-Search query
-Search result
Showing 1 - 50 of 184 items for (author: homa & fl)

EMDB-18779: 
Structure of the non-mitochondrial citrate synthase from Ananas comosus

EMDB-17309: 
In situ cryoEM structure of Prototype Foamy Virus Env trimer

EMDB-17311: 
In situ cryoEM structure of Prototype Foamy Virus Env dimer of trimers

EMDB-17312: 
In situ cryoEM structure of the Prototype Foamy Virus capsid, icosahedral map

EMDB-17313: 
In situ cryoEM structure of the Prototype Foamy Virus capsid, pentamer localised reconstruction

EMDB-17314: 
In situ cryoEM structure of the Prototype Foamy Virus capsid, hexamer 1 localised reconstruction

EMDB-17315: 
In situ cryoEM structure of the Prototype Foamy Virus capsid, hexamer 2 localised reconstruction

EMDB-17316: 
In situ subtomogram average of Prototype Foamy Virus Env trimer

EMDB-17317: 
In situ subtomogram average of Prototype Foamy Virus Env pentamer of trimers

EMDB-17318: 
In situ subtomogram average of Prototype Foamy Virus Env hexamer of trimers

EMDB-17319: 
In situ subtomogram average of the Prototype Foamy Virus capsid, wild-type Gag

EMDB-17320: 
In situ subtomogram average of the Prototype Foamy Virus capsid, p68 Gag
EMDB-17321: 
Cryotomogram of Prototype Foamy Virus particles, wild-type Gag
EMDB-17322: 
Cryotomogram of Prototype Foamy Virus particles, p68 Gag

PDB-8ozh: 
In situ cryoEM structure of Prototype Foamy Virus Env trimer

PDB-8ozj: 
In situ cryoEM structure of Prototype Foamy Virus Env dimer of trimers

PDB-8ozk: 
In situ cryoEM structure of the Prototype Foamy Virus capsid, icosahedral map

PDB-8ozl: 
In situ cryoEM structure of the Prototype Foamy Virus capsid, pentamer localised reconstruction

PDB-8ozm: 
In situ cryoEM structure of the Prototype Foamy Virus capsid, hexamer 1 localised reconstruction

PDB-8ozn: 
In situ cryoEM structure of the Prototype Foamy Virus capsid, hexamer 2 localised reconstruction

PDB-8ozp: 
In situ subtomogram average of Prototype Foamy Virus Env pentamer of trimers

PDB-8ozq: 
In situ subtomogram average of Prototype Foamy Virus Env hexamer of trimers

EMDB-28966: 
CryoEM map of de novo designed oligomeric protein C4-71_6x

EMDB-28967: 
CryoEM map of de novo designed oligomeric protein C4-71_8x
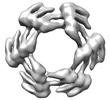
EMDB-28968: 
CryoEM map of de novo designed oligomeric protein C6-71

EMDB-28969: 
CryoEM map of de novo designed oligomeric protein C6-71_6x

EMDB-28970: 
CryoEM map of de novo designed oligomeric protein C6-71_8x

EMDB-28971: 
CryoEM map of de novo designed oligomeric protein C8-71_6x

EMDB-28972: 
CryoEM map of de novo designed oligomeric protein C8-71_8x
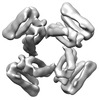
EMDB-28973: 
CryoEM map of de novo designed oligomeric protein C4-81

EMDB-28974: 
CryoEM map of designed oligomeric protein C4-71

EMDB-17449: 
S. cerevisiae nexus-sCMGE after DNA replication initiation

EMDB-17458: 
S. cerevisiae ssDNA-sCMGE after DNA replication initiation

EMDB-17459: 
S. cerevisiae consensus-sCMGE on ssDNA after DNA replication initiation

EMDB-17460: 
S. cerevisiae sCMGE with N-ter Mcm10 density

PDB-8p5e: 
S. cerevisiae nexus-sCMGE after DNA replication initiation

PDB-8p62: 
S. cerevisiae ssDNA-sCMGE after DNA replication initiation

PDB-8p63: 
S. cerevisiae consensus-sCMGE on ssDNA after DNA replication initiation

EMDB-19250: 
Pseudoatomic model of a second-order Sierpinski triangle formed by the citrate synthase from Synechococcus elongatus

EMDB-19251: 
Structure of a first order Sierpinski triangle formed by the H369R mutant of the citrate synthase from Synechococcus elongatus

EMDB-28958: 
CryoEM structure of designed modular protein oligomer C4-131

EMDB-28889: 
CryoEM structure of designed modular protein oligomer C6-79

PDB-8f6r: 
CryoEM structure of designed modular protein oligomer C6-79

EMDB-28888: 
CryoEM structure of designed modular protein oligomer C8-71

PDB-8f6q: 
CryoEM structure of designed modular protein oligomer C8-71

EMDB-16805: 
Cryo-EM structure of PcrV/Fab(30-B8)

EMDB-16807: 
Cryo-EM structure of PcrV/Fab(11-E5)

PDB-8cr9: 
Cryo-EM structure of PcrV/Fab(30-B8)

PDB-8crb: 
Cryo-EM structure of PcrV/Fab(11-E5)

EMDB-17154: 
Cryo-EM structure of CLOCK-BMAL1 bound to a nucleosomal E-box at position SHL+5.8 (consensus and constituent map 1)
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
