Services and Softwares
Yorodumi
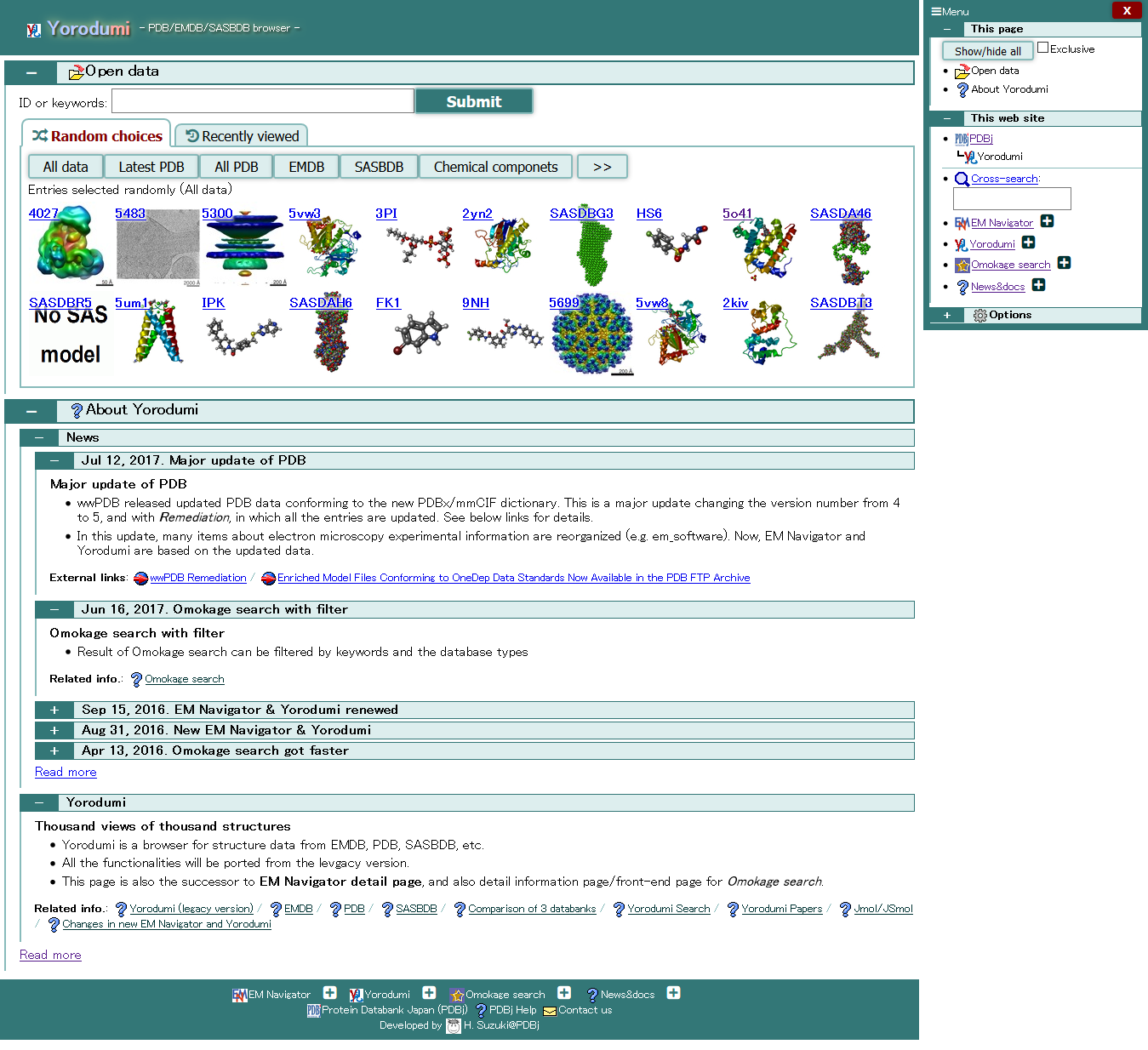
Yorodumi is a website to easily watch, move, rotate, learn, and enjoy 3D structures of biological molecules. See Yorodumi Docs for more details.
ASH

ASH is a program to perform three-dimensional structure alignment. From the ASH homepage you can run the program over the web or download the software for free.
