-Search query
-Search result
Showing 1 - 50 of 73 items for (author: heinz & s)

EMDB-28281: 
Subtomogram average of the T4SS of Coxiella Burnetii at pH 4.75
Method: subtomogram averaging / : Kaplan M, Ghosal D

EMDB-28282: 
Subtomogram average of T4SS of Coxiella burnetii at pH 7
Method: subtomogram averaging / : Kaplan M, Shepherd DC, Vankadari N, Kim KW, Larson CL, Przemyslaw D, Beare PA, Krzymowski E, Heinzen RA, Jensen GJ, Ghosal D

EMDB-28283: 
Subtomogram average of T4SS of Coxiella burnetii at pH7 with an inner membrane mask
Method: subtomogram averaging / : Kaplan M, Shepherd DC, Vankadari N, Kim KW, Larson CL, Przemyslaw D, Beare PA, Krzymowski E, Heinzen RA, Jensen GJ, Ghosal D

EMDB-36754: 
SLC15A4 inhibitor complex
Method: single particle / : Zhang SS, Chen XD, Xie M

PDB-7uih: 
PSMD2 Structure
Method: single particle / : Johnson MC, Bashore C, Ciferri C, Dueber EC

PDB-7ujd: 
PSMD2 Structure bound to MC1 and Fab8/14
Method: single particle / : Johnson MC, Bashore C, Ciferri C, Dueber EC

EMDB-28596: 
CryoEM Structure of NLRP3 NACHT domain in complex with G2394
Method: single particle / : Murray JM, Johnson MC

PDB-8etr: 
CryoEM Structure of NLRP3 NACHT domain in complex with G2394
Method: single particle / : Murray JM, Johnson MC

EMDB-24742: 
PSMD2 with bound macrocycle MC1
Method: single particle / : Johnson MC, Bashore C, Ciferri C, Dueber EC

EMDB-24743: 
PSMD2
Method: single particle / : Johnson MC, Bashore C, Ciferri C, Dueber EC

EMDB-13930: 
3D reconstruction of the membrane domains of the sialic acid TRAP transporter HiSiaQM from Haemophilus influenzae in lipid nanodiscs bound to a high affinity megabody
Method: single particle / : Peter MF, Hagelueken G

PDB-7qe5: 
Structure of the membrane domains of the sialic acid TRAP transporter HiSiaQM from Haemophilus influenzae
Method: single particle / : Peter MF, Hagelueken G

EMDB-12676: 
Vibrio vulnificus stressosome
Method: single particle / : Kaltwasser S, Heinz V

PDB-7o0i: 
Vibrio vulnificus stressosome
Method: single particle / : Kaltwasser S, Heinz V, Madej MG, Pane-Farre J, Ziegler C

EMDB-12324: 
In situ cryo-electron tomogram of the Synechocystis wild-type VIPP1 strain grown in high light
Method: electron tomography / : Wietrzynski W, Klumpe S, Heinz S, Spaniol B, Schaffer M, Rast A, Nickelsen J, Schroda M, Engel BD

EMDB-12325: 
In situ cryo-electron tomogram of the Synechocystis F4E VIPP1 mutant grown in high light
Method: electron tomography / : Wietrzynski W, Klumpe S, Heinz S, Spaniol B, Schaffer M, Rast A, Nickelsen J, Schroda M, Engel BD

EMDB-12326: 
In situ cryo-electron tomogram of the Synechocystis V11E VIPP1 mutant grown in high light
Method: electron tomography / : Wietrzynski W, Klumpe S, Heinz S, Spaniol B, Schaffer M, Rast A, Nickelsen J, Schroda M, Engel BD

EMDB-12327: 
In situ cryo-electron tomogram of a cluster of VIPP1-mCherry structures inside the Chlamydomonas chloroplast
Method: electron tomography / : Wietrzynski W, Klumpe S, Heinz S, Spaniol B, Schaffer M, Rast A, Nickelsen J, Schroda M, Engel BD

EMDB-12328: 
In situ cryo-electron tomogram of a cluster of VIPP1-mCherry structures inside the Chlamydomonas chloroplast
Method: electron tomography / : Wietrzynski W, Klumpe S, Heinz S, Spaniol B, Schaffer M, Rast A, Nickelsen J, Schroda M, Engel BD

EMDB-12329: 
In situ cryo-electron tomogram of a cluster of VIPP1-mCherry structures inside the Chlamydomonas chloroplast
Method: electron tomography / : Wietrzynski W, Klumpe S, Heinz S, Spaniol B, Schaffer M, Rast A, Nickelsen J, Schroda M, Engel BD

EMDB-12330: 
In situ cryo-electron tomogram of a cluster of VIPP1-mCherry structures inside the Chlamydomonas chloroplast
Method: electron tomography / : Wietrzynski W, Klumpe S, Heinz S, Spaniol B, Schaffer M, Rast A, Nickelsen J, Schroda M, Engel BD

EMDB-12710: 
Structural basis for VIPP1 oligomerization and maintenance of thylakoid membrane integrity
Method: single particle / : Gupta TK, Klumpe S, Gries K, Strauss M, Rudack T, Schuller JM, Schroda M, Engel BD

EMDB-12711: 
Structural basis for VIPP1 oligomerization and maintenance of thylakoid membrane integrity
Method: single particle / : Gupta TK, Klumpe S, Gries K, Strauss M, Rudack T, Schuller JM, Schroda M, Engel BD

EMDB-12712: 
Structural basis for VIPP1 oligomerization and maintenance of thylakoid membrane integrity
Method: single particle / : Gupta TK, Klumpe S, Gries K, Strauss M, Rudack T, Schuller JM, Schroda M, Engel BD

