-Search query
-Search result
Showing all 49 items for (author: doyle & la)

EMDB-74435: 
Dimer of BrxC-BrxB fusion complexed with PglZ from the Acinetobacter BREX system
Method: single particle / : Doyle LA, Stoddard BL, Kaiser B, Kaiser A

EMDB-74076: 
Dimer of ATPase BrxC containing a Walker B mutation and bound to ATP from the Acinetobacter BREX system
Method: single particle / : Doyle LA, Stoddard BL, Kaiser B, Kaiser A

EMDB-74400: 
Volume of PglZ in complex with BrxB-BrxC fusion from the Acinetobacter BREX system
Method: single particle / : Doyle LA, Stoddard BL, Kaiser B, Kaiser A

EMDB-44474: 
HIV-1 Env 16055 dGly4 NFL
Method: single particle / : Ozorowski G, Lee WH, Ward AB

PDB-9be9: 
HIV-1 Env 16055 dGly4 NFL
Method: single particle / : Ozorowski G, Lee WH, Ward AB

EMDB-51308: 
Cryo-EM Structure of Pentameric Outer Membrane Protein A from Bdellovibrio bacteriovorus
Method: single particle / : Parr RJ, Ratkeviciute G, Lovering AL

PDB-9gf0: 
Cryo-EM Structure of Pentameric Outer Membrane Protein A from Bdellovibrio bacteriovorus
Method: single particle / : Parr RJ, Ratkeviciute G, Lovering AL

EMDB-49827: 
Complex of BREX proteins BrxB and PglZ from Salmonella typhimurium
Method: single particle / : Doyle LA, Stoddard B, Blower TR, Kaiser B

EMDB-46047: 
HIV Env JRFL NFL TD CC3+ trimer in complex with NHP #1 polyclonal Fab (gp120 interface, gp41-FP, gp41-base epitopes)
Method: single particle / : Sewall LM, Ozorowski G, Ward AB

EMDB-46048: 
HIV Env ZM233 NFL TD CC3+ trimer in complex with NHP #1,2,3,4 polyclonal Fab (gp41-FP, gp41-base epitopes)
Method: single particle / : Sewall LM, Ozorowski G, Ward AB

EMDB-45928: 
LJF-085 Fab in complex with HIV Env JRFL NFL TD CC3+ trimer and 35O22 Fab
Method: single particle / : Ozorowski G, Ward AB

EMDB-45929: 
LJF-034 Fab in complex with HIV Env JRFL NFL TD CC3+ trimer and 35O22 Fab
Method: single particle / : Ozorowski G, Ward AB

EMDB-45957: 
LJF-085 Fab in complex with HIV Env ZM233 NFL TD CC3+ trimer
Method: single particle / : Ozorowski G, Ward AB

EMDB-45978: 
LJF-085 Fab in complex with HIV Env ZM233 NFL TD CC3+ dimer and 35O22 Fab
Method: single particle / : Ozorowski G, Ward AB

PDB-9cu5: 
LJF-085 Fab in complex with HIV Env JRFL NFL TD CC3+ trimer and 35O22 Fab
Method: single particle / : Ozorowski G, Ward AB

PDB-9cu6: 
LJF-034 Fab in complex with HIV Env JRFL NFL TD CC3+ trimer and 35O22 Fab
Method: single particle / : Ozorowski G, Ward AB

PDB-9cv7: 
LJF-085 Fab in complex with HIV Env ZM233 NFL TD CC3+ trimer
Method: single particle / : Ozorowski G, Ward AB

EMDB-43551: 
CCHFV GP38 bound with ADI-46143 and ADI-46158 Fabs
Method: single particle / : Hjorth CK, McLellan JS

EMDB-43552: 
CCHFV GP38 bound with ADI-58062 and ADI-63530 Fabs
Method: single particle / : Hjorth CK, McLellan JS

EMDB-43553: 
CCHFV GP38 bound with ADI-58026 and ADI-63547 Fabs
Method: single particle / : Hjorth CK, McLellan JS

EMDB-43604: 
CCHFV GP38 bound to ADI-46152 and ADI-58048 Fabs
Method: single particle / : Hjorth CK, McLellan JS

PDB-8vww: 
CCHFV GP38 bound to ADI-46152 and ADI-58048 Fabs
Method: single particle / : Hjorth CK, McLellan JS

EMDB-28244: 
CryoEM characterization of BrxL -- a unique AAA+ phage restriction Factor.
Method: single particle / : Shen BW, Stoddard BL

EMDB-28248: 
CryoEM characterization of a unique AAA+ BrxL phage restriction factor
Method: single particle / : Shen BW, Stoddard BL

