7ONP
 
 | | Wild type carbonic anhydrase II with bound IrCp* complex to generate an artificial transfer hydrogenase (ATHase) | | Descriptor: | 4-[2-(9-chloranyl-2',3',4',5',6'-pentamethyl-4-nitro-7-oxidanylidene-spiro[1$l^{4},8-diaza-9$l^{8}-iridabicyclo[4.3.0]nona-1,3,5-triene-9,1'-1$l^{8}-iridapentacyclo[2.2.0.0^{1,3}.0^{1,5}.0^{2,6}]hexane]-8-yl)ethyl]benzenesulfonamide, ACETATE ION, Carbonic anhydrase 2, ... | | Authors: | Stein, A, Dongping, C, Cotelle, Y, Rebelein, J.G, Ward, T.R. | | Deposit date: | 2021-05-25 | | Release date: | 2021-12-01 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.408 Å) | | Cite: | A Dual Anchoring Strategy for the Directed Evolution of Improved Artificial Transfer Hydrogenases Based on Carbonic Anhydrase.
Acs Cent.Sci., 7, 2021
|
|
7ONM
 
 | | Carbonic anhydrase II mutant (N67G-E69R-I91C) dually binding an IrCp* complex to generate an artificial transfer hydrogenase (ATHase) | | Descriptor: | 1,2-ETHANEDIOL, 4-[2-(4-azanyl-9-chloranyl-2',3',4',5',6'-pentamethyl-7-oxidanylidene-spiro[1$l^{4},8-diaza-9$l^{8}-iridabicyclo[4.3.0]nona-1,3,5-triene-9,1'-1$l^{8}-iridapentacyclo[2.2.0.0^{1,3}.0^{1,5}.0^{2,6}]hexane]-8-yl)ethyl]benzenesulfonamide, Carbonic anhydrase 2, ... | | Authors: | Stein, A, Dongping, C, Cotelle, Y, Rebelein, J.G, Ward, T.R. | | Deposit date: | 2021-05-25 | | Release date: | 2021-12-01 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.769 Å) | | Cite: | A Dual Anchoring Strategy for the Directed Evolution of Improved Artificial Transfer Hydrogenases Based on Carbonic Anhydrase.
Acs Cent.Sci., 7, 2021
|
|
7ONQ
 
 | | Carbonic anhydrase II mutant (E69C) dually binding an IrCp* complex to generate an artificial transfer hydrogenase (ATHase) | | Descriptor: | 1,2-ETHANEDIOL, 4-[2-(4-azanyl-9-chloranyl-2',3',4',5',6'-pentamethyl-7-oxidanylidene-spiro[1$l^{4},8-diaza-9$l^{8}-iridabicyclo[4.3.0]nona-1,3,5-triene-9,1'-1$l^{8}-iridapentacyclo[2.2.0.0^{1,3}.0^{1,5}.0^{2,6}]hexane]-8-yl)ethyl]benzenesulfonamide, Carbonic anhydrase 2, ... | | Authors: | Stein, A, Dongping, C, Cotelle, Y, Rebelein, J.G, Ward, T.R. | | Deposit date: | 2021-05-25 | | Release date: | 2021-12-01 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | A Dual Anchoring Strategy for the Directed Evolution of Improved Artificial Transfer Hydrogenases Based on Carbonic Anhydrase.
Acs Cent.Sci., 7, 2021
|
|
7ONV
 
 | | Carbonic anhydrase II mutant (I91C) dually binding an IrCp* complex to generate an artificial transfer hydrogenase (ATHase) | | Descriptor: | 1,2-ETHANEDIOL, 4-[2-(4-azanyl-9-chloranyl-2',3',4',5',6'-pentamethyl-7-oxidanylidene-spiro[1$l^{4},8-diaza-9$l^{8}-iridabicyclo[4.3.0]nona-1,3,5-triene-9,1'-1$l^{8}-iridapentacyclo[2.2.0.0^{1,3}.0^{1,5}.0^{2,6}]hexane]-8-yl)ethyl]benzenesulfonamide, Carbonic anhydrase 2, ... | | Authors: | Stein, A, Dongping, C, Cotelle, Y, Rebelein, J.G, Ward, T.R. | | Deposit date: | 2021-05-26 | | Release date: | 2021-12-01 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.04 Å) | | Cite: | A Dual Anchoring Strategy for the Directed Evolution of Improved Artificial Transfer Hydrogenases Based on Carbonic Anhydrase.
Acs Cent.Sci., 7, 2021
|
|
3QQZ
 
