-Search query
-Search result
Showing all 21 items for (author: chakrabarti & s)

EMDB-53417: 
Human UPF1 in complex with the histone stem loop RNA
Method: single particle / : Machado de Amorim A, Loll B, Hilal T, Chakrabarti S

PDB-9qwn: 
Human UPF1 in complex with the histone stem loop RNA
Method: single particle / : Machado de Amorim A, Loll B, Hilal T, Chakrabarti S

EMDB-43639: 
Cryo-EM structure of mammalian Thorase filament, 3-layer refinement
Method: single particle / : Dar MA, Louder RK, Umanah GK, Dawson TM, Dawson VL

EMDB-43640: 
Cryo-EM structure of mammalian Thorase filament, global helical refinement
Method: helical / : Dar MA, Louder RK, Umanah GK, Dawson TM, Dawson VL

PDB-8vxt: 
Cryo-EM structure of mammalian Thorase filament, 3-layer refinement
Method: single particle / : Dar MA, Louder RK, Umanah GK, Dawson TM, Dawson VL

EMDB-41816: 
Cryo-EM structure of the RAF1-HSP90-CDC37 complex in the closed state
Method: single particle / : Finci LI, Simanshu DK

EMDB-41817: 
Cryo-EM structure of the HSP90 dimer (NTD-MD) in the semi-open state
Method: single particle / : Finci LI, Simanshu DK

EMDB-41818: 
Cryo-EM structure of the cross-linked HSP90 dimer (NTD-MD) in the semi-open state
Method: single particle / : Finci LI, Simanshu DK

PDB-8u1l: 
Cryo-EM structure of the RAF1-HSP90-CDC37 complex in the closed state
Method: single particle / : Finci LI, Simanshu DK

PDB-8u1m: 
Cryo-EM structure of the HSP90 dimer (NTD-MD) in the semi-open state
Method: single particle / : Finci LI, Simanshu DK

PDB-8u1n: 
Cryo-EM structure of the cross-linked HSP90 dimer (NTD-MD) in the semi-open state
Method: single particle / : Finci LI, Simanshu DK

EMDB-25427: 
Structure of KRAS G12V/HLA-A*03:01 in complex with antibody fragment V2
Method: single particle / : Wright KM, Gabelli SB

PDB-7stf: 
Structure of KRAS G12V/HLA-A*03:01 in complex with antibody fragment V2
Method: single particle / : Wright KM, Gabelli SB, Miller M

EMDB-32497: 
SARS-CoV-2 spike in complex with the ZB8 neutralizing antibody Fab (focused refinement on Fab-RBD)
Method: single particle / : Zeng JW

EMDB-32498: 
SARS-CoV-2 spike in complex with the ZB8 neutralizing antibody Fab (3U)
Method: single particle / : Zeng JW, Ge JW

EMDB-32499: 
SARS-CoV-2 spike in complex with the ZB8 neutralizing antibody Fab (2u1d)
Method: single particle / : Zeng JW, Wang XW

PDB-7wh8: 
SARS-CoV-2 spike in complex with the ZB8 neutralizing antibody Fab (focused refinement on Fab-RBD)
Method: single particle / : Zeng JW

PDB-7whb: 
SARS-CoV-2 spike in complex with the ZB8 neutralizing antibody Fab (3U)
Method: single particle / : Zeng JW, Ge JW, Wang XQ

PDB-7whd: 
SARS-CoV-2 spike in complex with the ZB8 neutralizing antibody Fab (2u1d)
Method: single particle / : Zeng JW, Wang XW, Ge JW, Wang ZY
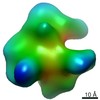
EMDB-9878: 
Cryo EM density map of Resveratrol-stabilized bioactive insulin oligomer
Method: single particle / : Sengupta J, Pathak BK

PDB-6jr3: 
Crystal structure of insulin hexamer fitted into cryo EM density map where each dimer was kept as rigid body
Method: single particle / : Sengupta J, Pathak BK, Bhakta S
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
