-Search query
-Search result
Showing all 39 items for (author: calcraft & t)

EMDB entry, No image
EMDB-17309: 
In situ cryoEM structure of Prototype Foamy Virus Env trimer

EMDB entry, No image
EMDB-17311: 
In situ cryoEM structure of Prototype Foamy Virus Env dimer of trimers

EMDB entry, No image
EMDB-17312: 
In situ cryoEM structure of the Prototype Foamy Virus capsid, icosahedral map

EMDB entry, No image
EMDB-17313: 
In situ cryoEM structure of the Prototype Foamy Virus capsid, pentamer localised reconstruction

EMDB entry, No image
EMDB-17314: 
In situ cryoEM structure of the Prototype Foamy Virus capsid, hexamer 1 localised reconstruction

EMDB entry, No image
EMDB-17315: 
In situ cryoEM structure of the Prototype Foamy Virus capsid, hexamer 2 localised reconstruction

EMDB entry, No image
EMDB-17316: 
In situ subtomogram average of Prototype Foamy Virus Env trimer

EMDB entry, No image
EMDB-17317: 
In situ subtomogram average of Prototype Foamy Virus Env pentamer of trimers

EMDB entry, No image
EMDB-17318: 
In situ subtomogram average of Prototype Foamy Virus Env hexamer of trimers

EMDB entry, No image
EMDB-17319: 
In situ subtomogram average of the Prototype Foamy Virus capsid, wild-type Gag

EMDB entry, No image
EMDB-17320: 
In situ subtomogram average of the Prototype Foamy Virus capsid, p68 Gag

EMDB entry, No image
EMDB-17321: 
Cryotomogram of Prototype Foamy Virus particles, wild-type Gag

EMDB entry, No image
EMDB-17322: 
Cryotomogram of Prototype Foamy Virus particles, p68 Gag

PDB-8ozh: 
In situ cryoEM structure of Prototype Foamy Virus Env trimer

PDB-8ozj: 
In situ cryoEM structure of Prototype Foamy Virus Env dimer of trimers

PDB-8ozk: 
In situ cryoEM structure of the Prototype Foamy Virus capsid, icosahedral map

PDB-8ozl: 
In situ cryoEM structure of the Prototype Foamy Virus capsid, pentamer localised reconstruction

PDB-8ozm: 
In situ cryoEM structure of the Prototype Foamy Virus capsid, hexamer 1 localised reconstruction

PDB-8ozn: 
In situ cryoEM structure of the Prototype Foamy Virus capsid, hexamer 2 localised reconstruction

PDB-8ozp: 
In situ subtomogram average of Prototype Foamy Virus Env pentamer of trimers

PDB-8ozq: 
In situ subtomogram average of Prototype Foamy Virus Env hexamer of trimers
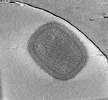
EMDB-18916: 
Cryotomogram of mature Vaccinia virus (WR) virion

EMDB-18917: 
Subtomogram average of the Vaccinia virus (WR) portal complex in mature virions

EMDB-18918: 
Subtomogram average of the Vaccinia virus (WR) A4/A10 palisade trimer in mature virions

PDB-8r5i: 
In situ structure of the Vaccinia virus (WR) A4/A10 palisade trimer in mature virions by flexible fitting into a cryoET map

EMDB-15596: 
In situ subtomogram average of Vaccinia virus (WR) palisade, all virions

EMDB-15597: 
In situ subtomogram average of Vaccinia virus (WR) palisade, from CEVs

EMDB-15598: 
In situ subtomogram average of Vaccinia virus (WR) palisade, from IMVs

EMDB-15599: 
In situ subtomogram average of Vaccinia virus (WR) palisade, from IEVs

EMDB-15600: 
Cryotomogram of Vaccinia virus (WR) infected HeLa cell (immature virions)

EMDB-15601: 
Cryotomogram of Vaccinia virus (WR) infected HeLa cell (mature virions)

EMDB-15602: 
In situ subtomogram average of Vaccinia virus (WR) D13 lattice, on immature virions

PDB-8arh: 
In situ subtomogram average of Vaccinia virus (WR) D13 lattice, on immature virions

EMDB-15181: 
Prefusion spike of SARS-CoV-2 (Wuhan), closed conformation

EMDB-15182: 
Electron cryotomography of SARS-CoV-2 virions

EMDB-15183: 
Whole SARS-CoV-2 (Wuhan) virion

EMDB-15185: 
Prefusion spike of SARS-CoV-2 (Wuhan), 1-RBD-up conformation

EMDB-11928: 
SctV (SsaV) cytoplasmic domain

PDB-7awa: 
SctV (SsaV) cytoplasmic domain
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