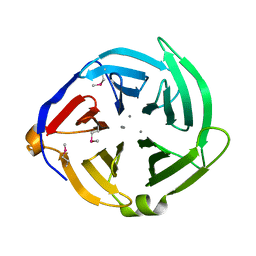 | | Crystal structure of the C-terminal domain of the yjiK protein from Escherichia coli CFT073 | | Descriptor: | CALCIUM ION, Putative uncharacterized protein yjiK | | Authors: | Stein, A, Chhor, G, Nocek, B, Fenske, R.J, Clancy, S, Joachimiak, A, Midwest Center for Structural Genomics (MCSG) | | Deposit date: | 2011-02-16 | | Release date: | 2011-03-09 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2.55 Å) | | Cite: | Crystal structure of the C-terminal domain of the yjiK protein from Escherichia coli CFT073
TO BE PUBLISHED
|
|
3HD7
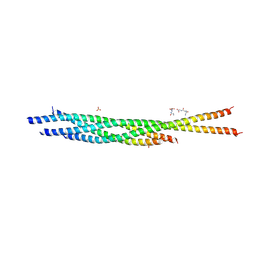 
 | | HELICAL EXTENSION OF THE NEURONAL SNARE COMPLEX INTO THE MEMBRANE, spacegroup C 1 2 1 | | Descriptor: | SULFATE ION, Synaptosomal-associated protein 25, Syntaxin-1A, ... | | Authors: | Stein, A, Weber, G, Wahl, M.C, Jahn, R. | | Deposit date: | 2009-05-07 | | Release date: | 2009-07-14 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (3.4 Å) | | Cite: | Helical extension of the neuronal SNARE complex into the membrane
Nature, 460, 2009
|
|
3IPD
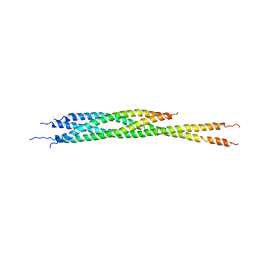 
 | | Helical extension of the neuronal SNARE complex into the membrane, spacegroup I 21 21 21 | | Descriptor: | Synaptosomal-associated protein 25, Syntaxin-1A, Vesicle-associated membrane protein 2 | | Authors: | Stein, A, Weber, G, Wahl, M.C, Jahn, R. | | Deposit date: | 2009-08-17 | | Release date: | 2009-09-01 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (4.8 Å) | | Cite: | Helical extension of the neuronal SNARE complex into the membrane
Nature, 460, 2009
|
|
6Z0I
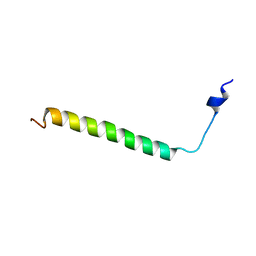 
 | | Structure of the TREM2 transmembrane helix in complex with DAP12 in DPC micelles | | Descriptor: | Triggering receptor expressed on myeloid cells 2 | | Authors: | Steiner, A, Schlepkow, K, Brunner, B, Steiner, H, Haass, C, Hagn, F. | | Deposit date: | 2020-05-08 | | Release date: | 2020-09-16 | | Last modified: | 2024-06-19 | | Method: | SOLUTION NMR | | Cite: | gamma-Secretase cleavage of the Alzheimer risk factor TREM2 is determined by its intrinsic structural dynamics.
Embo J., 2020
|
|
6Z0G
 
