-Search query
-Search result
Showing 1 - 50 of 4,078 items for (author: john & c)
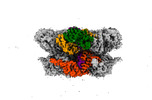
EMDB-42603: 
Human p97/VCP structure with a triazole inhibitor (NSC799462/hexamer)
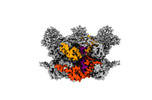
EMDB-42625: 
Human p97/VCP R155H mutant structure with a triazole inhibitor (NSC804515)
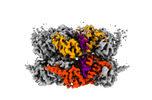
EMDB-42626: 
Human p97/VCP R155H mutant structure with a triazole inhibitor (NSC819701/up)
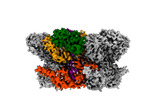
EMDB-42627: 
Human p97/VCP R155H mutant structure with a triazole inhibitor (NSC819701/down)
EMDB-44748: 
Human p97/VCP structure with a triazole inhibitor (NSC799462/dodecamer)
EMDB-45078: 
E.coli GroEL + PBZ1587 inhibitor
EMDB-45079: 
E.coli GroEL apoenzyme
EMDB-45080: 
E.Faecium GroEL

EMDB-45098: 
E.coli GroEL + PBZ1587 inhibitor C1 reconstruction

PDB-9c0b: 
E.coli GroEL + PBZ1587 inhibitor

PDB-9c0c: 
E.coli GroEL apoenzyme

PDB-9c0d: 
E.Faecium GroEL

EMDB-44482: 
Cryo-EM structure of the HIV-1 JR-FL IDL Env trimer in complex with PGT122 Fab
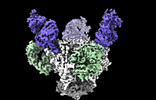
EMDB-44484: 
Cryo-EM structure of the HIV-1 BG505 IDL Env trimer in complex with 3BNC117 and 10-1074 Fabs
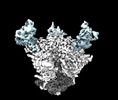
EMDB-44491: 
Cryo-EM structure of the HIV-1 WITO IDL Env trimer in complex with PGT122 Fab

PDB-9ber: 
Cryo-EM structure of the HIV-1 JR-FL IDL Env trimer in complex with PGT122 Fab

PDB-9bew: 
Cryo-EM structure of the HIV-1 BG505 IDL Env trimer in complex with 3BNC117 and 10-1074 Fabs

PDB-9bf6: 
Cryo-EM structure of the HIV-1 WITO IDL Env trimer in complex with PGT122 Fab
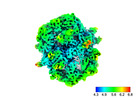
EMDB-45235: 
Subtomogram average (C1) of fatty acid synthase from S.cerevisiae prepared using cryo-plasmaFIB milling

EMDB-42400: 
RORC mRNA 3'UTR riboswitch A97G/G98A mutant class C

EMDB-42401: 
RORC mRNA 3'UTR riboswitch 77-GA mutant class A

EMDB-42403: 
RORC mRNA 3'UTR riboswitch 117-AC mutant class C

EMDB-42404: 
RORC mRNA 3'UTR riboswitch 117-AC mutant class B

EMDB-43139: 
SARS-CoV-2 Spike S2 bound to Fab 54043-5

PDB-8vcr: 
SARS-CoV-2 Spike S2 bound to Fab 54043-5

EMDB-43551: 
CCHFV GP38 bound with ADI-46143 and ADI-46158 Fabs

EMDB-43552: 
CCHFV GP38 bound with ADI-58062 and ADI-63530 Fabs

EMDB-43553: 
CCHFV GP38 bound with ADI-58026 and ADI-63547 Fabs

EMDB-43604: 
CCHFV GP38 bound to ADI-46152 and ADI-58048 Fabs

PDB-8vww: 
CCHFV GP38 bound to ADI-46152 and ADI-58048 Fabs

EMDB-43435: 
Prefusion stabilized structure of the SARS-CoV-2 fusion machinery

EMDB-43436: 
Prefusion stabilized structure of the SARS-CoV-2 fusion machinery

EMDB-43437: 
Prefusion stabilized structure of the SARS-CoV-2 fusion machinery

PDB-8vq9: 
Prefusion stabilized structure of the SARS-CoV-2 fusion machinery

PDB-8vqa: 
Prefusion stabilized structure of the SARS-CoV-2 fusion machinery

PDB-8vqb: 
Prefusion stabilized structure of the SARS-CoV-2 fusion machinery

EMDB-43011: 
Phosphorylated, ATP-bound, E1371Q human cystic fibrosis transmembrane conductance regulator (E1371Q-CFTR)

EMDB-43014: 
Phosphorylated, ATP-bound, inhibitor 172-bound E1371Q human cystic fibrosis transmembrane conductance regulator

EMDB-16904: 
Structure of the MlaCD complex (1:6 stoichiometry)

EMDB-16913: 
Structure of the MlaCD complex (2:6 stoichiometry)
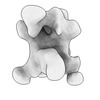
EMDB-42516: 
HIV-1 JR-FL NFL.664 soluble trimer in complex with polyclonal Fab from rabbit U5902

EMDB-42517: 
HIV-1 JR-FL NFL.664 soluble trimer in complex with polyclonal Fab from rabbit U5756

EMDB-42518: 
HIV-1 1086c NFL.664 soluble trimer in complex with polyclonal Fab from rabbit U5403

EMDB-42519: 
HIV-1 1086c NFL.664 soluble trimer in complex with polyclonal Fab from rabbit U5919

EMDB-44351: 
Synaptic Vesicle V-ATPase with synaptophysin and SidK, State 3, V1

PDB-9b8p: 
Synaptic Vesicle V-ATPase with synaptophysin and SidK, State 3, V1

EMDB-41041: 
Open human HCN1 F186C S264C bound to cAMP, reconstituted in LMNG + SPL

PDB-8t50: 
Open human HCN1 F186C S264C bound to cAMP, reconstituted in LMNG + SPL

EMDB-17295: 
Stabilised BA.1 SARS-CoV-2 spike with H6 nanobodies in '3 up' RBD conformation

PDB-8oyt: 
Stabilised BA.1 SARS-CoV-2 spike with H6 nanobodies in '3 up' RBD conformation
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
