5MJ6
 
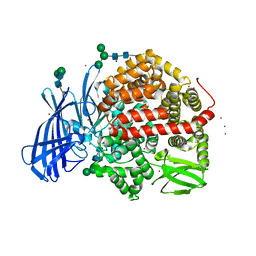 | | Ligand-induced conformational change of Insulin-regulated aminopeptidase: insights on catalytic mechanism and active site plasticity. | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, BROMIDE ION, ... | | Authors: | Mpakali, A, Stratikos, E, Saridakis, E, Giastas, P. | | Deposit date: | 2016-11-30 | | Release date: | 2017-04-05 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.53 Å) | | Cite: | Ligand-Induced Conformational Change of Insulin-Regulated Aminopeptidase: Insights on Catalytic Mechanism and Active Site Plasticity.
J. Med. Chem., 60, 2017
|
|
7B6H
 
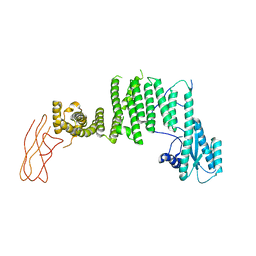 | | Drosophila melanogaster TRAPP C11 subunits region 1 to 718 | | Descriptor: | Trafficking protein particle complex subunit 11 | | Authors: | Galindo, A, Munro, S, Planelles-Herrero, V.J, Degliesposti, G. | | Deposit date: | 2020-12-07 | | Release date: | 2021-06-09 | | Last modified: | 2024-05-01 | | Method: | ELECTRON MICROSCOPY (5.4 Å) | | Cite: | Cryo-EM structure of metazoan TRAPPIII, the multi-subunit complex that activates the GTPase Rab1.
Embo J., 40, 2021
|
|
7B6E
 
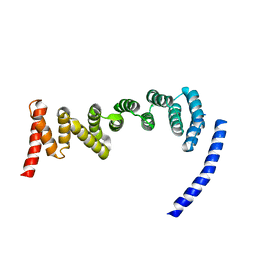 | | Drosophila melanogaster TRAPP C8 subunits region 355 to 661 | | Descriptor: | FI18195p1 | | Authors: | Galindo, A, Munro, S, Planelles-Herrero, V.J, Degliesposti, G. | | Deposit date: | 2020-12-07 | | Release date: | 2021-06-09 | | Last modified: | 2024-05-01 | | Method: | ELECTRON MICROSCOPY (4.5 Å) | | Cite: | Cryo-EM structure of metazoan TRAPPIII, the multi-subunit complex that activates the GTPase Rab1.
Embo J., 40, 2021
|
|
7B6R
 
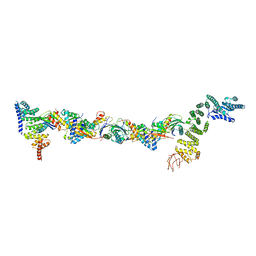 | | Drosophila melanogaster TRAPPIII partial complex: core plus C8 and C11 attached region | | Descriptor: | FI18195p1, GEO08327p1, Probable trafficking protein particle complex subunit 2, ... | | Authors: | Galindo, A, Munro, S, Planelles-Herrero, V.J, Degliesposti, G. | | Deposit date: | 2020-12-08 | | Release date: | 2021-06-09 | | Last modified: | 2021-07-28 | | Method: | ELECTRON MICROSCOPY (5.8 Å) | | Cite: | Cryo-EM structure of metazoan TRAPPIII, the multi-subunit complex that activates the GTPase Rab1.
Embo J., 40, 2021
|
|
7B6D
 
