+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Human nuclear pore complex (constricted) | |||||||||
 Map data Map data | Structure of the human nuclear pore complex from isolated nuclear envelopes | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | multiprotein complex / nucleocytoplasmic transport / TRANSPORT PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationcytoplasmic side of nuclear pore / nuclear pore transmembrane ring / nephron development / centriole assembly / positive regulation of centriole replication / GATOR2 complex / cytoplasmic periphery of the nuclear pore complex / regulation of protein import into nucleus / regulation of Ras protein signal transduction / positive regulation of mitotic cytokinetic process ...cytoplasmic side of nuclear pore / nuclear pore transmembrane ring / nephron development / centriole assembly / positive regulation of centriole replication / GATOR2 complex / cytoplasmic periphery of the nuclear pore complex / regulation of protein import into nucleus / regulation of Ras protein signal transduction / positive regulation of mitotic cytokinetic process / SUMO ligase complex / HuR (ELAVL1) binds and stabilizes mRNA / Seh1-associated complex / SUMO ligase activity / nuclear pore inner ring / COPII-coated vesicle budding / nuclear pore localization / annulate lamellae / protein exit from endoplasmic reticulum / nuclear pore central transport channel / transcription-dependent tethering of RNA polymerase II gene DNA at nuclear periphery / regulation of nucleocytoplasmic transport / telomere tethering at nuclear periphery / nuclear pore complex assembly / nuclear pore outer ring / protein localization to nuclear inner membrane / COPII-coated vesicle cargo loading / nuclear pore organization / somite development / nuclear envelope organization / nuclear pore cytoplasmic filaments / atrial cardiac muscle cell action potential / homologous chromosome pairing at meiosis / positive regulation of protein localization to centrosome / COPII vesicle coat / nuclear export signal receptor activity / paraxial mesoderm development / Nuclear Pore Complex (NPC) Disassembly / post-transcriptional tethering of RNA polymerase II gene DNA at nuclear periphery / nuclear inclusion body / Regulation of Glucokinase by Glucokinase Regulatory Protein / Defective TPR may confer susceptibility towards thyroid papillary carcinoma (TPC) / nuclear pore nuclear basket / Transport of Ribonucleoproteins into the Host Nucleus / attachment of mitotic spindle microtubules to kinetochore / miRNA processing / Transport of the SLBP independent Mature mRNA / Transport of the SLBP Dependant Mature mRNA / Amino acids regulate mTORC1 / NS1 Mediated Effects on Host Pathways / SUMOylation of SUMOylation proteins / negative regulation of Ras protein signal transduction / structural constituent of nuclear pore / positive regulation of mRNA splicing, via spliceosome / microtubule bundle formation / Transport of Mature mRNA Derived from an Intronless Transcript / Flemming body / Rev-mediated nuclear export of HIV RNA / SUMOylation of RNA binding proteins / nuclear export / Nuclear import of Rev protein / mitotic centrosome separation / NEP/NS2 Interacts with the Cellular Export Machinery / fertilization / Transferases; Acyltransferases; Aminoacyltransferases / Transport of Mature mRNA derived from an Intron-Containing Transcript / RNA export from nucleus / SUMO transferase activity / tRNA processing in the nucleus / Postmitotic nuclear pore complex (NPC) reformation / kinase activator activity / COPII-mediated vesicle transport / neural tube development / centrosome cycle / centrosome localization / protein-containing complex localization / nuclear localization sequence binding / regulation of gluconeogenesis / lamellipodium assembly / Viral Messenger RNA Synthesis / nucleocytoplasmic transport / negative regulation of programmed cell death / NLS-bearing protein import into nucleus / PTB domain binding / poly(A)+ mRNA export from nucleus / mitotic metaphase chromosome alignment / SUMOylation of ubiquitinylation proteins / Vpr-mediated nuclear import of PICs / female gonad development / macrophage chemotaxis / SUMOylation of DNA replication proteins / Hydrolases; Acting on peptide bonds (peptidases); Serine endopeptidases / positive regulation of SMAD protein signal transduction / negative regulation of epidermal growth factor receptor signaling pathway / protein sumoylation / RHOA GTPase cycle / Regulation of HSF1-mediated heat shock response / mitotic spindle assembly / regulation of signal transduction / positive regulation of TOR signaling Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | subtomogram averaging / cryo EM / Resolution: 12.0 Å | |||||||||
