-Search query
-Search result
Showing 1 - 50 of 629 items for (author: tai & l)

EMDB-19986: 
Structure of recombinant alpha-synuclein fibrils 1B capable of seeding GCIs in vivo
Method: helical / : Burger D, Kashyrina M, Lewis A, De Nuccio F, Mohammed I, de La Seigliere H, van den Heuvel L, Feuillie C, Verchere J, Berbon M, Arotcarena M, Retailleau A, Bezard E, Laferriere F, Loquet A, Bousset L, Baron T, Lofrumento DD, De Giorgi F, Stahlberg H, Ichas F

PDB-9euu: 
Structure of recombinant alpha-synuclein fibrils 1B capable of seeding GCIs in vivo
Method: helical / : Burger D, Kashyrina M, Lewis A, De Nuccio F, Mohammed I, de La Seigliere H, van den Heuvel L, Feuillie C, Verchere J, Berbon M, Arotcarena M, Retailleau A, Bezard E, Laferriere F, Loquet A, Bousset L, Baron T, Lofrumento DD, De Giorgi F, Stahlberg H, Ichas F

EMDB-35943: 
Cryo-EM structure of FFAR2 complex in apo state
Method: single particle / : Tai L, Li F, Sun X, Tang W, Wang J

PDB-8j23: 
Cryo-EM structure of FFAR2 complex in apo state
Method: single particle / : Tai L, Li F, Sun X, Tang W, Wang J
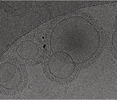
EMDB-16813: 
Tomogram of GBP1 coatomers assembled on brain polar lipid-derived small unilamellar vesicles.
Method: electron tomography / : Kuhm TI, Jakobi AJ

EMDB-16814: 
Tomogram of GBP1 coatomers assembled on brain polar lipid-derived small unilamellar vesicles.
Method: electron tomography / : Kuhm TI, Jakobi AJ

EMDB-16815: 
Tomogram of GBP1 coatomers assembled on brain polar lipid-derived small unilamellar vesicles.
Method: electron tomography / : Kuhm TI, Jakobi AJ

EMDB-41625: 
Type IV pilus from Pseudomonas PAO1 strain
Method: single particle / : Thongchol J, Zhang J, Zeng L

EMDB-41632: 
Asymmetric reconstruction of mature PP7 virions
Method: single particle / : Thongchol J, Zhang J, Zeng L

EMDB-41633: 
Type IV pilus from Pseudomonas PAO1 strain with PP7 Maturation protein
Method: single particle / : Thongchol J, Zhang J, Zeng L
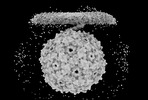
EMDB-41675: 
The original consensus map of PP7 binding to T4P from PAO1
Method: single particle / : Thongchol J, Zhang J, Zeng L

EMDB-41780: 
Composite map of PP7 binding to type IV pilus from PAO1
Method: single particle / : Thongchol J, Zhang J, Zeng L

PDB-8tum: 
Type IV pilus from Pseudomonas PAO1 strain
Method: single particle / : Thongchol J, Zhang J, Zeng L

PDB-8tuw: 
Type IV pilus from Pseudomonas PAO1 strain with PP7 Maturation protein
Method: single particle / : Thongchol J, Zhang J, Zeng L

PDB-8tux: 
Capsid of mature PP7 virion with 3'end region of PP7 genomic RNA
Method: single particle / : Thongchol J, Zhang J, Zeng L

EMDB-16794: 
Cryo-EM structure of the human GBP1 dimer bound to GDP-AlF3
Method: single particle / : Kuhm TI, Jakobi AJ

EMDB-40687: 
PS3 F1 Rotorless, no ATP
Method: single particle / : Sobti M, Stewart AG

EMDB-40688: 
PS3 F1 Rotorless, low ATP
Method: single particle / : Sobti M, Stewart AG

EMDB-40689: 
PS3 F1 Rotorless, high ATP
Method: single particle / : Sobti M, Stewart AG

PDB-8spv: 
PS3 F1 Rotorless, no ATP
Method: single particle / : Sobti M, Stewart AG

PDB-8spw: 
PS3 F1 Rotorless, low ATP
Method: single particle / : Sobti M, Stewart AG

PDB-8spx: 
PS3 F1 Rotorless, high ATP
Method: single particle / : Sobti M, Stewart AG

EMDB-35940: 
Cryo-EM structure of FFAR3 bound with valeric acid and AR420626
Method: single particle / : Tai L, Li F, Sun X, Tang W, Wang J

EMDB-35941: 
Cryo-EM structure of FFAR3 complex bound with butyrate acid
Method: single particle / : Tai L, Li F, Sun X, Tang W, Wang J

