+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | CryoEM map of designed oligomeric protein C4-71 | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Synthetic / Self-assembling / Oligomeric / Helical repeats / DE NOVO PROTEIN | |||||||||
| Biological species | synthetic construct (others) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 5.1 Å | |||||||||
 Authors Authors | Redler RL / Edman NI / Baker D / Ekiert DC / Bhabha G | |||||||||
| Funding support |  United States, 2 items United States, 2 items
| |||||||||
 Citation Citation |  Journal: Cell / Year: 2024 Journal: Cell / Year: 2024Title: Modulation of FGF pathway signaling and vascular differentiation using designed oligomeric assemblies. Authors: Natasha I Edman / Ashish Phal / Rachel L Redler / Thomas Schlichthaerle / Sanjay R Srivatsan / Devon Duron Ehnes / Ali Etemadi / Seong J An / Andrew Favor / Zhe Li / Florian Praetorius / Max ...Authors: Natasha I Edman / Ashish Phal / Rachel L Redler / Thomas Schlichthaerle / Sanjay R Srivatsan / Devon Duron Ehnes / Ali Etemadi / Seong J An / Andrew Favor / Zhe Li / Florian Praetorius / Max Gordon / Thomas Vincent / Silvia Marchiano / Leslie Blakely / Chuwei Lin / Wei Yang / Brian Coventry / Derrick R Hicks / Longxing Cao / Neville Bethel / Piper Heine / Analisa Murray / Stacey Gerben / Lauren Carter / Marcos Miranda / Babak Negahdari / Sangwon Lee / Cole Trapnell / Ying Zheng / Charles E Murry / Devin K Schweppe / Benjamin S Freedman / Lance Stewart / Damian C Ekiert / Joseph Schlessinger / Jay Shendure / Gira Bhabha / Hannele Ruohola-Baker / David Baker /   Abstract: Many growth factors and cytokines signal by binding to the extracellular domains of their receptors and driving association and transphosphorylation of the receptor intracellular tyrosine kinase ...Many growth factors and cytokines signal by binding to the extracellular domains of their receptors and driving association and transphosphorylation of the receptor intracellular tyrosine kinase domains, initiating downstream signaling cascades. To enable systematic exploration of how receptor valency and geometry affect signaling outcomes, we designed cyclic homo-oligomers with up to 8 subunits using repeat protein building blocks that can be modularly extended. By incorporating a de novo-designed fibroblast growth factor receptor (FGFR)-binding module into these scaffolds, we generated a series of synthetic signaling ligands that exhibit potent valency- and geometry-dependent Ca release and mitogen-activated protein kinase (MAPK) pathway activation. The high specificity of the designed agonists reveals distinct roles for two FGFR splice variants in driving arterial endothelium and perivascular cell fates during early vascular development. Our designed modular assemblies should be broadly useful for unraveling the complexities of signaling in key developmental transitions and for developing future therapeutic applications. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_28974.map.gz emd_28974.map.gz | 24.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-28974-v30.xml emd-28974-v30.xml emd-28974.xml emd-28974.xml | 17.1 KB 17.1 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_28974.png emd_28974.png | 76.4 KB | ||
| Filedesc metadata |  emd-28974.cif.gz emd-28974.cif.gz | 5.4 KB | ||
| Others |  emd_28974_half_map_1.map.gz emd_28974_half_map_1.map.gz emd_28974_half_map_2.map.gz emd_28974_half_map_2.map.gz | 24.8 MB 24.8 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-28974 http://ftp.pdbj.org/pub/emdb/structures/EMD-28974 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-28974 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-28974 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_28974.map.gz / Format: CCP4 / Size: 36.3 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_28974.map.gz / Format: CCP4 / Size: 36.3 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.825 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #1
| File | emd_28974_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
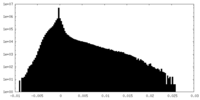
| Density Histograms |
-Half map: #2
| File | emd_28974_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
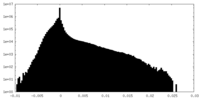
| Density Histograms |
- Sample components
Sample components
-Entire : Self-assembled homo-tetramer of de novo designed protein C4-71
| Entire | Name: Self-assembled homo-tetramer of de novo designed protein C4-71 |
|---|---|
| Components |
|
-Supramolecule #1: Self-assembled homo-tetramer of de novo designed protein C4-71
| Supramolecule | Name: Self-assembled homo-tetramer of de novo designed protein C4-71 type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism: synthetic construct (others) |
-Macromolecule #1: C4-71
| Macromolecule | Name: C4-71 / type: protein_or_peptide / ID: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism: synthetic construct (others) |
| Recombinant expression | Organism:  |
| Sequence | String: MGPEEILERA RESLERAREA SERGDEEEFR KAAEKALELA KRLVEQAKKE GDPWMVMWAA LVALWVALLA LRNGDKEVFK KAAESALEVA KRLVEVASKE GDPEMVLLAA WVALFVAWLA WLFGDKEVFK KAAESALEVA KRLVEVASKE GDPELVEEAA KVAEEVEKLA ...String: MGPEEILERA RESLERAREA SERGDEEEFR KAAEKALELA KRLVEQAKKE GDPWMVMWAA LVALWVALLA LRNGDKEVFK KAAESALEVA KRLVEVASKE GDPEMVLLAA WVALFVAWLA WLFGDKEVFK KAAESALEVA KRLVEVASKE GDPELVEEAA KVAEEVEKLA EKQGDEEVRE KAWETWMEVW LLWLEVRLRK GGGGSLEHHH HHH |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 1.0 mg/mL |
|---|---|
| Buffer | pH: 8 |
| Grid | Model: Quantifoil R2/2 / Material: COPPER / Mesh: 300 / Support film - Material: CARBON / Support film - topology: HOLEY ARRAY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 5 sec. |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 295 K / Instrument: FEI VITROBOT MARK IV / Details: blot time = 4s; blot force = 0. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Number grids imaged: 1 / Number real images: 5221 / Average exposure time: 1.6 sec. / Average electron dose: 46.77 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 1.3 µm / Nominal defocus min: 0.3 µm / Nominal magnification: 105000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: NONE / Details: Ab initio model generated in Cryosparc |
|---|---|
| Final reconstruction | Applied symmetry - Point group: C4 (4 fold cyclic) / Resolution.type: BY AUTHOR / Resolution: 5.1 Å / Resolution method: FSC 0.143 CUT-OFF / Software - Name: RELION (ver. 2) / Details: RELION v.2 3D auto-refine / Number images used: 107483 |
| Initial angle assignment | Type: MAXIMUM LIKELIHOOD / Software - Name: cryoSPARC |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD / Software - Name: RELION (ver. 2) |
 Movie
Movie Controller
Controller

















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)