EMDB-27098: 
Cryo-EM structure of the SARS-CoV-2 HR1HR2 fusion core complex with extended HR2
Method: single particle / : Yang K, Brunger AT

PDB-8czi: 
Cryo-EM structure of the SARS-CoV-2 HR1HR2 fusion core complex with extended HR2
Method: single particle / : Yang K, Brunger AT

EMDB-26105: 
BamABCDE bound to substrate EspP class 1
Method: single particle / : Doyle MT, Jimah JR

EMDB-26106: 
BamABCDE bound to substrate EspP class 2
Method: single particle / : Doyle MT, Jimah JR

EMDB-26107: 
BamABCDE bound to substrate EspP class 4
Method: single particle / : Doyle MT, Jimah JR

EMDB-26108: 
BamABCDE bound to substrate EspP class 3
Method: single particle / : Doyle MT, Jimah JR

EMDB-26109: 
BamABCDE bound to substrate EspP class 5
Method: single particle / : Doyle MT, Jimah JR

EMDB-26110: 
BamABCDE bound to substrate EspP class 6
Method: single particle / : Doyle MT, Jimah JR

EMDB-26111: 
BamABCDE bound to substrate EspP in the open-sheet EspP state
Method: single particle / : Doyle MT, Jimah JR

EMDB-26112: 
BamABCDE bound to substrate EspP in the intermediate-open EspP state
Method: single particle / : Doyle MT, Jimah JR

EMDB-26113: 
BamABCDE bound to substrate EspP in the barrelized EspP/continuous open BamA state
Method: single particle / : Doyle MT, Jimah JR

EMDB-26114: 
BamABCDE bound to substrate EspP
Method: single particle / : Doyle MT, Jimah JR

PDB-7tsz: 
BamABCDE bound to substrate EspP class 1
Method: single particle / : Doyle MT, Jimah JR, Dowdy T, Ohlemacher SI, Larion M, Hinshaw JE, Bernstein HD

PDB-7tt0: 
BamABCDE bound to substrate EspP class 2
Method: single particle / : Doyle MT, Jimah JR, Dowdy T, Ohlemacher SI, Larion M, Hinshaw JE, Bernstein HD

PDB-7tt1: 
BamABCDE bound to substrate EspP class 4
Method: single particle / : Doyle MT, Jimah JR, Dowdy T, Ohlemacher SI, Larion M, Hinshaw JE, Bernstein HD

PDB-7tt2: 
BamABCDE bound to substrate EspP class 3
Method: single particle / : Doyle MT, Jimah JR, Dowdy T, Ohlemacher SI, Larion M, Hinshaw JE, Bernstein HD

PDB-7tt3: 
BamABCDE bound to substrate EspP class 5
Method: single particle / : Doyle MT, Jimah JR, Dowdy T, Ohlemacher SI, Larion M, Hinshaw JE, Bernstein HD

PDB-7tt4: 
BamABCDE bound to substrate EspP class 6
Method: single particle / : Doyle MT, Jimah JR, Dowdy T, Ohlemacher SI, Larion M, Hinshaw JE, Bernstein HD

PDB-7tt5: 
BamABCDE bound to substrate EspP in the open-sheet EspP state
Method: single particle / : Doyle MT, Jimah JR, Dowdy T, Ohlemacher SI, Larion M, Hinshaw JE, Bernstein HD

PDB-7tt6: 
BamABCDE bound to substrate EspP in the intermediate-open EspP state
Method: single particle / : Doyle MT, Jimah JR, Dowdy T, Ohlemacher SI, Larion M, Hinshaw JE, Bernstein HD

PDB-7tt7: 
BamABCDE bound to substrate EspP in the barrelized EspP/continuous open BamA state
Method: single particle / : Doyle MT, Jimah JR, Dowdy T, Ohlemacher SI, Larion M, Hinshaw JE, Bernstein HD

PDB-7ttc: 
BamABCDE bound to substrate EspP
Method: single particle / : Doyle MT, Jimah JR, Dowdy T, Ohlemacher SI, Larion M, Hinshaw JE, Bernstein HD

EMDB-23488: 
Hendra virus receptor binding protein in complex with HENV-103 and HENV-117 Fabs
Method: single particle / : Binshtein E, Crowe JE
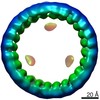
EMDB-24425: 
Circular tandem repeat protein with novel repeat topology and enhanced subunit contact surfaces
Method: single particle / : Shen BW, Stoddard BL

PDB-7rdr: 
Circular tandem repeat protein with novel repeat topology and enhanced subunit contact surfaces
Method: single particle / : Shen BW, Stoddard BL
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
