-Search query
-Search result
Showing all 40 items for (author: foe & it)

EMDB-29941: 
HPV16 E6-E6AP-p53 complex
Method: single particle / : Bratkowski MA, Wang JCK, Hao Q, Nile AH

EMDB-12749: 
In situ cryo-electron tomogram of a pyrenoid inside a Chlamydomonas reinhardtii cell
Method: electron tomography / : Cuellar LK, Schaffer M, Strauss M, Martinez-Sanchez A, Plitzko JM, Foerster F, Engel BD

EMDB-11992: 
the molecular sociology at the HeLa cell nuclear periphery
Method: electron tomography / : Mahamid J, Pfeffer S, Schaffer M, Villa E, Danev R, Kuhn-Cuellar L, Foerster F, Hyman A, Plitzko J, Baumeister W

EMDB-4306: 
Subtomogram average of unsorted ribosome-translocon complexes from STT3B(-/-) HEK cells
Method: subtomogram averaging / : Pfeffer S, Foerster F

EMDB-4307: 
Subtomogram average of OST-free ribosome-translocon complexes from STT3B(-/-) HEK cells
Method: subtomogram averaging / : Pfeffer S, Foerster F

EMDB-4308: 
Subtomogram average of OST-containing ribosome-translocon complexes from STT3B(-/-) HEK cells
Method: subtomogram averaging / : Pfeffer S, Foerster F
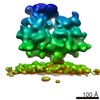
EMDB-4309: 
Subtomogram average of unsorted ribosome-translocon complexes from STT3A(-/-) HEK cells
Method: subtomogram averaging / : Pfeffer S, Foerster F
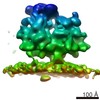
EMDB-4310: 
Subtomogram average of OST-free ribosome-translocon complexes from STT3A(-/-) HEK cells
Method: subtomogram averaging / : Pfeffer S, Foerster F

EMDB-4311: 
Subtomogram average of ribosome-translocon complexes from STT3A(-/-) HEK cells containing an unidentified ER-lumenal component
Method: subtomogram averaging / : Pfeffer S, Foerster F
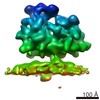
EMDB-4312: 
Subtomogram average of unsorted ribosome-translocon complexes from wild type HEK cells
Method: subtomogram averaging / : Pfeffer S, Foerster F
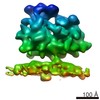
EMDB-4313: 
Subtomogram average of OST-free ribosome-translocon complexes from wild type HEK cells
Method: subtomogram averaging / : Pfeffer S, Foerster F
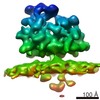
EMDB-4314: 
Subtomogram average of OST-containing ribosome-translocon complexes from wild type HEK cells
Method: subtomogram averaging / : Pfeffer S, Foerster F

EMDB-4315: 
Subtomogram average of OST-containing ribosome-translocon complexes from canine rough microsomal membranes
Method: subtomogram averaging / : Pfeffer S, Foerster F

PDB-6ftg: 
Subtomogram average of OST-containing ribosome-translocon complexes from canine rough microsomal membranes
Method: subtomogram averaging / : Pfeffer S, Foerster F

EMDB-3694: 
In situ subtomogram average of Rubisco within the Chlamydomonas pyrenoid
Method: subtomogram averaging / : Cuellar LK, Schaffer M, Strauss M, Martinez-Sanchez A, Plitzko JM, Foerster F, Engel BD

EMDB-3603: 
KaiCB circadian clock backbone model based on a Cryo-EM density
Method: single particle / : Schuller JM, Snijder J, Loessl P, Heck AJR, Foerster F

PDB-5n8y: 
KaiCBA circadian clock backbone model based on a Cryo-EM density
Method: single particle / : Schuller JM, Snijder J, Loessl P, Heck AJR, Foerster F

PDB-5mp9: 
26S proteasome in presence of ATP (s1)
Method: single particle / : Wehmer M, Rudack T, Beck F, Aufderheide A, Pfeifer G, Plitzko JM, Foerster F, Schulten K, Baumeister W, Sakata E

PDB-5mpa: 
26S proteasome in presence of ATP (s2)
Method: single particle / : Wehmer M, Rudack T, Beck F, Aufderheide A, Pfeifer G, Plitzko JM, Foerster F, Schulten K, Baumeister W, Sakata E

PDB-5mpb: 
26S proteasome in presence of AMP-PNP (s3)
Method: single particle / : Wehmer M, Rudack T, Beck F, Aufderheide A, Pfeifer G, Plitzko JM, Foerster F, Schulten K, Baumeister W, Sakata E

PDB-5mpc: 
26S proteasome in presence of BeFx (s4)
Method: single particle / : Wehmer M, Rudack T, Beck F, Aufderheide A, Pfeifer G, Plitzko JM, Foerster F, Schulten K, Baumeister W, Sakata E

PDB-5mpd: 
26S proteasome in presence of ATP (s1)
Method: single particle / : Wehmer M, Rudack T, Beck F, Aufderheide A, Pfeifer G, Plitzko JM, Foerster F, Schulten K, Baumeister W, Sakata E