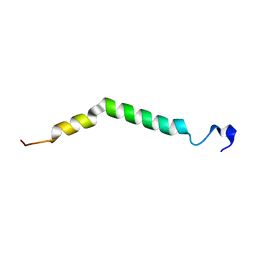 | | Structure of TREM2 transmembrane helix in DPC micelles | | Descriptor: | Triggering receptor expressed on myeloid cells 2 | | Authors: | Steiner, A, Schlepkow, K, Brunner, B, Steiner, H, Haass, C, Hagn, F. | | Deposit date: | 2020-05-08 | | Release date: | 2020-09-16 | | Last modified: | 2024-06-19 | | Method: | SOLUTION NMR | | Cite: | gamma-Secretase cleavage of the Alzheimer risk factor TREM2 is determined by its intrinsic structural dynamics.
Embo J., 39, 2020
|
|
6Z0H
 
 | | Structure of TREM2 transmembrane helix K186A variant in DPC micelles | | Descriptor: | Triggering receptor expressed on myeloid cells 2 | | Authors: | Steiner, A, Schlepkow, K, Brunner, B, Steiner, H, Haass, C, Hagn, F. | | Deposit date: | 2020-05-08 | | Release date: | 2020-09-16 | | Last modified: | 2024-06-19 | | Method: | SOLUTION NMR | | Cite: | gamma-Secretase cleavage of the Alzheimer risk factor TREM2 is determined by its intrinsic structural dynamics.
Embo J., 39, 2020
|
|
8S4T
 
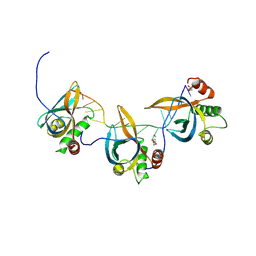 | |
8S4S
 
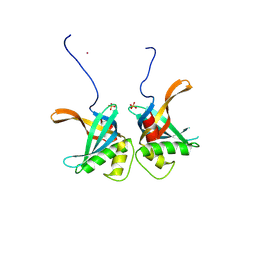 | | PrgE from plasmid pCF10 | | Descriptor: | POTASSIUM ION, PrgE, SULFATE ION | | Authors: | Breidenstein, A, Berntsson, R.P.-A. | | Deposit date: | 2024-02-22 | | Release date: | 2024-05-29 | | Last modified: | 2024-07-03 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | PrgE: an OB-fold protein from plasmid pCF10 with striking differences to prototypical bacterial SSBs.
Life Sci Alliance, 7, 2024
|
|
6Z2T
 
 | |
2Z2C
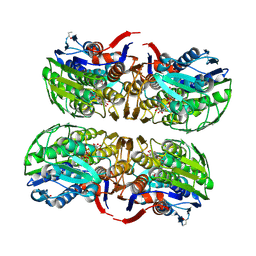 
 | | MURA inhibited by unag-cnicin adduct | | Descriptor: | 2-acetamido-3-O-[(2S,3S)-2-carboxy-3,4-dihydroxybutan-2-yl]-2-deoxy-alpha-D-glucopyranose, UDP-N-acetylglucosamine 1-carboxyvinyltransferase, URIDINE-5'-DIPHOSPHATE | | Authors: | Steinbach, A, Skarzynski, T, Scheidig, A.J, Klein, C.D. | | Deposit date: | 2007-05-17 | | Release date: | 2008-05-20 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.05 Å) | | Cite: | The Antibacterial Target Enzyme Mura Synthesizes its Own Non-Covalent Suicide Inhibitor from the Natural Product Cnicin
To be Published
|
|
8A1B
 
 | |
8A1C
 
 | |
7OND
 
 | |
7OO4
 
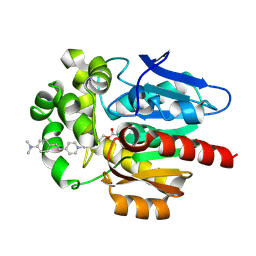 | |
3HS9
 
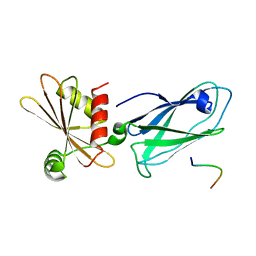 | | Intersectin 1-peptide-AP2 beta ear complex | | Descriptor: | AP-2 complex subunit beta-1, peptide from Intersectin-1, residues 841-851 | | Authors: | Vahedi-Faridi, A, Pechstein, A, Schaefer, J.G, Saenger, W, Haucke, V. | | Deposit date: | 2009-06-10 | | Release date: | 2010-02-23 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.15 Å) | | Cite: | Regulation of synaptic vesicle recycling by complex formation between intersectin 1 and the clathrin adaptor complex AP2.
Proc.Natl.Acad.Sci.USA, 107, 2010
|
|
3HS8
 
