-検索条件
-検索結果
検索 (著者・登録者: hagen & c)の結果235件中、1から50件目までを表示しています
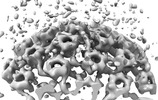
EMDB-18482: 
Herpes simplex virus 1 capsid (WT) vertices in perinuclear NEC-coated vesicles determined in situ

EMDB-18484: 
Herpes simplex virus 1 nuclear egress complex (WT) determined in situ from perinuclear vesicles

EMDB-17974: 
Pseudorabies virus cytosolic C-capsid (US3 KO) vertices determined in situ

EMDB-17975: 
Pseudorabies virus primary enveloped (perinuclear) C-capsid (US3 KO) vertices determined in situ

EMDB-17976: 
Pseudorabies nuclear C-capsids (US3 KO) vertices determined in situ

EMDB-18473: 
Subtomogram average of pseudorabies virus nuclear egress complex helical form (UL31/34) determined in situ

EMDB-18474: 
Subtomogram average of pseudorabies virus nuclear egress complex (UL31/34) determined in situ
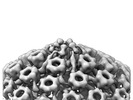

EMDB-18479: 
Pseudorabies virus cytosolic C-capsid (WT) vertices determined in situ

EMDB-18480: 
Pseudorabies virus nuclear C-capsid (WT) vertices determined in situ
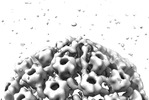

EMDB-18481: 
Herpes simplex virus 1 cytosolic C-capsid (WT) vertices determined in situ
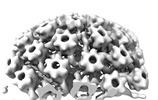
EMDB-18483: 
Herpes simplex virus 1 nuclear C-capsid (WT) vertices determined in situ

EMDB-18212: 
Cryo-EM structure of Adenovirus C5 hexon

PDB-8q7c: 
Cryo-EM structure of Adenovirus C5 hexon

EMDB-14707: 
Trypanosoma brucei gambiense ISG65 in complex with human complement component C3

EMDB-14708: 
Trypanosoma brucei gambiense ISG65 in complex with human complement component C3b

PDB-7zgj: 
Trypanosoma brucei gambiense ISG65 in complex with human complement component C3

PDB-7zgk: 
Trypanosoma brucei gambiense ISG65 in complex with human complement component C3b

EMDB-16454: 
Electron cryo-tomography of HeLa cells expressing untagged Cidec
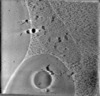
EMDB-16455: 
Electron cryo-tomography of HeLa cells expressing Cidec-EGFP

EMDB-14325: 
Cytoplasmic ring of the human nuclear pore complex from isolated HeLa nuclear envelopes

EMDB-14326: 
Structure of nuclear pore complex from HEK cells with GP210 knockout

EMDB-14327: 
Nuclear pore complex from intact HeLa cells

EMDB-14328: 
Inner/spoke ring of the human nuclear pore complex from isolated HeLa nuclear envelopes.

EMDB-14330: 
Nuclear ring of the human nuclear pore complex from isolated HeLa nuclear envelopes

EMDB-14321: 
Human nuclear pore complex (dilated)

PDB-7r5j: 
Human nuclear pore complex (dilated)

EMDB-14322: 
Human nuclear pore complex (constricted)

PDB-7r5k: 
Human nuclear pore complex (constricted)

EMDB-14243: 
cryoEM structure of human Nup155 (residues 19-981)

PDB-7r1y: 
cryoEM structure of human Nup155 (residues 19-981)

EMDB-14032: 
Structure of the EIAV CA-SP2 hexamer (C2), acquired on Krios G4, Selectris/Falcon4, processed with Relion/M

EMDB-14034: 
Structure of the EIAV CA-SP hexamer (C2), acquired on Krios 3Gi/Gatan Bioquantum K3, processed with Relion/M

EMDB-14033: 
Structure of the EIAV CA-SP hexamer (C6), acquired on Krios 3Gi/Bioquantum K3, processed with Relion/M

EMDB-14035: 
Structure of the EIAV CA-SP hexamer (C6), acquired on Krios4, Selectris/Falcon4, processed with AV3

EMDB-14036: 
Cryo-electron tomogram of EIAV CASPNC VLPs, acquired on Krios G4, Selectris/Falcon4

EMDB-14037: 
Cryo-electron tomogram of EIAV CASPNC VLPs, acquired on Krios 3Gi, Bioquantum K3

EMDB-14031: 
Structure of the EIAV CA-SP hexamer (C6), acquired on a Krios 4, Selectris/Falcon4, processed with Relion/M

EMDB-23542: 
Structure of full-length GluK1 with L-Glu

PDB-7lvt: 
Structure of full-length GluK1 with L-Glu

EMDB-11222: 
Structure of SARS-CoV-2 spike glycoprotein (S) trimer determined by sub-tomogram averaging

EMDB-11223: 
Structure of SARS-CoV-2 spike glycoprotein (S) monomer in a closed conformation determined by sub-tomogram averaging

EMDB-11347: 
Structure of SARS-CoV-2 spike glycoprotein (S) trimer with one receptor binding domain (RBD) in open-state determined by subtomogram averaging

EMDB-10660: 
Nup116_delta_NPC_25C

EMDB-10661: 
Nup116delta_NPC_37C

EMDB-10198: 
In-cell S cerevisiae nuclear pore complex

EMDB-22190: 
Subtomogram averaging map of yeast polysome

EMDB-22196: 
Cryo-EM Structure of K63R Ubiquitin Mutant Ribosome under Oxidative Stress

EMDB-22197: 
Cryo-EM Structure of K63R Ubiquitin Mutant Consensus Ribosome under Oxidative Stress

EMDB-22198: 
Cryo-EM Structure of K63 Ubiquitinated Yeast Translocating Ribosome under Oxidative Stress

PDB-6xiq: 
Cryo-EM Structure of K63R Ubiquitin Mutant Ribosome under Oxidative Stress
ページ:
 ムービー
ムービー コントローラー
コントローラー 構造ビューア
構造ビューア EMN検索について
EMN検索について



 wwPDBはEMDBデータモデルのバージョン3へ移行します
wwPDBはEMDBデータモデルのバージョン3へ移行します