PDB-5mpe: 
26S proteasome in presence of ATP (s2)
Method: single particle / : Wehmer M, Rudack T, Beck F, Aufderheide A, Pfeifer G, Plitzko JM, Foerster F, Schulten K, Baumeister W, Sakata E

EMDB-4143: 
Subtomogram average of the ER membrane-associated ribosome from human TRAPdelta-deficient fibroblasts
Method: subtomogram averaging / : Pfeffer S, Dudek J, Ng BG, Zimmermann R, Freeze HH, Foerster F

EMDB-4144: 
Subtomogram average of the ER membrane-associated ribosome from human TRAPgamma-deficient fibroblasts
Method: subtomogram averaging / : Pfeffer S, Dudek J, Ng BG, Zimmermann R, Freeze HH, Foerster F

EMDB-4145: 
Subtomogram average of the ER membrane-associated ribosome from FIB-milled C. reinhardtii cells
Method: subtomogram averaging / : Pfeffer S, Schaffer M, Albert S, Engel BD, Foerster F

EMDB-8054: 
In situ sub-tomogram average of HeLa nuclear pore complex from a single cell
Method: subtomogram averaging / : Mahamid J, Pfeffer S, Schaffer M, Villa E, Danev R, Kuhn-Cuellar L, Foerster F, Hyman A, Plitzko J, Baumeister W

EMDB-8055: 
In situ sub-tomogram average of HeLa nuclear pore complex
Method: subtomogram averaging / : Mahamid J, Pfeffer S, Schaffer M, Villa E, Danev R, Kuhn-Cuellar L, Foerster F, Hyman A, Plitzko J, Baumeister W

EMDB-8056: 
In situ sub-tomogram average of HeLa ER-associated ribosomes
Method: subtomogram averaging / : Mahamid J, Pfeffer S, Schaffer M, Villa E, Danev R, Kuhn-Cuellar L, Foerster F, Hyman A, Plitzko J, Baumeister W

EMDB-8057: 
In situ sub-tomogram average of HeLa cytosolic ribosomes
Method: subtomogram averaging / : Mahamid J, Pfeffer S, Schaffer M, Villa E, Danev R, Kuhn-Cuellar L, Foerster F, Hyman A, Plitzko J, Baumeister W

EMDB-3231: 
Structure and function based design of Plasmodium-selective proteasome inhibitors
Method: single particle / : Li H, O' Donoghue AJ, van der Linden WA, Xie SC, Yoo E, Foe IT, Tilley L, Craik CS, da Fonseca PCA, Bogyo M

PDB-5a5b: 
Structure of the 26S proteasome-Ubp6 complex
Method: single particle / : Aufderheide A, Beck F, Stengel F, Hartwig M, Schweitzer A, Pfeifer G, Goldberg AL, Sakata E, Baumeister W, Foerster F

EMDB-3034: 
Structure of the 26S proteasome-Ubp6 complex
Method: single particle / : Aufderheide A, Beck F, Stengel F, Hartwig M, Schweitzer A, Pfeifer G, Goldberg AL, Sakata E, Baumeister W, Foerster F

EMDB-2594: 
Deep classification of a large cryo-EM dataset defines the conformational landscape of the 26S proteasome
Method: single particle / : Unverdorben P, Beck F, Sledz P, Schweitzer A, Pfeifer G, Plitzko JM, Baumeister W, Foerster F

EMDB-2595: 
Deep classification of a large cryo-EM dataset defines the conformational landscape of the 26S proteasome
Method: single particle / : Unverdorben P, Beck F, Sledz P, Schweitzer A, Pfeifer G, Plitzko JM, Baumeister W, Foerster F

EMDB-2596: 
Deep classification of a large cryo-EM dataset defines the conformational landscape of the 26S proteasome
Method: single particle / : Unverdorben P, Beck F, Sledz P, Schweitzer A, Pfeifer G, Plitzko JM, Baumeister W, Foerster F

PDB-4cr2: 
Deep classification of a large cryo-EM dataset defines the conformational landscape of the 26S proteasome
Method: single particle / : Unverdorben P, Beck F, Sledz P, Schweitzer A, Pfeifer G, Plitzko JM, Baumeister W, Foerster F

PDB-4cr3: 
Deep classification of a large cryo-EM dataset defines the conformational landscape of the 26S proteasome
Method: single particle / : Unverdorben P, Beck F, Sledz P, Schweitzer A, Pfeifer G, Plitzko JM, Baumeister W, Foerster F

PDB-4cr4: 
Deep classification of a large cryo-EM dataset defines the conformational landscape of the 26S proteasome
Method: single particle / : Unverdorben P, Beck F, Sledz P, Schweitzer A, Pfeifer G, Plitzko JM, Baumeister W, Foerster F

EMDB-2348: 
Structure of the 26S proteasome with ATP-yS bound provides insights into the mechanism of nucleotide-dependent substrate translocation
Method: single particle / : Sledz P, Unverdorben P, Beck F, Pfeifer G, Schweitzer A, Foerster F, Baumeister W
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