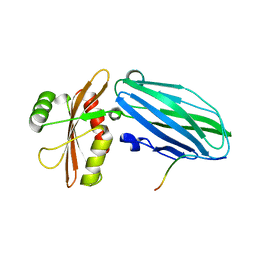 | | Intersectin 1-peptide-AP2 alpha ear complex | | Descriptor: | Adaptor protein complex AP-2, alpha 2 subunit, peptide from Intersectin-1, ... | | Authors: | Vahedi-Faridi, A, Pechstein, A, Schaefer, J.G, Saenger, W, Haucke, V. | | Deposit date: | 2009-06-10 | | Release date: | 2010-02-23 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Regulation of synaptic vesicle recycling by complex formation between intersectin 1 and the clathrin adaptor complex AP2.
Proc.Natl.Acad.Sci.USA, 107, 2010
|
|
6GOL
 
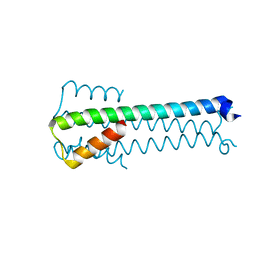 | |
3RPF
 
 | | Protein-protein complex of subunit 1 and 2 of Molybdopterin-converting factor from Helicobacter pylori 26695 | | Descriptor: | 1,2-ETHANEDIOL, Molybdopterin converting factor, subunit 1 (MoaD), ... | | Authors: | Nocek, B, Stein, A, Marshall, N, Jedrzejczak, R, Babnigg, G, Joachimiak, A, Midwest Center for Structural Genomics (MCSG) | | Deposit date: | 2011-04-26 | | Release date: | 2011-06-29 | | Last modified: | 2012-01-11 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Protein-protein complex of subunit 1 and 2 of Molybdopterin-converting factor from Helicobacter pylori 26695
TO BE PUBLISHED
|
|
2KOG
 
 | | lipid-bound synaptobrevin solution NMR structure | | Descriptor: | Vesicle-associated membrane protein 2 | | Authors: | Ellena, J.F, Liang, B, Wiktor, M, Stein, A, Cafiso, D.S, Jahn, R, Tamm, L.K. | | Deposit date: | 2009-09-22 | | Release date: | 2009-12-01 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Dynamic structure of lipid-bound synaptobrevin suggests a nucleation-propagation mechanism for trans-SNARE complex formation.
Proc.Natl.Acad.Sci.USA, 106, 2009
|
|
3PES
 
 | | Crystal structure of uncharacterized protein from Pseudomonas phage YuA | | Descriptor: | Uncharacterized protein gp49 | | Authors: | Nocek, B, Stein, A, Evdokimove, A, Egorova, O, Savchenko, A, Edwards, A, Joachimiak, A, Midwest Center for Structural Genomics (MCSG) | | Deposit date: | 2010-10-27 | | Release date: | 2010-12-08 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.3 Å) | | Cite: | Crystal structure of uncharacterized protein from Pseudomonas phage YuA
TO BE PUBLISHED
|
|
5V7V
 
 | | Cryo-EM structure of ERAD-associated E3 ubiquitin-protein ligase component HRD3 | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, ERAD-associated E3 ubiquitin-protein ligase component HRD3, beta-D-mannopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose | | Authors: | Mi, W, Schoebel, S, Stein, A, Rapoport, T.A, Liao, M. | | Deposit date: | 2017-03-20 | | Release date: | 2017-08-16 | | Last modified: | 2020-07-29 | | Method: | ELECTRON MICROSCOPY (3.9 Å) | | Cite: | Cryo-EM structure of the protein-conducting ERAD channel Hrd1 in complex with Hrd3.
Nature, 548, 2017
|
|