EMDB-12713: 
Structural basis for VIPP1 oligomerization and maintenance of thylakoid membrane integrity
Method: single particle / : Gupta TK, Klumpe S, Gries K, Strauss M, Rudack T, Schuller JM, Schroda M, Engel BD

EMDB-12714: 
Structural basis for VIPP1 oligomerization and maintenance of thylakoid membrane integrity
Method: single particle / : Gupta TK, Klumpe S, Gries K, Strauss M, Rudack T, Schuller JM, Schroda M, Engel BD

PDB-7o3w: 
Structural basis for VIPP1 oligomerization and maintenance of thylakoid membrane integrity
Method: single particle / : Gupta TK, Klumpe S, Gries K, Strauss M, Rudack T, Schuller JM, Schroda M, Engel BD

PDB-7o3x: 
Structural basis for VIPP1 oligomerization and maintenance of thylakoid membrane integrity
Method: single particle / : Gupta TK, Klumpe S, Gries K, Strauss M, Rudack T, Schuller JM, Schroda M, Engel BD

PDB-7o3y: 
Structural basis for VIPP1 oligomerization and maintenance of thylakoid membrane integrity
Method: single particle / : Gupta TK, Klumpe S, Gries K, Strauss M, Rudack T, Schuller JM, Schroda M, Engel BD

PDB-7o3z: 
Structural basis for VIPP1 oligomerization and maintenance of thylakoid membrane integrity
Method: single particle / : Gupta TK, Klumpe S, Gries K, Strauss M, Rudack T, Schuller JM, Schroda M, Engel BD

PDB-7o40: 
Structural basis for VIPP1 oligomerization and maintenance of thylakoid membrane integrity
Method: single particle / : Gupta TK, Klumpe S, Gries K, Strauss M, Rudack T, Schuller JM, Schroda M, Engel BD

EMDB-30835: 
cryo EM map of the LAT1-4F2hc bound with JX-075
Method: single particle / : Yan RH, Li YN, Zhang YY, Zhong XY, Zhou Q

EMDB-30836: 
cryo EM map of the LAT1-4F2hc bound with JX-075, focused refined on transmembrane region
Method: single particle / : Yan RH, Li YN, Zhang YY, Zhong XY, Zhou Q

EMDB-30837: 
cryo EM map of the LAT1-4F2hc bound with JX-078
Method: single particle / : Yan RH, Li YN, Zhang YY, Zhong XY, Zhou Q

EMDB-30838: 
cryo EM map of the LAT1-4F2hc bound with JX-078, focused refined on transmembrane region
Method: single particle / : Yan RH, Li YN, Zhang YY, Zhong XY, Zhou Q

EMDB-30839: 
cryo EM map of the LAT1-4F2hc bound with JX-119
Method: single particle / : Yan RH, Li YN, Zhang YY, Zhong XY, Zhou Q

EMDB-30840: 
cryo EM map of the LAT1-4F2hc bound with JX-119, focused refined on transmembrane region
Method: single particle / : Yan RH, Li YN, Zhang YY, Zhong XY, Zhou Q

EMDB-30841: 
cryo EM map of the LAT1-4F2hc bound with 3,5-diiodo-L-tyrosine
Method: single particle / : Yan RH, Li YN, Zhang YY, Zhong XY, Zhou Q

EMDB-30842: 
cryo EM map of the LAT1-4F2hc bound with 3,5-diiodo-L-tyrosine, focused refined on transmembrane region
Method: single particle / : Yan RH, Li YN, Zhang YY, Zhong XY, Zhou Q

PDB-7dsk: 
Overall structure of the LAT1-4F2hc bound with JX-075
Method: single particle / : Yan RH, Li YN, Zhang YY, Zhong XY, Zhou Q

PDB-7dsl: 
Overall structure of the LAT1-4F2hc bound with JX-078
Method: single particle / : Yan RH, Li YN, Zhang YY, Zhong XY, Zhou Q

PDB-7dsn: 
Overall structure of the LAT1-4F2hc bound with JX-119
Method: single particle / : Yan RH, Li YN, Zhang YY, Zhong XY, Zhou Q

PDB-7dsq: 
Overall structure of the LAT1-4F2hc bound with 3,5-diiodo-L-tyrosine
Method: single particle / : Yan RH, Li YN, Zhang YY, Zhong XY, Zhou Q

EMDB-23156: 
SARS-CoV 2 Spike Protein bound to LY-CoV555
Method: single particle / : Goldsmith JA, McLellan JS

PDB-7l3n: 
SARS-CoV 2 Spike Protein bound to LY-CoV555
Method: single particle / : Goldsmith JA, McLellan JS

EMDB-10266: 
Homo sapiens WRB/CAML heterotetramer in complex with a TRC40 dimer
Method: single particle / : McDowell MA, Heimes M, Wild K, Flemming D, Sinning I

EMDB-11607: 
Saccharomyces cerevisiae Get1/Get2 heterotetramer in complex with a Get3 dimer
Method: single particle / : McDowell MA, Pfeffer S, Flemming D, Sinning I

PDB-6so5: 
Homo sapiens WRB/CAML heterotetramer in complex with a TRC40 dimer
Method: single particle / : McDowell MA, Heimes M, Wild K, Flemming D, Sinning I
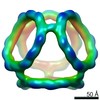
EMDB-21026: 
Truncated Tetrahedral RNA Nanostructures
Method: single particle / : Wu W, Zakrevsky P, Heinz W, Khant H, Kasprzak WK, Bindewald E, Dorjsuren N, Fields E, De Val N, Jaeger L, Shapiro BA
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search




 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