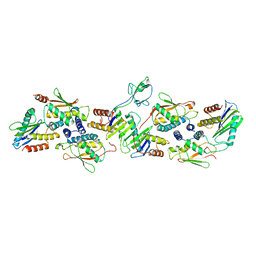 | | Drosophila melanogaster TRAPPCore (C1, C2, C2L, C3a/b, C4, C5, C6 subunits) | | Descriptor: | GEO08327p1, Probable trafficking protein particle complex subunit 2, TRAPPC2L, ... | | Authors: | Galindo, A, Munro, S, Planelles-Herrero, V.J, Degliesposti, G. | | Deposit date: | 2020-12-07 | | Release date: | 2021-06-09 | | Last modified: | 2024-05-01 | | Method: | ELECTRON MICROSCOPY (4.27 Å) | | Cite: | Cryo-EM structure of metazoan TRAPPIII, the multi-subunit complex that activates the GTPase Rab1.
Embo J., 40, 2021
|
|
7B70
 
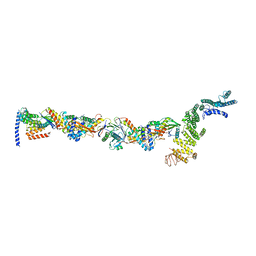 | | TRAPPCore plus C8 (355-596) and C11 (1-718) from MiniTRAPPIII | | Descriptor: | FI18195p1, GEO08327p1, Probable trafficking protein particle complex subunit 2, ... | | Authors: | Galindo, A, Munro, S, Planelles-Herrero, V.J. | | Deposit date: | 2020-12-09 | | Release date: | 2021-06-09 | | Last modified: | 2024-05-01 | | Method: | ELECTRON MICROSCOPY (4 Å) | | Cite: | Cryo-EM structure of metazoan TRAPPIII, the multi-subunit complex that activates the GTPase Rab1.
Embo J., 40, 2021
|
|
4Z7I
 
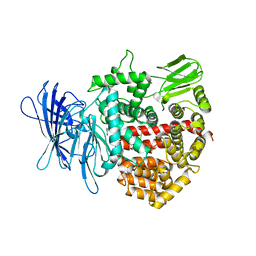 | | Crystal structure of insulin regulated aminopeptidase in complex with ligand | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, DG025 transition-state analogue enzyme inhibitor, ... | | Authors: | Mpakali, A, Saridakis, E, Harlos, K, Zhao, Y, Stratikos, E. | | Deposit date: | 2015-04-07 | | Release date: | 2015-08-26 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (3.31 Å) | | Cite: | Crystal Structure of Insulin-Regulated Aminopeptidase with Bound Substrate Analogue Provides Insight on Antigenic Epitope Precursor Recognition and Processing.
J Immunol., 195, 2015
|
|
5AB2
 
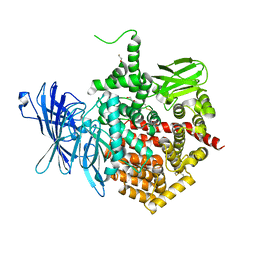 | | Crystal structure of aminopeptidase ERAP2 with ligand | | Descriptor: | 1,2-ETHANEDIOL, 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Mpakali, A, Giastas, P, Saridakis, E, Mavridis, I.M, Stratikos, E. | | Deposit date: | 2015-07-31 | | Release date: | 2015-09-30 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (2.729 Å) | | Cite: | Structural Basis for Antigenic Peptide Recognition and Processing by Endoplasmic Reticulum (Er) Aminopeptidase 2.
J.Biol.Chem., 290, 2015
|
|
5AB0
 
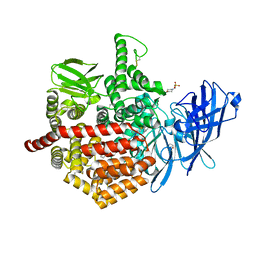 | | Crystal structure of aminopeptidase ERAP2 with ligand | | Descriptor: | 1,2-ETHANEDIOL, 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, 2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Mpakali, A, Giastas, P, Saridakis, E, Stratikos, E. | | Deposit date: | 2015-07-31 | | Release date: | 2015-09-30 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Structural Basis for Antigenic Peptide Recognition and Processing by Endoplasmic Reticulum (Er) Aminopeptidase 2.
J.Biol.Chem., 290, 2015
|
|
7NUP
 
