8C7D
 | | Structure of the GEF Kalirin DH2 Domain | | Descriptor: | Kalirin | | Authors: | Callens, M.C, Thompson, A.P, Gray, J.L, Bountra, C, von Delft, F, Brennan, P.E. | | Deposit date: | 2023-01-14 | | Release date: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (1.86 Å) | | Cite: | Structure-based alignment and analysis of RHO selectivity of RHO-DBL GTPase exchange factors
to be published
|
|
2KR9
 
 | |
3EO2
 
 | | Crystal structure of the RhoGEF domain of human neuroepithelial cell-transforming gene 1 protein | | Descriptor: | Neuroepithelial cell-transforming gene 1 protein, UNKNOWN ATOM OR ION | | Authors: | Nedyalkova, L, Tong, Y, Tempel, W, MacKenzie, F, Arrowsmith, C.H, Edwards, A.M, Bountra, C, Weigelt, J, Bochkarev, A, Park, H, Structural Genomics Consortium (SGC) | | Deposit date: | 2008-09-26 | | Release date: | 2008-10-07 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Crystal structure of the RhoGEF domain of human neuroepithelial cell-transforming gene 1 protein
To be Published
|
|
1BY1
 
 | | DBL homology domain from beta-PIX | | Descriptor: | PROTEIN (PIX) | | Authors: | Aghazadeh, B, Zhu, K, Kubiseski, T.J, Liu, G.A, Pawson, T, Zheng, Y, Rosen, M.K. | | Deposit date: | 1998-10-22 | | Release date: | 1999-10-24 | | Last modified: | 2023-12-27 | | Method: | SOLUTION NMR | | Cite: | Structure and Mutagenesis of the Dbl Homology Domain
Nat.Struct.Biol., 5, 1998
|
|
5N7E
 
 | | Crystal structure of the Dbl-homology domain of Bcr-Abl in complex with monobody Mb(Bcr-DH_4). | | Descriptor: | Breakpoint cluster region protein, Mb(Bcr-DH_4) | | Authors: | Reckel, S, Reynaud, A, Pojer, F, Hantschel, O. | | Deposit date: | 2017-02-20 | | Release date: | 2017-12-27 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (1.647 Å) | | Cite: | Structural and functional dissection of the DH and PH domains of oncogenic Bcr-Abl tyrosine kinase.
Nat Commun, 8, 2017
|
|
5N6R
 
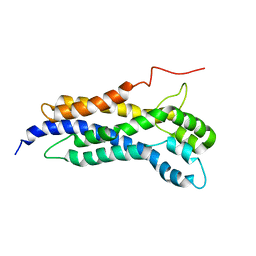 | | Solution structure of the Dbl-homology domain of Bcr-Abl | | Descriptor: | Breakpoint cluster region protein | | Authors: | Reckel, S, Lohr, F, Buchner, L, Guntert, P, Dotsch, V, Hantschel, O. | | Deposit date: | 2017-02-16 | | Release date: | 2017-12-27 | | Last modified: | 2024-06-19 | | Method: | SOLUTION NMR | | Cite: | Structural and functional dissection of the DH and PH domains of oncogenic Bcr-Abl tyrosine kinase.
Nat Commun, 8, 2017
|
|
1F5X
 
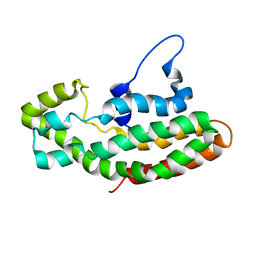 | | NMR STRUCTURE OF THE Y174 AUTOINHIBITED DBL HOMOLOGY DOMAIN | | Descriptor: | RHO-GEF VAV | | Authors: | Aghazadeh, B, Rosen, M.K, Lowry, W.E, Huang, X.Y. | | Deposit date: | 2000-06-18 | | Release date: | 2000-09-15 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Structural basis for relief of autoinhibition of the Dbl homology domain of proto-oncogene Vav by tyrosine phosphorylation.
Cell(Cambridge,Mass.), 102, 2000
|
|
4D0O
 | | AKAP13 (AKAP-Lbc) DH domain | | Descriptor: | A-KINASE ANCHOR PROTEIN 13 | | Authors: | Abdul Azeez, K.R, Shrestha, L, Krojer, T, Allerston, C, von Delft, F, Bountra, C, Arrowsmith, C, Edwards, A.M, Knapp, S, Klussmann, E, Elkins, J.M. | | Deposit date: | 2014-04-29 | | Release date: | 2014-05-21 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2.75 Å) | | Cite: | The Crystal Structure of the Rhoa : Akap-Lbc Dh-Ph Domain Complex.
Biochem.J., 464, 2014
|
|
4GYV
 
