-検索条件
-検索結果
検索 (著者・登録者: chen & yh)の結果114件中、1から50件目までを表示しています

EMDB-39098: 
Cryo-electron microscopic structure of an amide hydrolase from Pseudoxanthomonas wuyuanensis

EMDB-39454: 
structure of phage T6 full-length topoisomerase II bound with DNA

EMDB-39433: 
structure of phage T6 topoisomerase II ATPase domain bound with AMPPNP

EMDB-39436: 
structure of phage T6 topoisomerase II central domain

EMDB-39438: 
structure of phage T4 topoisomerase II gp52 subunit WHD-open state

EMDB-39437: 
structure of phage T6 topoisomerase II central domain bound with DNA and m-AMSA

EMDB-39391: 
structure of phage T6 topoisomerase II central domain bound with DNA

EMDB-39434: 
structure of phage T4 topoisomerase II central domain

EMDB-39435: 
structure of phage T4 topoisomerase II central domain bound with DNA

EMDB-39444: 
structure of phage T6 apo full-length topoisomerase II

EMDB-61053: 
cryo-EM structure of human cystic fibrosis transmembrane conductance regulator (CFTR) from Biortus

EMDB-41849: 
Structure of 310-18A5 Fab in complex with A/Solomon Islands/3/2006(H1N1) influenza virus hemagglutinin

EMDB-43011: 
Phosphorylated, ATP-bound, E1371Q human cystic fibrosis transmembrane conductance regulator (E1371Q-CFTR)

EMDB-43014: 
Phosphorylated, ATP-bound, inhibitor 172-bound E1371Q human cystic fibrosis transmembrane conductance regulator

PDB-8v7z: 
Phosphorylated, ATP-bound, E1371Q human cystic fibrosis transmembrane conductance regulator (E1371Q-CFTR)

PDB-8v81: 
Phosphorylated, ATP-bound, inhibitor 172-bound E1371Q human cystic fibrosis transmembrane conductance regulator

EMDB-38216: 
Cryo-EM structure of SARS-CoV-2 S-BQ.1 in complex with antibody O5C2

EMDB-17362: 
Homotypic interacting B1 fab bound to Chondroitin Sulfate A

EMDB-37604: 
Structural basis of translation inhibition by a valine tRNA-derived fragment

EMDB-37733: 
Structural basis of translation inhibition by a valine tRNA-derived fragment

EMDB-37734: 
Structural basis of translation inhibition by a valine tRNA-derived fragment
EMDB-39012: 
Representative tomogram of primary glioblastoma stem cell with circular inter-mitochondrial junctions.
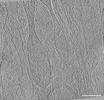

EMDB-39015: 
Representative tomogram of microglia cell with nanotunnel-like structures resembling mitochondrial fission.
EMDB-39019: 
Representative tomogram of glioblastoma cell with nanotunnel-like structure and inter-mitochondrial junction.

EMDB-39021: 
Representative tomogram of normal human astrocyte with nanotunnel-like structure which is an extension of the mitochondrial outer membrane.
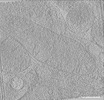

EMDB-39023: 
Representative tomogram of primary glioblastoma differentiated cell with parallel inter-mitochondrial junction.
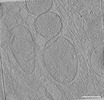

EMDB-39024: 
Representative tomogram of primary glioblastoma stem cell with clustered mitochondria bearing various long-short axis ratios.

EMDB-34866: 
Cryo-EM Structures and Translocation Mechanism of Crenarchaeota Ribosome

EMDB-34869: 
Cryo-EM Structures and Translocation Mechanism of Crenarchaeota Ribosome

EMDB-35775: 
The rice Na+/H+ antiporter SOS1 in an auto-inhibited state

EMDB-35950: 
The truncated rice Na+/H+ antiporter SOS1 (1-976) in a constitutively active state

EMDB-34870: 
Cryo-EM Structures and Translocation Mechanism of Crenarchaeota Ribosome

EMDB-34867: 
Cryo-EM Structures and Translocation Mechanism of Crenarchaeota Ribosome

EMDB-34868: 
Cryo-EM Structures and Translocation Mechanism of Crenarchaeota Ribosome

EMDB-34863: 
Cryo-EM Structures and Translocation Mechanism of Crenarchaeota Ribosome

EMDB-34864: 
Cryo-EM Structures and Translocation Mechanism of Crenarchaeota Ribosome

EMDB-34860: 
Cryo-EM Structures and Translocation Mechanism of Crenarchaeota Ribosome

EMDB-34861: 
Cryo-EM Structures and Translocation Mechanism of Crenarchaeota Ribosome

EMDB-34862: 
Cryo-EM Structures and Translocation Mechanism of Crenarchaeota Ribosome

EMDB-34710: 
Cryo-EM structure of WeiTsing

EMDB-33145: 
Cryo-EM structures of human mitochondrial NAD(P)+-dependent malic enzyme in apo form

EMDB-33146: 
Cryo-EM structures of human mitochondrial NAD(P)+-dependent malic enzyme in a ternary complex with NAD+ and allosteric inhibitor EA

EMDB-33147: 
Cryo-EM structures of human mitochondrial NAD(P)+-dependent malic enzyme in a ternary complex with NAD+ and allosteric inhibitor MDSA

EMDB-35522: 
Cryo-EM structure of the TUG891 bound GPR120-Giq complex(mask on receptor)

EMDB-35523: 
Cryo-EM structure of the TUG891 bound GPR120-Giq complex(mask on Giq-scFV16 complex)

EMDB-35524: 
Cryo-EM structure of the eicosapentaenoic acid bound GPR120-Gi1 complex(mask on receptor)

EMDB-35525: 
Cryo-EM structure of the eicosapentaenoic acid bound GPR120-Gi1 complex(mask on Gil-scFV16 complex)

EMDB-35529: 
Cryo-EM structure of the TUG891 bound GPR120-Giq complex (consensus map)

EMDB-35533: 
Cryo-EM structure of the eicosapentaenoic acid bound GPR120-Gi complex(consensus map)

EMDB-35356: 
Cryo-EM structure of the 9-hydroxystearic acid bound GPR120-Gi complex
ページ:
 ムービー
ムービー コントローラー
コントローラー 構造ビューア
構造ビューア EMN検索について
EMN検索について



 wwPDBはEMDBデータモデルのバージョン3へ移行します
wwPDBはEMDBデータモデルのバージョン3へ移行します
