4LBM
 
 | | Crystal structure of Human galectin-3 CRD in complex with LNT | | Descriptor: | CHLORIDE ION, Galectin-3, beta-D-galactopyranose-(1-3)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-3)-beta-D-galactopyranose-(1-4)-beta-D-glucopyranose | | Authors: | Collins, P.M, Blanchard, H. | | Deposit date: | 2013-06-20 | | Release date: | 2014-01-22 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.55 Å) | | Cite: | Galectin-3 interactions with glycosphingolipids.
J.Mol.Biol., 426, 2014
|
|
4LBO
 
 | |
4LBN
 
 | | Crystal structure of Human galectin-3 CRD in complex with LNnT | | Descriptor: | CHLORIDE ION, Galectin-3, beta-D-galactopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-3)-beta-D-galactopyranose-(1-4)-alpha-D-glucopyranose | | Authors: | Collins, P.M, Blanchard, H. | | Deposit date: | 2013-06-20 | | Release date: | 2014-01-22 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Galectin-3 interactions with glycosphingolipids.
J.Mol.Biol., 426, 2014
|
|
6EW3
 
 | | Crystal structure of the metallo-beta-lactamase VIM-2 with ML302F | | Descriptor: | (2Z)-2-sulfanyl-3-(2,3,6-trichlorophenyl)prop-2-enoic acid, CHLORIDE ION, DIMETHYL SULFOXIDE, ... | | Authors: | Collins, P.M, Brem, J, McDonough, M.A, van Berkel, S.S, von Delft, F, Schofield, C.J. | | Deposit date: | 2017-11-03 | | Release date: | 2018-10-03 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.14 Å) | | Cite: | Structure activity relationship studies on rhodanines and derived enethiol inhibitors of metallo-beta-lactamases.
Bioorg. Med. Chem., 26, 2018
|
|
6F83
 
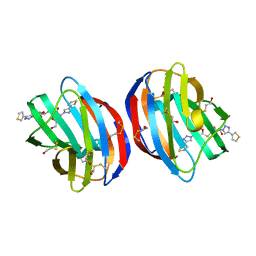 | | Crystal Structure of Human Galectin-1 in Complex With Thienyl-1,2, 3-triazolyl Thiodigalactoside Inhibitor | | Descriptor: | 3-deoxy-3-[4-(thiophen-3-yl)-1H-1,2,3-triazol-1-yl]-beta-D-galactopyranosyl 3-deoxy-1-thio-3-[4-(thiophen-3-yl)-1H-1,2,3-triazol-1-yl]-beta-D-galactopyranoside, Galectin-1 | | Authors: | Collins, P.M, Blanchard, H. | | Deposit date: | 2017-12-12 | | Release date: | 2019-01-30 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Aromatic heterocycle galectin-1 interactions for selective single-digit nM affinity ligands
To be published
|
|
3T1M
 
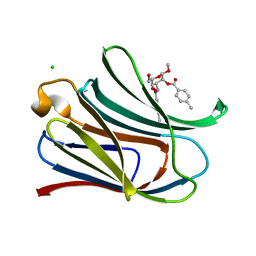 | |
3T1L
 
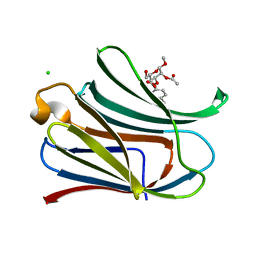 | |
3T2T
 
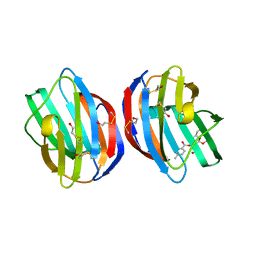 | |
4JC1
 
 | |
7RGY
 
 | |
7RH0
 
 | |
7RH1
 
 | |
7RGZ
 
 | |
7RH3
 
 | |
7RH4
 
 | |
2NN8
 
 | | Crystal structure of human galectin-3 carbohydrate-recognition domain with lactose bound, at 1.35 angstrom resolution | | Descriptor: | CHLORIDE ION, GLYCEROL, Galectin-3, ... | | Authors: | Blanchard, H, Collins, P.M. | | Deposit date: | 2006-10-24 | | Release date: | 2007-03-06 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.35 Å) | | Cite: | Slow diffusion of lactose out of galectin-3 crystals monitored by X-ray crystallography: possible implications for ligand-exchange protocols.
Acta Crystallogr.,Sect.D, 63, 2007
|
|
2NMN
 