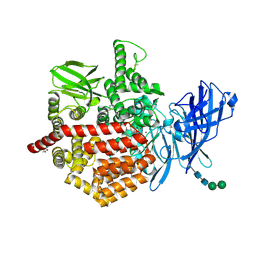 | | Endoplasmic reticulum aminopeptidase 2 complexed with a mixed hydroxamic and sulfonyl ligand | | Descriptor: | (3~{S})-4-(4-hydroxyphenyl)-~{N}-oxidanyl-3-[5-[[(5-phenylthiophen-2-yl)sulfonylamino]methyl]-1,2,3-triazol-1-yl]butanamide, 1,2-ETHANEDIOL, 2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Mpakali, A, Giastas, P, Stratikos, E. | | Deposit date: | 2021-03-12 | | Release date: | 2022-06-22 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (3.1 Å) | | Cite: | Endoplasmic reticulum aminopeptidase 2 complexed with a mixed hydroxamic and sulfonyl ligand
To Be Published
|
|
7NSK
 
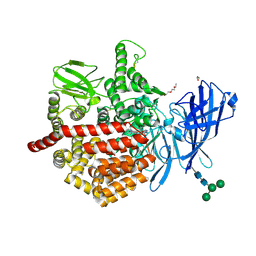 | | Endoplasmic reticulum aminopeptidase 2 complexed with a hydroxamic ligand | | Descriptor: | (2~{S})-3-(4-hydroxyphenyl)-~{N}-oxidanyl-2-[4-[[(5-pyridin-2-ylthiophen-2-yl)sulfonylamino]methyl]-1,2,3-triazol-1-yl]propanamide, 1,2-ETHANEDIOL, 1-ETHOXY-2-(2-ETHOXYETHOXY)ETHANE, ... | | Authors: | Mpakali, A, Giastas, P, Stratikos, E. | | Deposit date: | 2021-03-08 | | Release date: | 2022-06-22 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (3.1 Å) | | Cite: | Endoplasmic reticulum aminopeptidase 2 in complex with a phosphinic ligand
To Be Published
|
|
1BX0
 
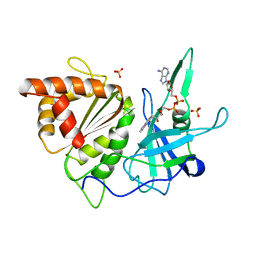 | | Ferredoxin:nadp+ oxidoreductase (ferredoxin reductase) mutant e312l | | Descriptor: | FLAVIN-ADENINE DINUCLEOTIDE, PHOSPHATE ION, PROTEIN (FERREDOXIN:NADP+ OXIDOREDUCTASE), ... | | Authors: | Aliverti, A, Deng, Z, Ravasi, D, Piubelli, L, Karplus, P.A, Zanetti, G. | | Deposit date: | 1998-10-10 | | Release date: | 1998-10-14 | | Last modified: | 2023-08-09 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Probing the function of the invariant glutamyl residue 312 in spinach ferredoxin-NADP+ reductase.
J.Biol.Chem., 273, 1998
|
|
1BX1
 
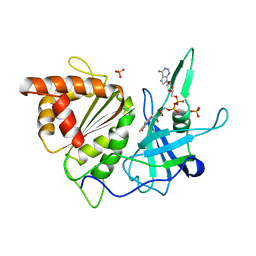 | | FERREDOXIN:NADP+ OXIDOREDUCTASE (FERREDOXIN REDUCTASE) MUTANT E312Q | | Descriptor: | FLAVIN-ADENINE DINUCLEOTIDE, PHOSPHATE ION, PROTEIN (FERREDOXIN:NADP+ OXIDOREDUCTASE), ... | | Authors: | Aliverti, A, Deng, Z, Ravasi, D, Piubelli, L, Karplus, P.A, Zanetti, G. | | Deposit date: | 1998-10-10 | | Release date: | 1998-10-14 | | Last modified: | 2023-08-09 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Probing the function of the invariant glutamyl residue 312 in spinach ferredoxin-NADP+ reductase.
J.Biol.Chem., 273, 1998
|
|
3BT7
 