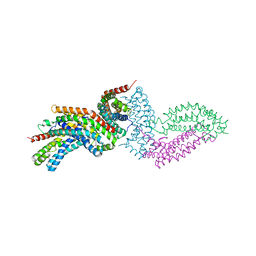 | | Crystal structure of the DH domain of FARP2 | | Descriptor: | FERM, RhoGEF and pleckstrin domain-containing protein 2 | | Authors: | He, X, Zhang, X. | | Deposit date: | 2012-09-05 | | Release date: | 2013-03-13 | | Last modified: | 2017-11-15 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Structural Basis for Autoinhibition of the Guanine Nucleotide Exchange Factor FARP2.
Structure, 21, 2013
|
|
2Z0Q
 
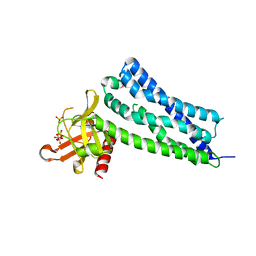 | | Crystal structure of DH-PH domain of RhoGEF3(Xpln) | | Descriptor: | Rho guanine nucleotide exchange factor 3, SULFATE ION | | Authors: | Murayama, K, Kato-Murayama, M, Terada, T, Shirouzu, M, Yokoyama, S, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2007-05-07 | | Release date: | 2008-05-13 | | Last modified: | 2012-12-12 | | Method: | X-RAY DIFFRACTION (1.79 Å) | | Cite: | Structure of the Rho-specific guanine nucleotide-exchange factor Xpln
Acta Crystallogr.,Sect.F, 68, 2012
|
|
4MT6
 
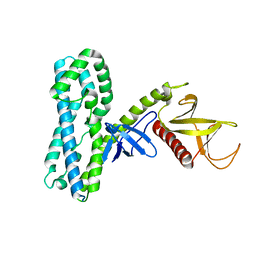 | |
1RJ2
 
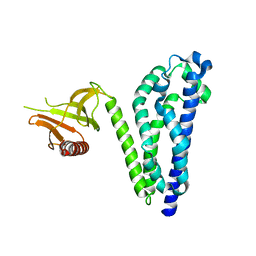 | |
1TXD
 | |
4MT7
 
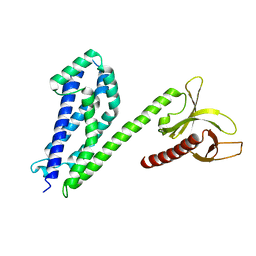 | | Crystal structure of collybistin I | | Descriptor: | Rho guanine nucleotide exchange factor 9 | | Authors: | Schneeberger, D, Schindelin, H. | | Deposit date: | 2013-09-19 | | Release date: | 2014-08-13 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (3.5 Å) | | Cite: | A conformational switch in collybistin determines the differentiation of inhibitory postsynapses.
Embo J., 33, 2014
|
|
1DBH
 
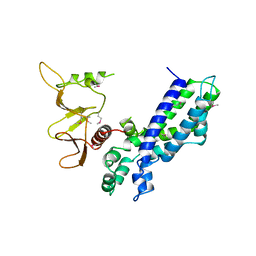 | | DBL AND PLECKSTRIN HOMOLOGY DOMAINS FROM HSOS1 | | Descriptor: | PROTEIN (HUMAN SOS 1) | | Authors: | Soisson, S.M, Kuriyan, J. | | Deposit date: | 1998-12-17 | | Release date: | 1998-12-23 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Crystal structure of the Dbl and pleckstrin homology domains from the human Son of sevenless protein.
Cell(Cambridge,Mass.), 95, 1998
|
|
5HZJ
 