 | |
2NMO
 
 | |
3OY8
 
 | |
3OYW
 
 | |
5ORO
 
 | | Crystal structure of Aurora-A kinase in complex with an allosterically binding fragment | | Descriptor: | 3-(4-chlorophenyl)-5,6-dihydroimidazo[2,1-b][1,3]thiazole, ADENOSINE-5'-DIPHOSPHATE, Aurora kinase A, ... | | Authors: | McIntyre, P.J, Collins, P.M, von Delft, F, Bayliss, R. | | Deposit date: | 2017-08-16 | | Release date: | 2017-11-01 | | Last modified: | 2019-10-16 | | Method: | X-RAY DIFFRACTION (2.12 Å) | | Cite: | Characterization of Three Druggable Hot-Spots in the Aurora-A/TPX2 Interaction Using Biochemical, Biophysical, and Fragment-Based Approaches.
ACS Chem. Biol., 12, 2017
|
|
5ORR
 
 | | Crystal structure of Aurora-A kinase in complex with an allosterically binding fragment | | Descriptor: | 4-[4-(trifluoromethyl)phenyl]-1,2,3-thiadiazol-5-amine, ADENOSINE-5'-DIPHOSPHATE, Aurora kinase A, ... | | Authors: | McIntyre, P.J, Collins, P.M, von Delft, F, Bayliss, R. | | Deposit date: | 2017-08-16 | | Release date: | 2017-11-01 | | Last modified: | 2017-11-29 | | Method: | X-RAY DIFFRACTION (2.09 Å) | | Cite: | Characterization of Three Druggable Hot-Spots in the Aurora-A/TPX2 Interaction Using Biochemical, Biophysical, and Fragment-Based Approaches.
ACS Chem. Biol., 12, 2017
|
|
5OS0
 
 | | Crystal structure of Aurora-A kinase in complex with an allosterically binding fragment | | Descriptor: | 2-[4-(3-chlorophenyl)piperazin-1-ium-1-yl]ethanenitrile, ADENOSINE-5'-DIPHOSPHATE, Aurora kinase A, ... | | Authors: | McIntyre, P.J, Collins, P.M, von Delft, F, Bayliss, R. | | Deposit date: | 2017-08-16 | | Release date: | 2017-11-01 | | Last modified: | 2017-11-29 | | Method: | X-RAY DIFFRACTION (1.74 Å) | | Cite: | Characterization of Three Druggable Hot-Spots in the Aurora-A/TPX2 Interaction Using Biochemical, Biophysical, and Fragment-Based Approaches.
ACS Chem. Biol., 12, 2017
|
|
5ORV
 
 | | Crystal structure of Aurora-A kinase in complex with an allosterically binding fragment | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, Aurora kinase A, CHLORIDE ION, ... | | Authors: | McIntyre, P.J, Collins, P.M, von Delft, F, Bayliss, R. | | Deposit date: | 2017-08-16 | | Release date: | 2017-11-01 | | Last modified: | 2017-11-29 | | Method: | X-RAY DIFFRACTION (1.88 Å) | | Cite: | Characterization of Three Druggable Hot-Spots in the Aurora-A/TPX2 Interaction Using Biochemical, Biophysical, and Fragment-Based Approaches.
ACS Chem. Biol., 12, 2017
|
|
5OS3
 
 | | Crystal structure of Aurora-A kinase in complex with an allosterically binding fragment | | Descriptor: | (1~{R})-1-(4-ethoxyphenyl)ethanamine, ADENOSINE-5'-DIPHOSPHATE, Aurora kinase A, ... | | Authors: | McIntyre, P.J, Collins, P.M, von Delft, F, Bayliss, R. | | Deposit date: | 2017-08-16 | | Release date: | 2017-11-01 | | Last modified: | 2017-11-29 | | Method: | X-RAY DIFFRACTION (1.81 Å) | | Cite: | Characterization of Three Druggable Hot-Spots in the Aurora-A/TPX2 Interaction Using Biochemical, Biophysical, and Fragment-Based Approaches.
ACS Chem. Biol., 12, 2017
|
|