 Authors Authors | Mosalaganti S / Obarska-Kosinska A | |||||||||
| Funding support | European Union,  Germany, 2 items Germany, 2 items
| |||||||||
 Citation Citation |  Journal: Science / Year: 2022 Journal: Science / Year: 2022Title: AI-based structure prediction empowers integrative structural analysis of human nuclear pores. Authors: Shyamal Mosalaganti / Agnieszka Obarska-Kosinska / Marc Siggel / Reiya Taniguchi / Beata Turoňová / Christian E Zimmerli / Katarzyna Buczak / Florian H Schmidt / Erica Margiotta / Marie- ...Authors: Shyamal Mosalaganti / Agnieszka Obarska-Kosinska / Marc Siggel / Reiya Taniguchi / Beata Turoňová / Christian E Zimmerli / Katarzyna Buczak / Florian H Schmidt / Erica Margiotta / Marie-Therese Mackmull / Wim J H Hagen / Gerhard Hummer / Jan Kosinski / Martin Beck /   Abstract: INTRODUCTION The eukaryotic nucleus pro-tects the genome and is enclosed by the two membranes of the nuclear envelope. Nuclear pore complexes (NPCs) perforate the nuclear envelope to facilitate ...INTRODUCTION The eukaryotic nucleus pro-tects the genome and is enclosed by the two membranes of the nuclear envelope. Nuclear pore complexes (NPCs) perforate the nuclear envelope to facilitate nucleocytoplasmic transport. With a molecular weight of ∼120 MDa, the human NPC is one of the larg-est protein complexes. Its ~1000 proteins are taken in multiple copies from a set of about 30 distinct nucleoporins (NUPs). They can be roughly categorized into two classes. Scaf-fold NUPs contain folded domains and form a cylindrical scaffold architecture around a central channel. Intrinsically disordered NUPs line the scaffold and extend into the central channel, where they interact with cargo complexes. The NPC architecture is highly dynamic. It responds to changes in nuclear envelope tension with conforma-tional breathing that manifests in dilation and constriction movements. Elucidating the scaffold architecture, ultimately at atomic resolution, will be important for gaining a more precise understanding of NPC function and dynamics but imposes a substantial chal-lenge for structural biologists. RATIONALE Considerable progress has been made toward this goal by a joint effort in the field. A synergistic combination of complementary approaches has turned out to be critical. In situ structural biology techniques were used to reveal the overall layout of the NPC scaffold that defines the spatial reference for molecular modeling. High-resolution structures of many NUPs were determined in vitro. Proteomic analysis and extensive biochemical work unraveled the interaction network of NUPs. Integra-tive modeling has been used to combine the different types of data, resulting in a rough outline of the NPC scaffold. Previous struc-tural models of the human NPC, however, were patchy and limited in accuracy owing to several challenges: (i) Many of the high-resolution structures of individual NUPs have been solved from distantly related species and, consequently, do not comprehensively cover their human counterparts. (ii) The scaf-fold is interconnected by a set of intrinsically disordered linker NUPs that are not straight-forwardly accessible to common structural biology techniques. (iii) The NPC scaffold intimately embraces the fused inner and outer nuclear membranes in a distinctive topol-ogy and cannot be studied in isolation. (iv) The conformational dynamics of scaffold NUPs limits the resolution achievable in structure determination. RESULTS In this study, we used artificial intelligence (AI)-based prediction to generate an exten-sive repertoire of structural models of human NUPs and their subcomplexes. The resulting models cover various domains and interfaces that so far remained structurally uncharac-terized. Benchmarking against previous and unpublished x-ray and cryo-electron micros-copy structures revealed unprecedented accu-racy. We obtained well-resolved cryo-electron tomographic maps of both the constricted and dilated conformational states of the hu-man NPC. Using integrative modeling, we fit-ted the structural models of individual NUPs into the cryo-electron microscopy maps. We explicitly included several linker NUPs and traced their trajectory through the NPC scaf-fold. We elucidated in great detail how mem-brane-associated and transmembrane NUPs are distributed across the fusion topology of both nuclear membranes. The resulting architectural model