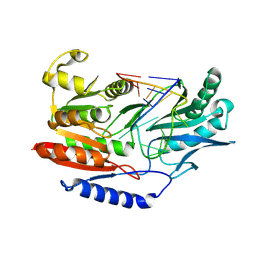 | |
8AA8
 
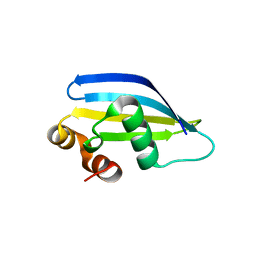 | |
8A8S
 | |
3DH3
 
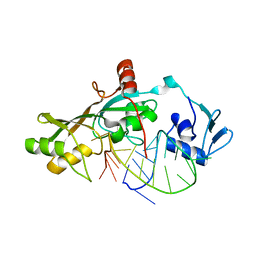 | | Crystal Structure of RluF in complex with a 22 nucleotide RNA substrate | | Descriptor: | Ribosomal large subunit pseudouridine synthase F, stem loop fragment of E. Coli 23S RNA | | Authors: | Alian, A, DeGiovanni, A, Stroud, R.M, Finer-Moore, J.S. | | Deposit date: | 2008-06-16 | | Release date: | 2009-04-28 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Crystal structure of an RluF-RNA complex: a base-pair rearrangement is the key to selectivity of RluF for U2604 of the ribosome.
J.Mol.Biol., 388, 2009
|
|
4EOR
 
 | | Thr 160 phosphorylated CDK2 WT - human cyclin A3 complex with the inhibitor NU6102 | | Descriptor: | Cyclin-A2, Cyclin-dependent kinase 2, O6-CYCLOHEXYLMETHOXY-2-(4'-SULPHAMOYLANILINO) PURINE | | Authors: | Echalier, A, Cot, E, Camasses, A, Hodimont, E, Hoh, F, Sheinerman, F, Krasinska, L, Fisher, D. | | Deposit date: | 2012-04-14 | | Release date: | 2013-02-06 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | An integrated chemical biology approach provides insight into Cdk2 functional redundancy and inhibitor sensitivity.
Chem.Biol., 19, 2012
|
|
4EON
 
 | | Thr 160 phosphorylated CDK2 H84S, Q85M, Q131E - human cyclin A3 complex with the inhibitor RO3306 | | Descriptor: | (5E)-5-(quinolin-6-ylmethylidene)-2-[(thiophen-2-ylmethyl)amino]-1,3-thiazol-4(5H)-one, Cyclin-A2, Cyclin-dependent kinase 2, ... | | Authors: | Echalier, A, Cot, E, Camasses, A, Hodimont, E, Hoh, F, Sheinerman, F, Krasinska, L, Fisher, D. | | Deposit date: | 2012-04-14 | | Release date: | 2013-02-06 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | An integrated chemical biology approach provides insight into Cdk2 functional redundancy and inhibitor sensitivity.
Chem.Biol., 19, 2012
|
|
4EOK
 
 | | Thr 160 phosphorylated CDK2 H84S, Q85M, K89D - human cyclin A3 complex with the inhibitor NU6102 | | Descriptor: | Cyclin-A2, Cyclin-dependent kinase 2, MONOTHIOGLYCEROL, ... | | Authors: | Echalier, A, Cot, E, Camasses, A, Hodimont, E, Hoh, F, Sheinerman, F, Krasinska, L, Fisher, D. | | Deposit date: | 2012-04-14 | | Release date: | 2013-02-06 | | Method: | X-RAY DIFFRACTION (2.57 Å) | | Cite: | An integrated chemical biology approach provides insight into Cdk2 functional redundancy and inhibitor sensitivity.
Chem.Biol., 19, 2012
|
|
4EOS
 
