-Search query
-Search result
Showing all 41 items for (author: pancera & m)

EMDB-28135: 
Cryo-EM structure of L9 Fab in complex with rsCSP
Method: single particle / : Martin GM, Ward AB

PDB-8eh5: 
Cryo-EM structure of L9 Fab in complex with rsCSP
Method: single particle / : Martin GM, Ward AB
EMDB-28192: 
Cryo-EM structure of a potent anti-malarial antibody L9 in complex with Plasmodium falciparum circumsporozoite protein (PfCSP)(dominant class)
Method: single particle / : Tripathi P, Kwong PD

EMDB-28196: 
Cryo-EM structure of a potent anti-malarial antibody L9 in complex with Plasmodium falciparum circumsporozoite protein (PfCSP)(class 2)
Method: single particle / : Tripathi P, Kwong PD

PDB-8ek1: 
Cryo-EM structure of a potent anti-malarial antibody L9 in complex with Plasmodium falciparum circumsporozoite protein (PfCSP)(dominant class)
Method: single particle / : Tripathi P, Kwong PD

PDB-8eka: 
Cryo-EM structure of a potent anti-malarial antibody L9 in complex with Plasmodium falciparum circumsporozoite protein (PfCSP)(class 2)
Method: single particle / : Tripathi P, Kwong PD

EMDB-27418: 
Cryo-EM Structure of HPIV3 prefusion F trimer in complex with 3x1 Fab
Method: single particle / : Rodarte JV, Pancera M

EMDB-27419: 
Cryo-EM Structure of RSV prefusion F trimer in complex with three MxR Fabs
Method: single particle / : Rodarte JV, Pancera M

PDB-8dg8: 
Cryo-EM Structure of HPIV3 prefusion F trimer in complex with 3x1 Fab
Method: single particle / : Rodarte JV, Pancera M

PDB-8dg9: 
Cryo-EM Structure of RSV prefusion F trimer in complex with three MxR Fabs
Method: single particle / : Rodarte JV, Pancera M

EMDB-27194: 
CIS43 Variant 10 bound to pfCSP Class 4
Method: single particle / : Gorman J, Kwong PD

EMDB-27195: 
CIS43 Variant 10 bound to pfCSP Class 2
Method: single particle / : Gorman J, Kwong PD
EMDB-27202: 
CIS43 Variant 10 bound to pfCSP Class 1
Method: single particle / : Gorman J, Kwong PD

EMDB-27222: 
CIS43 Variant 10 bound to pfCSP Class 3
Method: single particle / : Gorman J, Kwong PD

EMDB-25498: 
CV3-25 IgG in complex with SARS-CoV-2 6P-D614G S protein
Method: single particle / : Jackson AM, Ozorowski G, Ward AB
EMDB-20344: 
Polyclonal serum of rabbit C3 post 5th immunization in complex with heterologous Env trimer 45_01dG5 NFL TD 2CC+
Method: single particle / : Turner HL, Andrew AB

EMDB-20259: 
HIV Env BG505 NFL TD+ in complex with antibody E70 fragment antigen binding
Method: single particle / : Ozorowski G, Torres JL, Ward AB

EMDB-20260: 
HIV Env 16055 NFL TD 2CC+ in complex with antibody 1C2 fragment antigen binding
Method: single particle / : Ozorowski G, Torres JL, Ward AB
EMDB-20273: 
Negative-stain reconstruction of HIV-1 Env BG505 NFL CC+ trimer in complex with rabbit antibody E70 fragment antigen binding
Method: single particle / : Torres JL, Ozorowski G, Andrew AB
EMDB-20274: 
Negative stain reconstruction of HIV-1 Env 16055 NFL TD 2CC+ trimer in complex with rabbit antibody 1C2 fragment antigen binding
Method: single particle / : Torres JL, Ozorowski G, Andrew AB
EMDB-20279: 
Polyclonal Fabs from rabbit C3 post 5th immunization in complex with heterologous Env trimer 45_01dH5
Method: single particle / : Cottrell CA, Andrew AB

