and the other in an outward-facing conformation

EMDB-36051: 
Cryo-EM structure of hZnT7-Fab complex in zinc state 2, determined in heterogeneous conformations- one subunit in an inward-facing zinc-bound and the other in an outward-facing zinc-bound conformation

EMDB-36052: 
Cryo-EM structure of hZnT7DeltaHis-loop-Fab complex in zinc-unbound state, determined in outward-facing conformation

EMDB-36053: 
Cryo-EM structure of hZnT7DeltaHis-loop-Fab complex in zinc-bound state, determined in outward-facing conformation

EMDB-36055: 
Cryo-EM structure of hZnT7-Fab complex in zinc state 1, determined in heterogeneous conformations- one subunit in an inward-facing zinc-bound and the other in an outward-facing zinc-unbound conformation

PDB-8j7t: 
Cryo-EM structure of hZnT7-Fab complex in zinc-unbound state, determined in outward-facing conformation

PDB-8j7u: 
Cryo-EM structure of hZnT7-Fab complex in zinc-bound state, determined in outward-facing conformation

PDB-8j7v: 
Cryo-EM structure of hZnT7-Fab complex in zinc-unbound state, determined in heterogeneous conformations- one subunit in an inward-facing and the other in an outward-facing conformation

PDB-8j7w: 
Cryo-EM structure of hZnT7-Fab complex in zinc state 2, determined in heterogeneous conformations- one subunit in an inward-facing zinc-bound and the other in an outward-facing zinc-bound conformation

PDB-8j7x: 
Cryo-EM structure of hZnT7DeltaHis-loop-Fab complex in zinc-unbound state, determined in outward-facing conformation

PDB-8j7y: 
Cryo-EM structure of hZnT7DeltaHis-loop-Fab complex in zinc-bound state, determined in outward-facing conformation

PDB-8j80: 
Cryo-EM structure of hZnT7-Fab complex in zinc state 1, determined in heterogeneous conformations- one subunit in an inward-facing zinc-bound and the other in an outward-facing zinc-unbound conformation

EMDB-32407: 
RNA polymerase II elongation complex bound with Elf1 and Spt4/5, stalled at SHL(-4) of the nucleosome

EMDB-32408: 
RNA polymerase II elongation complex bound with Elf1 and Spt4/5, stalled at SHL(-3.5) of the nucleosome

EMDB-32409: 
RNA polymerase II elongation complex bound with Elf1 and Spt4/5, stalled at SHL(-3) of the nucleosome

EMDB-34685: 
RNA polymerase II elongation complex bound with Rad26 and Elf1, stalled at SHL(-3.5) of the nucleosome

PDB-7wbv: 
RNA polymerase II elongation complex bound with Elf1 and Spt4/5, stalled at SHL(-4) of the nucleosome

PDB-7wbw: 
RNA polymerase II elongation complex bound with Elf1 and Spt4/5, stalled at SHL(-3.5) of the nucleosome

PDB-7wbx: 
RNA polymerase II elongation complex bound with Elf1 and Spt4/5, stalled at SHL(-3) of the nucleosome

PDB-8he5: 
RNA polymerase II elongation complex bound with Rad26 and Elf1, stalled at SHL(-3.5) of the nucleosome

EMDB-34430: 
Cryo-EM structure of the human RAD52 protein

PDB-8h1p: 
Cryo-EM structure of the human RAD52 protein

EMDB-30360: 
34-fold symmetry salmonella MS ring

EMDB-30361: 
11 fold-syymetry of salmonella MS ring

EMDB-30363: 
Without imposing symmetry of salmonella MS ring

EMDB-30613: 
Structure of the bacterial flagellar basal body

EMDB-30940: 
Salmonella flagella MS-ring protein FliF 1-503 having C33 symmetry

EMDB-30941: 
Salmonella flagella MS-ring protein FliF 1-456 having C35 symmetry

EMDB-30942: 
Salmonella flagella MS-ring protein FliF 1-456 having C34 symmetry
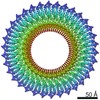
EMDB-30612: 
34-fold symmetry Salmonella S ring formed by full-length FliF

PDB-7d84: 
34-fold symmetry Salmonella S ring formed by full-length FliF

EMDB-30378: 
Salmonella FliF-FliG ring complex

EMDB-30379: 
11-fold symmetry of Salmonella FliF-FliG ring complex
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
