+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 7d84 | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | 34-fold symmetry Salmonella S ring formed by full-length FliF | ||||||||||||||||||||||||||||||||||||
 Components Components | Flagellar M-ring protein | ||||||||||||||||||||||||||||||||||||
 Keywords Keywords | MEMBRANE PROTEIN / Flagellar motor / Type III protein export system / Transmembrane protein Complex / Torque generation | ||||||||||||||||||||||||||||||||||||
| Function / homology |  Function and homology information Function and homology informationbacterial-type flagellum basal body, MS ring / cytoskeletal motor activity / bacterial-type flagellum-dependent cell motility / plasma membrane Similarity search - Function | ||||||||||||||||||||||||||||||||||||
| Biological species |  Salmonella enterica serovar Typhimurium (bacteria) Salmonella enterica serovar Typhimurium (bacteria) | ||||||||||||||||||||||||||||||||||||
| Method | ELECTRON MICROSCOPY / single particle reconstruction / cryo EM / Resolution: 3.7 Å | ||||||||||||||||||||||||||||||||||||
 Authors Authors | Kawamoto, A. / Miyata, T. / Makino, F. / Kinoshita, M. / Minamino, T. / Imada, K. / Kato, T. / Namba, K. | ||||||||||||||||||||||||||||||||||||
| Funding support |  Japan, 11items Japan, 11items
| ||||||||||||||||||||||||||||||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2021 Journal: Nat Commun / Year: 2021Title: Native flagellar MS ring is formed by 34 subunits with 23-fold and 11-fold subsymmetries. Authors: Akihiro Kawamoto / Tomoko Miyata / Fumiaki Makino / Miki Kinoshita / Tohru Minamino / Katsumi Imada / Takayuki Kato / Keiichi Namba /  Abstract: The bacterial flagellar MS ring is a transmembrane complex acting as the core of the flagellar motor and template for flagellar assembly. The C ring attached to the MS ring is involved in torque ...The bacterial flagellar MS ring is a transmembrane complex acting as the core of the flagellar motor and template for flagellar assembly. The C ring attached to the MS ring is involved in torque generation and rotation switch, and a large symmetry mismatch between these two rings has been a long puzzle, especially with respect to their role in motor function. Here, using cryoEM structural analysis of the flagellar basal body and the MS ring formed by full-length FliF from Salmonella enterica, we show that the native MS ring is formed by 34 FliF subunits with no symmetry variation. Symmetry analysis of the C ring shows a variation with a peak at 34-fold, suggesting flexibility in C ring assembly. Finally, our data also indicate that FliF subunits assume two different conformations, contributing differentially to the inner and middle parts of the M ring and thus resulting in 23- and 11-fold subsymmetries in the inner and middle M ring, respectively. The internal core of the M ring, formed by 23 subunits, forms a hole of the right size to accommodate the protein export gate. | ||||||||||||||||||||||||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  7d84.cif.gz 7d84.cif.gz | 1 MB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb7d84.ent.gz pdb7d84.ent.gz | 812 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  7d84.json.gz 7d84.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/d8/7d84 https://data.pdbj.org/pub/pdb/validation_reports/d8/7d84 ftp://data.pdbj.org/pub/pdb/validation_reports/d8/7d84 ftp://data.pdbj.org/pub/pdb/validation_reports/d8/7d84 | HTTPS FTP |
|---|
-Related structure data
| Related structure data | 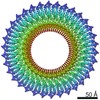 30612MC M: map data used to model this data C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
|
|---|---|
| 1 |
|
- Components
Components
| #1: Protein | Mass: 61295.645 Da / Num. of mol.: 34 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Salmonella enterica serovar Typhimurium (bacteria) Salmonella enterica serovar Typhimurium (bacteria)Gene: fliF, fla AII.1, fla BI, STM1969 / Production host:  |
|---|
-Experimental details
-Experiment
| Experiment | Method: ELECTRON MICROSCOPY |
|---|---|
| EM experiment | Aggregation state: PARTICLE / 3D reconstruction method: single particle reconstruction |
- Sample preparation
Sample preparation
| Component | Name: transmembrane protein complex made of FliF / Type: COMPLEX Details: the MS ring formed by full-length FliF from Salmonella enterica serovar Typhimurium Entity ID: all / Source: RECOMBINANT | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Molecular weight | Experimental value: NO | ||||||||||||||||||||
| Source (natural) | Organism:  Salmonella typhimurium (strain LT2 / SGSC1412 / ATCC 700720) (bacteria) Salmonella typhimurium (strain LT2 / SGSC1412 / ATCC 700720) (bacteria) | ||||||||||||||||||||
| Source (recombinant) | Organism:  | ||||||||||||||||||||
| Buffer solution | pH: 8 | ||||||||||||||||||||
| Buffer component |
| ||||||||||||||||||||
| Specimen | Embedding applied: NO / Shadowing applied: NO / Staining applied: NO / Vitrification applied: YES | ||||||||||||||||||||
| Specimen support | Grid material: COPPER / Grid mesh size: 200 divisions/in. / Grid type: Quantifoil R1.2/1.3 | ||||||||||||||||||||
| Vitrification | Instrument: FEI VITROBOT MARK III / Cryogen name: ETHANE / Humidity: 100 % / Chamber temperature: 277 K |
- Electron microscopy imaging
Electron microscopy imaging
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
|---|---|
| Microscopy | Model: FEI TITAN KRIOS |
| Electron gun | Electron source:  FIELD EMISSION GUN / Accelerating voltage: 300 kV / Illumination mode: FLOOD BEAM FIELD EMISSION GUN / Accelerating voltage: 300 kV / Illumination mode: FLOOD BEAM |
| Electron lens | Mode: BRIGHT FIELD / Nominal magnification: 75000 X / Nominal defocus max: 3000 nm / Nominal defocus min: 1000 nm / Cs: 2.7 mm / Alignment procedure: COMA FREE |
| Specimen holder | Cryogen: NITROGEN / Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER |
| Image recording | Electron dose: 90 e/Å2 / Film or detector model: FEI FALCON II (4k x 4k) / Num. of grids imaged: 1 / Num. of real images: 1589 |
| Image scans | Width: 4096 / Height: 4096 / Movie frames/image: 7 / Used frames/image: 1-6 |
- Processing
Processing
| Software | Name: PHENIX / Version: 1.15.2_3472: / Classification: refinement | ||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| EM software |
| ||||||||||||||||||||||||||||||||||||||||||||||||
| CTF correction | Type: PHASE FLIPPING AND AMPLITUDE CORRECTION | ||||||||||||||||||||||||||||||||||||||||||||||||
| Particle selection | Num. of particles selected: 339861 | ||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Point symmetry: C34 (34 fold cyclic) | ||||||||||||||||||||||||||||||||||||||||||||||||
| 3D reconstruction | Resolution: 3.7 Å / Resolution method: FSC 0.143 CUT-OFF / Num. of particles: 38889 / Num. of class averages: 1 / Symmetry type: POINT | ||||||||||||||||||||||||||||||||||||||||||||||||
| Atomic model building | Protocol: RIGID BODY FIT / Space: REAL | ||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
|
 Movie
Movie Controller
Controller

















 PDBj
PDBj