EMDB-35942: 
Cryo-EM structure of FFAR2 complex bound with TUG-1375
Method: single particle / : Tai L, Li F, Sun X, Tang W, Wang J

EMDB-35944: 
Cryo-EM structure of FFAR2 complex bound with acetic acid
Method: single particle / : Tai L, Li F, Tang W, Sun X, Wang J

PDB-8j20: 
Cryo-EM structure of FFAR3 bound with valeric acid and AR420626
Method: single particle / : Tai L, Li F, Sun X, Tang W, Wang J

PDB-8j21: 
Cryo-EM structure of FFAR3 complex bound with butyrate acid
Method: single particle / : Tai L, Li F, Sun X, Tang W, Wang J

PDB-8j22: 
Cryo-EM structure of FFAR2 complex bound with TUG-1375
Method: single particle / : Tai L, Li F, Sun X, Tang W, Wang J

PDB-8j24: 
Cryo-EM structure of FFAR2 complex bound with acetic acid
Method: single particle / : Tai L, Li F, Tang W, Sun X, Wang J

EMDB-35932: 
Local refined cryo-EM reconstruction of Omicron BA.5 RBD in complex with 8-9D Fab
Method: single particle / : Xiang Y, Li R

EMDB-35934: 
Cryo-EM reconstruction of SARS-CoV2 Omicron BA.5 spike in complex with 8-9D Fabs
Method: single particle / : Xiang Y, Li R

PDB-8j1t: 
Local refined cryo-EM structure of Omicron BA.5 RBD in complex with 8-9D Fab
Method: single particle / : Xiang Y, Li R

PDB-8j1v: 
Cryo-EM structure of SARS-CoV2 Omicron BA.5 spike in complex with 8-9D Fabs
Method: single particle / : Xiang Y, Li R

EMDB-41983: 
Structure of the phage immune evasion protein Gad1 bound to the Gabija GajAB complex
Method: single particle / : Antine SP, Johnson AG, Mooney SE, Mayer ML, Kranzsuch PJ

PDB-8u7i: 
Structure of the phage immune evasion protein Gad1 bound to the Gabija GajAB complex
Method: single particle / : Antine SP, Johnson AG, Mooney SE, Mayer ML, Kranzusch PJ

EMDB-36636: 
Cryo-EM structure of SARS-CoV-1 2P spike protein in complex with W328-6A1 IgG
Method: single particle / : Zhu JY

EMDB-36647: 
Cryo-EM structure of SARS-CoV-1 2P spike protein in complex with W328-6E10 Fab
Method: single particle / : Zhu JY

EMDB-36648: 
Cryo-EM structure of SARS-CoV-1 2P spike protein protomer in complex with W328-6E10 (local refine)
Method: single particle / : Zhu JY

EMDB-34550: 
Immune complex of W322-3D10 IgG binding the RBD of SARS-CoV-1 2P spike protein
Method: single particle / : Zhu JY, Zhou BN

EMDB-34553: 
Immune complex of W328-5B1 IgG binding the RBD of SARS-CoV-1 2P spike protein
Method: single particle / : Zhu JY, Zhou BN

EMDB-34557: 
Immune complex of W322-3E12 IgG binding the RBD of SARS-CoV-1 2P spike protein
Method: single particle / : Zhu JY, Zhou BN

EMDB-34558: 
Immune complex of W305-2A5 IgG binding the RBD of SARS-CoV-1 2P spike protein
Method: single particle / : Zhu JY, Zhou BN

EMDB-34559: 
Immune complex of W313-6D11 IgG binding the RBD of SARS-CoV-1 2P spike protein
Method: single particle / : Zhu JY, Zhou BN

EMDB-34560: 
Immune complex of W322-3B1 Fab binding the RBD of SARS-CoV-1 2P spike protein
Method: single particle / : Zhu JY, Zhou BN

EMDB-34561: 
Immune complex of W322-3G5 IgG binding the RBD of SARS-CoV-1 2P spike protein
Method: single particle / : Zhu JY, Zhou BN

EMDB-34562: 
Immune complex of W328-6F1 IgG binding the RBD of SARS-CoV-1 2P spike protein
Method: single particle / : Zhu JY, Zhou BN

EMDB-34599: 
Immune complex of W305-2G7 IgG binding the RBD of SARS-CoV-1 2P spike protein
Method: single particle / : Zhu JY, Zhou BN

EMDB-34600: 
Immune complex of W305-2A4 Fab binding the RBD of SARS-CoV-1 2P spike protein
Method: single particle / : Zhu JY, Zhou BN

EMDB-34601: 
Immune complex of W328-6G6 Fab binding the RBD of SARS-CoV-1 2P spike protein
Method: single particle / : Zhu JY, Zhou BN
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