increases the structural coverage of the human NPC scaffold by about twofold. We extensively validated our model against both earlier and new experimental data. The completeness of our model has enabled microsecond-long coarse-grained molecular dynamics simulations of the NPC scaffold within an explicit membrane en-vironment and solvent. These simulations reveal that the NPC scaffold prevents the constriction of the otherwise stable double-membrane fusion pore to small diameters in the absence of membrane tension. CONCLUSION Our 70-MDa atomically re-solved model covers >90% of the human NPC scaffold. It captures conforma-tional changes that occur during dilation and constriction. It also reveals the precise anchoring sites for intrinsically disordered NUPs, the identification of which is a prerequisite for a complete and dy-namic model of the NPC. Our study exempli-fies how AI-based structure prediction may accelerate the elucidation of subcellular ar-chitecture at atomic resolution. [Figure: see text]. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_14322.map.gz emd_14322.map.gz | 306.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-14322-v30.xml emd-14322-v30.xml emd-14322.xml emd-14322.xml | 56.8 KB 56.8 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_14322.png emd_14322.png | 40 KB | ||
| Filedesc metadata |  emd-14322.cif.gz emd-14322.cif.gz | 22.7 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-14322 http://ftp.pdbj.org/pub/emdb/structures/EMD-14322 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-14322 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-14322 | HTTPS FTP |
-Related structure data
| Related structure data |  7r5kMC  7tbjM  7tblM  7r1yC  7r5jC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_14322.map.gz / Format: CCP4 / Size: 729 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_14322.map.gz / Format: CCP4 / Size: 729 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Structure of the human nuclear pore complex from isolated nuclear envelopes | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 3.37 Å | ||||||||||||||||||||||||||||||||||||
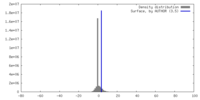
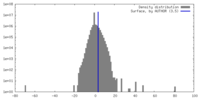
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
- Sample components
Sample components
+Entire : Nuclear pore complex
+Supramolecule #1: Nuclear pore complex
+Macromolecule #1: E3 SUMO-protein ligase RanBP2
+Macromolecule #2: Nuclear pore membrane glycoprotein 210
+Macromolecule #3: Aladin
+Macromolecule #4: Nuclear pore complex protein Nup93
+Macromolecule #5: Nucleoporin NUP188 homolog
+Macromolecule #6: Nuclear pore complex protein Nup205
+Macromolecule #7: Nuclear pore complex protein Nup155
+Macromolecule #8: Nucleoporin NDC1
+Macromolecule #9: Nucleoporin NUP35
+Macromolecule #10: Nucleoporin p54
+Macromolecule #11: Nucleoporin p58/p45
+Macromolecule #12: Nuclear pore glycoprotein p62
+Macromolecule #13: Nuclear pore complex protein Nup133
+Macromolecule #14: Nuclear pore complex protein Nup107
+Macromolecule #15: Nuclear pore complex protein Nup96
+Macromolecule #16: Protein SEC13 homolog
+Macromolecule #17: Nucleoporin SEH1
+Macromolecule #18: Nuclear pore complex protein Nup85
+Macromolecule #19: Nucleoporin Nup43
+Macromolecule #20: Nuclear pore complex protein Nup160
+Macromolecule #21: Nucleoporin Nup37
+Macromolecule #22: Protein ELYS
+Macromolecule #23: Nuclear pore complex protein Nup98
+Macromolecule #24: Nuclear pore complex protein Nup214
+Macromolecule #25: Nuclear pore complex protein Nup88
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | subtomogram averaging |
| Aggregation state | cell |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE / Instrument: LEICA EM GP |
- Electron microscopy
Electron microscopy
| Microscope | TFS KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 QUANTUM (4k x 4k) / Average electron dose: 2.93 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 70.0 µm / Illumination mode: OTHER / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 4.0 µm / Nominal defocus min: 2.0 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Final reconstruction | Applied symmetry - Point group: C1 (asymmetric) / Resolution.type: BY AUTHOR / Resolution: 12.0 Å / Resolution method: FSC 0.143 CUT-OFF / Number subtomograms used: 7711 |
|---|---|
| Extraction | Number tomograms: 554 / Number images used: 7711 |
| Final angle assignment | Type: OTHER |
 Movie
Movie Controller
Controller












































 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)