[English] 日本語
 Yorodumi
Yorodumi- EMDB-9294: Germline VRC01 antibody recognition of a modified clade C HIV-1 e... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-9294 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Germline VRC01 antibody recognition of a modified clade C HIV-1 envelope trimer, 3 Fabs bound, sharpened map | |||||||||
 Map data Map data | Germline VRC01 antibody recognition of a modified clade C HIV-1 envelope trimer, 3 Fabs bound | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | HIV envelope / germline VRC01 antibody / viral fusion protein / glycan shield / VIRAL PROTEIN-IMMUNE SYSTEM complex | |||||||||
| Function / homology |  Function and homology information Function and homology informationimmunoglobulin complex / symbiont-mediated perturbation of host defense response / positive regulation of plasma membrane raft polarization / positive regulation of receptor clustering / host cell endosome membrane / clathrin-dependent endocytosis of virus by host cell / adaptive immune response / viral protein processing / fusion of virus membrane with host plasma membrane / fusion of virus membrane with host endosome membrane ...immunoglobulin complex / symbiont-mediated perturbation of host defense response / positive regulation of plasma membrane raft polarization / positive regulation of receptor clustering / host cell endosome membrane / clathrin-dependent endocytosis of virus by host cell / adaptive immune response / viral protein processing / fusion of virus membrane with host plasma membrane / fusion of virus membrane with host endosome membrane / viral envelope / virion attachment to host cell / host cell plasma membrane / virion membrane / structural molecule activity / extracellular region / membrane / plasma membrane Similarity search - Function | |||||||||
| Biological species |   Human immunodeficiency virus 1 / Human immunodeficiency virus 1 /  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.8 Å | |||||||||
 Authors Authors | Borst AJ / Weidle CE | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Elife / Year: 2018 Journal: Elife / Year: 2018Title: Germline VRC01 antibody recognition of a modified clade C HIV-1 envelope trimer and a glycosylated HIV-1 gp120 core. Authors: Andrew J Borst / Connor E Weidle / Matthew D Gray / Brandon Frenz / Joost Snijder / M Gordon Joyce / Ivelin S Georgiev / Guillaume Be Stewart-Jones / Peter D Kwong / Andrew T McGuire / Frank ...Authors: Andrew J Borst / Connor E Weidle / Matthew D Gray / Brandon Frenz / Joost Snijder / M Gordon Joyce / Ivelin S Georgiev / Guillaume Be Stewart-Jones / Peter D Kwong / Andrew T McGuire / Frank DiMaio / Leonidas Stamatatos / Marie Pancera / David Veesler /  Abstract: VRC01 broadly neutralizing antibodies (bnAbs) target the CD4-binding site (CD4) of the human immunodeficiency virus-1 (HIV-1) envelope glycoprotein (Env). Unlike mature antibodies, corresponding ...VRC01 broadly neutralizing antibodies (bnAbs) target the CD4-binding site (CD4) of the human immunodeficiency virus-1 (HIV-1) envelope glycoprotein (Env). Unlike mature antibodies, corresponding VRC01 germline precursors poorly bind to Env. Immunogen design has mostly relied on glycan removal from trimeric Env constructs and has had limited success in eliciting mature VRC01 bnAbs. To better understand elicitation of such bnAbs, we characterized the inferred germline precursor of VRC01 in complex with a modified trimeric 426c Env by cryo-electron microscopy and a 426c gp120 core by X-ray crystallography, biolayer interferometry, immunoprecipitation, and glycoproteomics. Our results show VRC01 germline antibodies interacted with a wild-type 426c core lacking variable loops 1-3 in the presence and absence of a glycan at position Asn276, with the latter form binding with higher affinity than the former. Interactions in the presence of an Asn276 oligosaccharide could be enhanced upon carbohydrate shortening, which should be considered for immunogen design. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_9294.map.gz emd_9294.map.gz | 88.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-9294-v30.xml emd-9294-v30.xml emd-9294.xml emd-9294.xml | 14.6 KB 14.6 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_9294.png emd_9294.png | 58.4 KB | ||
| Filedesc metadata |  emd-9294.cif.gz emd-9294.cif.gz | 6.9 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-9294 http://ftp.pdbj.org/pub/emdb/structures/EMD-9294 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-9294 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-9294 | HTTPS FTP |
-Related structure data