 | | Thr 160 phosphorylated CDK2 WT - human cyclin A3 complex with the inhibitor RO3306 | | Descriptor: | (5E)-5-(quinolin-6-ylmethylidene)-2-[(thiophen-2-ylmethyl)amino]-1,3-thiazol-4(5H)-one, Cyclin-A2, Cyclin-dependent kinase 2 | | Authors: | Echalier, A, Cot, E, Camasses, A, Hodimont, E, Hoh, F, Sheinerman, F, Krasinska, L, Fisher, D. | | Deposit date: | 2012-04-14 | | Release date: | 2013-02-06 | | Method: | X-RAY DIFFRACTION (2.57 Å) | | Cite: | An integrated chemical biology approach provides insight into Cdk2 functional redundancy and inhibitor sensitivity.
Chem.Biol., 19, 2012
|
|
8P0I
 
 | | Crystal structure of the open conformation of insulin-regulated aminopeptidase in complex with a small-MW inhibitor | | Descriptor: | 2-[2-(3,5-dimethylpyrazol-1-yl)-6-(4-methoxyphenyl)pyrimidin-4-yl]oxy-N-methyl-N-[(2-methylpyridin-3-yl)methyl]ethanamide, 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Mpakali, A, Giastas, P, Stratikos, E. | | Deposit date: | 2023-05-10 | | Release date: | 2024-09-04 | | Method: | X-RAY DIFFRACTION (3.5 Å) | | Cite: | Stabilization of the open conformation omicron f insulin-regulated aminopeptidase by a novel substrate-selective small-molecule inhibitor.
Protein Sci., 33, 2024
|
|
4EOM
 | | Thr 160 phosphorylated CDK2 H84S, Q85M, Q131E - human cyclin A3 complex with ATP | | Descriptor: | ADENOSINE-5'-TRIPHOSPHATE, Cyclin-A2, Cyclin-dependent kinase 2, ... | | Authors: | Echalier, A, Cot, E, Camasses, A, Hodimont, E, Hoh, F, Sheinerman, F, Krasinska, L, Fisher, D. | | Deposit date: | 2012-04-14 | | Release date: | 2013-02-06 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | An integrated chemical biology approach provides insight into Cdk2 functional redundancy and inhibitor sensitivity.
Chem.Biol., 19, 2012
|
|
4EOI
 
 | | Thr 160 phosphorylated CDK2 K89D, Q131E - human cyclin A3 complex with the inhibitor RO3306 | | Descriptor: | (5E)-5-(quinolin-6-ylmethylidene)-2-[(thiophen-2-ylmethyl)amino]-1,3-thiazol-4(5H)-one, Cyclin-A2, Cyclin-dependent kinase 2, ... | | Authors: | Echalier, A, Cot, E, Camasses, A, Hodimont, E, Hoh, F, Sheinerman, F, Krasinska, L, Fisher, D. | | Deposit date: | 2012-04-14 | | Release date: | 2013-02-06 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | An integrated chemical biology approach provides insight into Cdk2 functional redundancy and inhibitor sensitivity.
Chem.Biol., 19, 2012
|
|
4EOP
 
 | | Thr 160 phosphorylated CDK2 Q131E - human cyclin A3 complex with the inhibitor RO3306 | | Descriptor: | (5E)-5-(quinolin-6-ylmethylidene)-2-[(thiophen-2-ylmethyl)amino]-1,3-thiazol-4(5H)-one, Cyclin-A2, Cyclin-dependent kinase 2, ... | | Authors: | Echalier, A, Cot, E, Camasses, A, Hodimont, E, Hoh, F, Sheinerman, F, Krasinska, L, Fisher, D. | | Deposit date: | 2012-04-14 | | Release date: | 2013-02-06 | | Method: | X-RAY DIFFRACTION (1.99 Å) | | Cite: | An integrated chemical biology approach provides insight into Cdk2 functional redundancy and inhibitor sensitivity.
Chem.Biol., 19, 2012
|
|