 | | Crystal structure of photoinhibitable Intersectin1 containing wildtype LOV2 domain | | Descriptor: | FLAVIN MONONUCLEOTIDE, Intersectin-1,NPH1-1,Intersectin-1 | | Authors: | Tarnawski, M, Dagliyan, O, Chu, P.H, Shirvanyants, D, Dokholyan, N.V, Hahn, K.M, Schlichting, I. | | Deposit date: | 2016-02-02 | | Release date: | 2016-12-21 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Engineering extrinsic disorder to control protein activity in living cells.
Science, 354, 2016
|
|
5HZI
 
 | | Crystal structure of photoinhibitable Intersectin1 containing C450M mutant LOV2 domain | | Descriptor: | FLAVIN MONONUCLEOTIDE, Intersectin-1,NPH1-1,Intersectin-1 | | Authors: | Tarnawski, M, Dagliyan, O, Chu, P.H, Shirvanyants, D, Dokholyan, N.V, Hahn, K.M, Schlichting, I. | | Deposit date: | 2016-02-02 | | Release date: | 2016-12-21 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Engineering extrinsic disorder to control protein activity in living cells.
Science, 354, 2016
|
|
3GF9
 
 | | Crystal structure of human Intersectin 2 RhoGEF domain | | Descriptor: | Intersectin 2, UNKNOWN ATOM OR ION | | Authors: | Shen, Y, Tong, Y, Tempel, W, Li, Y, Arrowsmith, C.H, Edwards, A.M, Bountra, C, Weigelt, J, Bochkarev, A, Park, H, Structural Genomics Consortium (SGC) | | Deposit date: | 2009-02-26 | | Release date: | 2009-03-10 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Crystal structure of human Intersectin 2 RhoGEF domain
To be Published
|
|
3ODX
 
 | |
3ODO
 
 | |
3ODW
 
 | |
3P6A
 
 | |
5FI1
 
 | | Crystal Structure of the P-Rex1 DH/PH tandem in complex with Cdc42 | | Descriptor: | Cell division control protein 42 homolog, Phosphatidylinositol 3,4,5-trisphosphate-dependent Rac exchanger 1 protein,Phosphatidylinositol 3,4,5-trisphosphate-dependent Rac exchanger 1 protein | | Authors: | Cash, J.N, Tesmer, J.J.G. | | Deposit date: | 2015-12-22 | | Release date: | 2016-04-20 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (3.203 Å) | | Cite: | Structural and Biochemical Characterization of the Catalytic Core of the Metastatic Factor P-Rex1 and Its Regulation by PtdIns(3,4,5)P3.
Structure, 24, 2016
|
|
4YON
 
 | | P-Rex1:Rac1 complex | | Descriptor: | Phosphatidylinositol 3,4,5-trisphosphate-dependent Rac exchanger 1 protein, Ras-related C3 botulinum toxin substrate 1 | | Authors: | Lucato, C.M, Whisstock, J.C, Ellisdon, A.M. | | Deposit date: | 2015-03-11 | | Release date: | 2015-07-01 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | The Phosphatidylinositol (3,4,5)-Trisphosphate-dependent Rac Exchanger 1Ras-related C3 Botulinum Toxin Substrate 1 (P-Rex1Rac1) Complex Reveals the Basis of Rac1 Activation in Breast Cancer Cells.
J.Biol.Chem., 290, 2015
|
|
6PCV
 
 | | Single Particle Reconstruction of Phosphatidylinositol (3,4,5) trisphosphate-dependent Rac exchanger 1 bound to G protein beta gamma subunits | | Descriptor: | Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2, Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1, Phosphatidylinositol (3,4,5) trisphosphate-dependent Rac exchanger 1 | | Authors: | Cash, J.N, Cianfrocco, M.A, Tesmer, J.J.G. | | Deposit date: | 2019-06-18 | | Release date: | 2019-10-23 | | Last modified: | 2024-03-20 | | Method: | ELECTRON MICROSCOPY (3.2 Å) | | Cite: | Cryo-electron microscopy structure and analysis of the P-Rex1-G beta gamma signaling scaffold.
Sci Adv, 5, 2019
|
|