| Related structure data |  6myyMC  9295C  9303C  9304C  6mftC  6mzjC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_9294.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_9294.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Germline VRC01 antibody recognition of a modified clade C HIV-1 envelope trimer, 3 Fabs bound | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.36 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
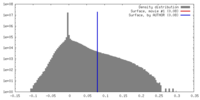
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : 426c DS-SOSIP D3
| Entire | Name: 426c DS-SOSIP D3 |
|---|---|
| Components |
|
-Supramolecule #1: 426c DS-SOSIP D3
| Supramolecule | Name: 426c DS-SOSIP D3 / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#3 |
|---|---|
| Source (natural) | Organism:   Human immunodeficiency virus 1 Human immunodeficiency virus 1 |
| Molecular weight | Theoretical: 450 KDa |
-Macromolecule #1: 426c DS-SOSIP D3
| Macromolecule | Name: 426c DS-SOSIP D3 / type: protein_or_peptide / ID: 1 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Human immunodeficiency virus 1 Human immunodeficiency virus 1 |
| Molecular weight | Theoretical: 70.243812 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: AENLWVTVYY GVPVWKEAKT TLFCASDAKA YEKEVHNVWA THACVPTDPN PQEVVLENVT ENFNMWKNDM VDQMQEDVIS IWDQSLKPC VKLTPLCVTL NCTNVNVTSN STNVNSSSTD NTTLGEIKNC SFDITTEIRD KTRKEYALFY RLDIVPLDNS S NPNSSNTY ...String: AENLWVTVYY GVPVWKEAKT TLFCASDAKA YEKEVHNVWA THACVPTDPN PQEVVLENVT ENFNMWKNDM VDQMQEDVIS IWDQSLKPC VKLTPLCVTL NCTNVNVTSN STNVNSSSTD NTTLGEIKNC SFDITTEIRD KTRKEYALFY RLDIVPLDNS S NPNSSNTY RLINCNTSTC TQACPKVTFD PIPIHYCAPA GYAILKCNNK TFNGKGPCNN VSTVQCTHGI KPVVSTQLLL NG SLAEEEI VIRSKNLADN AKIIIVQLNK SVEIVCTRPN NNTRRSIRIG PGQTFYATDI IGDIRQAYCN ISGRNWSEAV NQV KKKLKE HFPHKNISFQ SSSGGDLEIT THSFNCGGEF FYCNTSGLFN DTISNATIML PCRIKQIINM WQEVGKCIYA PPIK GNITC KSDITGLLLL RDGCNTANNA EIFRPGGGDM RDNWRSELYK YKVVKIEPLG VAPTRCKRRV VGRRRRRRAV GIGAV FLGF LGAAGSTMGA ASMTLTVQAR NLLSGIVQQQ SNLLRAPEAQ QHLLKLTVWG IKQLQARVLA VERYLRDQQL LGIWGC SGK LICCTNVPWN SSWSNRNLSE IWDNMTWLQW DKEISNYTQI IYGLLEESQN QQEKNEQDLL ALD |
-Macromolecule #2: VRC01GL light chain
| Macromolecule | Name: VRC01GL light chain / type: protein_or_peptide / ID: 2 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 25.008906 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MGWSCIILFL VATATGVHSE IVLTQSPATL SLSPGERATL SCRASQSVSS YLAWYQQKPG QAPRLLIYDA SNRATGIPAR FSGSGSGTD FTLTISSLEP EDFAVYYCQQ YEFFGQGTKL EIKRTVAAPS VFIFPPSDEQ LKSGTASVVC LLNNFYPREA K VQWKVDNA ...String: MGWSCIILFL VATATGVHSE IVLTQSPATL SLSPGERATL SCRASQSVSS YLAWYQQKPG QAPRLLIYDA SNRATGIPAR FSGSGSGTD FTLTISSLEP EDFAVYYCQQ YEFFGQGTKL EIKRTVAAPS VFIFPPSDEQ LKSGTASVVC LLNNFYPREA K VQWKVDNA LQSGNSQESV TEQDSKDSTY SLSSTLTLSK ADYEKHKVYA CEVTHQGLSS PVTKSFNRGE C UniProtKB: IGK@ protein |
-Macromolecule #3: VRC01GL heavy chain
| Macromolecule | Name: VRC01GL heavy chain / type: protein_or_peptide / ID: 3 / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 25.334305 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: (PCA)VQLVQSGAE VKKPGASVKV SCKASGYTFT GYYMHWVRQA PGQGLEWMGW INPNSGGTNY CQKFQGRVTM TRDTSI STA YMELSRLRSD DTAVYYCARG KNSDYNWDFQ HWGQGTLVTV SSASTKGPSV FPLAPSSKST SGGTAALGCL VKDYFPE PV TVSWNSGALT ...String: (PCA)VQLVQSGAE VKKPGASVKV SCKASGYTFT GYYMHWVRQA PGQGLEWMGW INPNSGGTNY CQKFQGRVTM TRDTSI STA YMELSRLRSD DTAVYYCARG KNSDYNWDFQ HWGQGTLVTV SSASTKGPSV FPLAPSSKST SGGTAALGCL VKDYFPE PV TVSWNSGALT SGVHTFPAVL QSSGLYSLSS VVTVPSSSLG TQTYICNVNH KPSNTKVDKR VEPKSCDKTH HHHHH UniProtKB: IgG H chain |
-Macromolecule #11: 2-acetamido-2-deoxy-beta-D-glucopyranose
| Macromolecule | Name: 2-acetamido-2-deoxy-beta-D-glucopyranose / type: ligand / ID: 11 / Number of copies: 12 / Formula: NAG |
|---|---|
| Molecular weight | Theoretical: 221.208 Da |
| Chemical component information |  ChemComp-NAG: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7 |
|---|---|
| Grid | Model: C-flat-1.2/1.3 4C / Pretreatment - Type: GLOW DISCHARGE |
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: GIF Bioquantum / Energy filter - Slit width: 20 eV |
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Average electron dose: 1.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: OTHER |
|---|---|
| Final reconstruction | Applied symmetry - Point group: C3 (3 fold cyclic) / Resolution.type: BY AUTHOR / Resolution: 3.8 Å / Resolution method: FSC 0.143 CUT-OFF / Software - Name: RELION / Details: Frequency-limited refinement / Number images used: 134443 |
| Initial angle assignment | Type: PROJECTION MATCHING / Software - Name: RELION |
| Final angle assignment | Type: PROJECTION MATCHING / Software - Name: RELION |
-Atomic model buiding 1
| Refinement | Protocol: OTHER |
|---|---|
| Output model |  PDB-6myy: |
 Movie
Movie Controller
Controller













 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)