EMDB-20280: 
Polyclonal Fabs from rabbit C3 post 5th immunization in complex with heterologous Env trimer 45_01dH5
Method: single particle / : Cottrell CA, Andrew AB
EMDB-20342: 
Polyclonal serum of rabbit C3 boost 5 in complex with heterologous Env trimer SC422 NFL TD CC+
Method: single particle / : Turner HL, Andrew AB
EMDB-20343: 
Polyclonal serum from rabbit C3 post 4th immunization in complex with heterologous Env trimer 45_01dG5 NFL TD 2CC+
Method: single particle / : Turner HL, Andrew AB
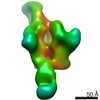
EMDB-20345: 
Polyclonal serum of rabbit A1 post 6th immunization in complex with homologous Env trimer 16055 NFL TD CC+
Method: single particle / : Turner HL, Andrew AB
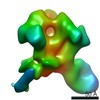
EMDB-20346: 
Polyclonal serum of rabbit A1 post 6th immunization in complex with heterologous Env trimer 45_01dG5 NFL TD 2CC+, class 1
Method: single particle / : Turner HL, Andrew AB
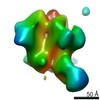
EMDB-20347: 
Polyclonal serum of rabbit A1 post 6th immunization in complex with heterologous Env trimer 45_01dG5 NFL TD 2CC+, class 2
Method: single particle / : Turner HL, Andrew AB

PDB-6p62: 
HIV Env BG505 NFL TD+ in complex with antibody E70 fragment antigen binding
Method: single particle / : Ozorowski G, Torres JL, Ward AB

PDB-6p65: 
HIV Env 16055 NFL TD 2CC+ in complex with antibody 1C2 fragment antigen binding
Method: single particle / : Ozorowski G, Torres JL, Ward AB

EMDB-9294: 
Germline VRC01 antibody recognition of a modified clade C HIV-1 envelope trimer, 3 Fabs bound, sharpened map
Method: single particle / : Borst AJ, Weidle CE, Gray MD, Frenz B, Snijder J, Joyce MG, Georgiev IS, Stewart-Jones GBE, Kwong PD, McGuire AT, DiMaio F, Stamatatos L, Pancera M, Veesler D

EMDB-9295: 
Germline VRC01 antibody recognition of a modified clade C HIV-1 envelope trimer, 3 Fabs bound, unsharpened map
Method: single particle / : Borst AJ, Weidle CE

EMDB-9303: 
Germline VRC01 antibody recognition of a modified clade C HIV-1 envelope trimer, 2 Fabs bound, sharpened map
Method: single particle / : Borst AJ, Weidle CE, Gray MD, Frenz B, Snijder J, Joyce MG, Georgiev IS, Stewart-Jones GBE, Kwong PD, McGuire AT, DiMaio F, Stamatatos L, Pancera M, Veesler D

EMDB-9304: 
Germline VRC01 antibody recognition of a modified clade C HIV-1 envelope trimer, 2 Fabs bound, unsharpened map
Method: single particle / : Borst AJ, Weidle CE

PDB-6myy: 
Germline VRC01 antibody recognition of a modified clade C HIV-1 envelope trimer, 3 Fabs bound, sharpened map
Method: single particle / : Borst AJ, Weidle CE, Gray MD, Frenz B, Snijder J, Joyce MG, Georgiev IS, Stewart-Jones GBE, Kwong PD, McGuire AT, DiMaio F, Stamatatos L, Pancera M, Veesler D

PDB-6mzj: 
Germline VRC01 antibody recognition of a modified clade C HIV-1 envelope trimer, 2 Fabs bound, sharpened map
Method: single particle / : Borst AJ, Weidle CE, Gray MD, Frenz B, Snijder J, Joyce MG, Georgiev IS, Stewart-Jones GBE, Kwong PD, McGuire AT, DiMaio F, Stamatatos L, Pancera M, Veesler D

EMDB-7344: 
An anti-gH/gL antibody that neutralizes dual-tropic infection defines a site of vulnerability on Epstein-Barr virus
Method: single particle / : Snijder J, Ortego MS, Weidle C, Stuart AB, Gray MA, McElrath MJ, Pancera M, Veesler D, McGuire AT, Seattle Structural Genomics Center for Infectious Disease (SSGCID)

EMDB-7345: 
An anti-gH/gL antibody that neutralizes dual-tropic infection defines a site of vulnerability on Epstein-Barr virus
Method: single particle / : Snijder J, Ortego MS

PDB-6c5v: 
An anti-gH/gL antibody that neutralizes dual-tropic infection defines a site of vulnerability on Epstein-Barr virus
Method: single particle / : Snijder J, Ortego MS, Weidle C, Stuart AB, Gray MA, McElrath MJ, Pancera M, Veesler D, McGuire AT, Seattle Structural Genomics Center for Infectious Disease (SSGCID)

EMDB-8125: 
BG505 SOSIP.664 HIV-1 Env trimer in complex with anti-HIV fusion peptide targeting N123-VRC34.01 Fab
Method: single particle / : Ozorowski G, Ward AB
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search





 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