MAFFTash
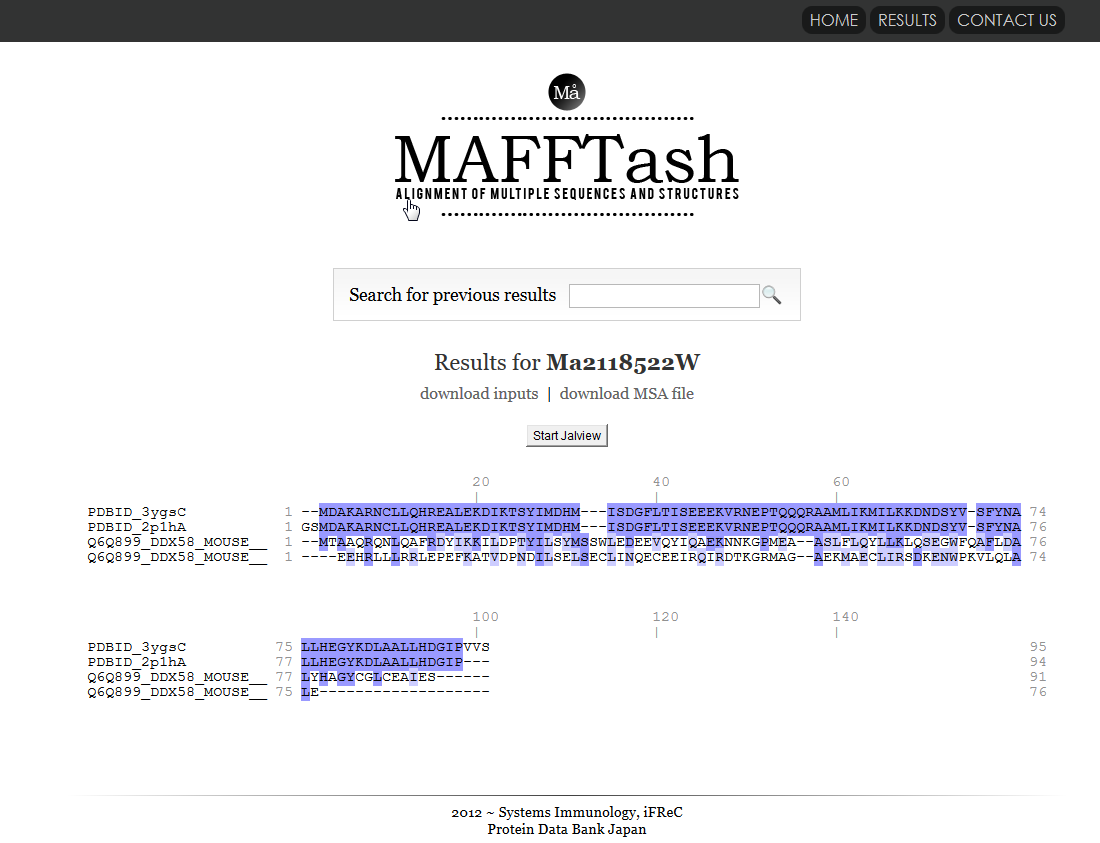
MAFFTash is a tool that calculates multple sequence alignments from sequences and structures.
CRNPRED
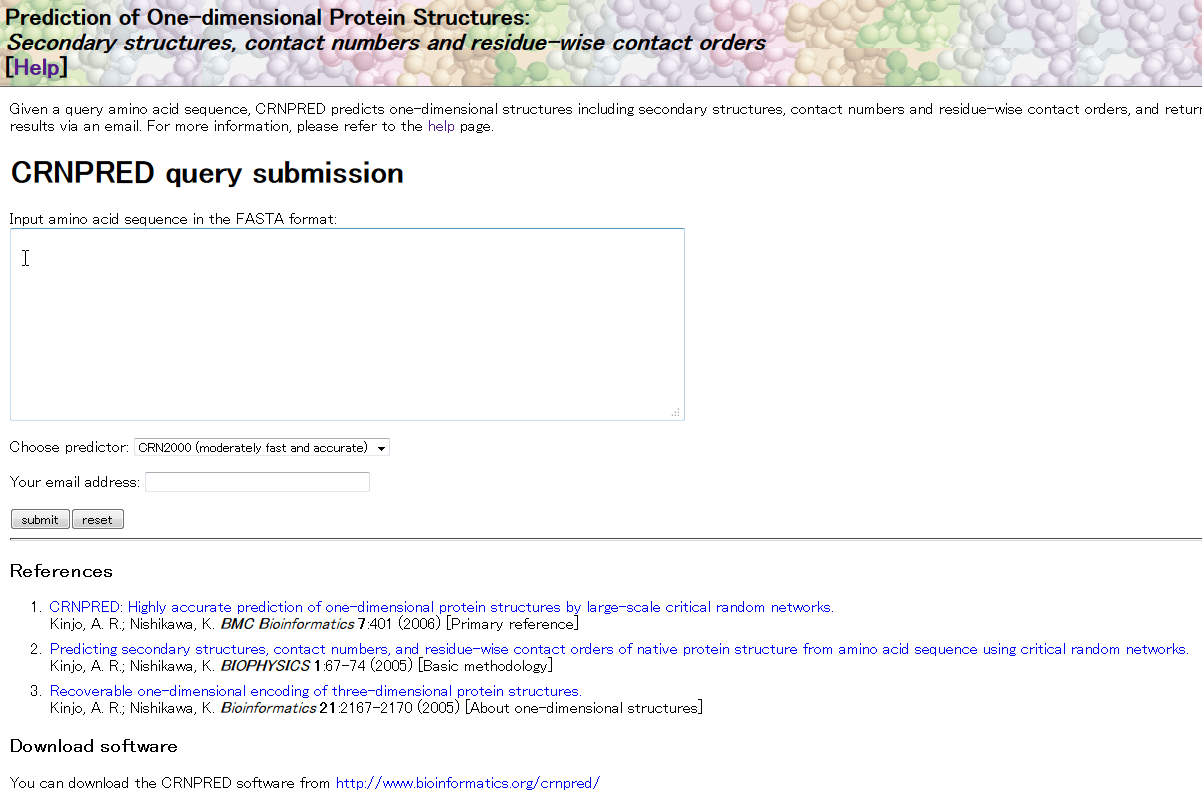
CRNPRED is a web-based service that predicts one-dimensional protein structures including secondary structures, contact numbers, and residue-wise contact orders from amino acid sequence.
Spanner
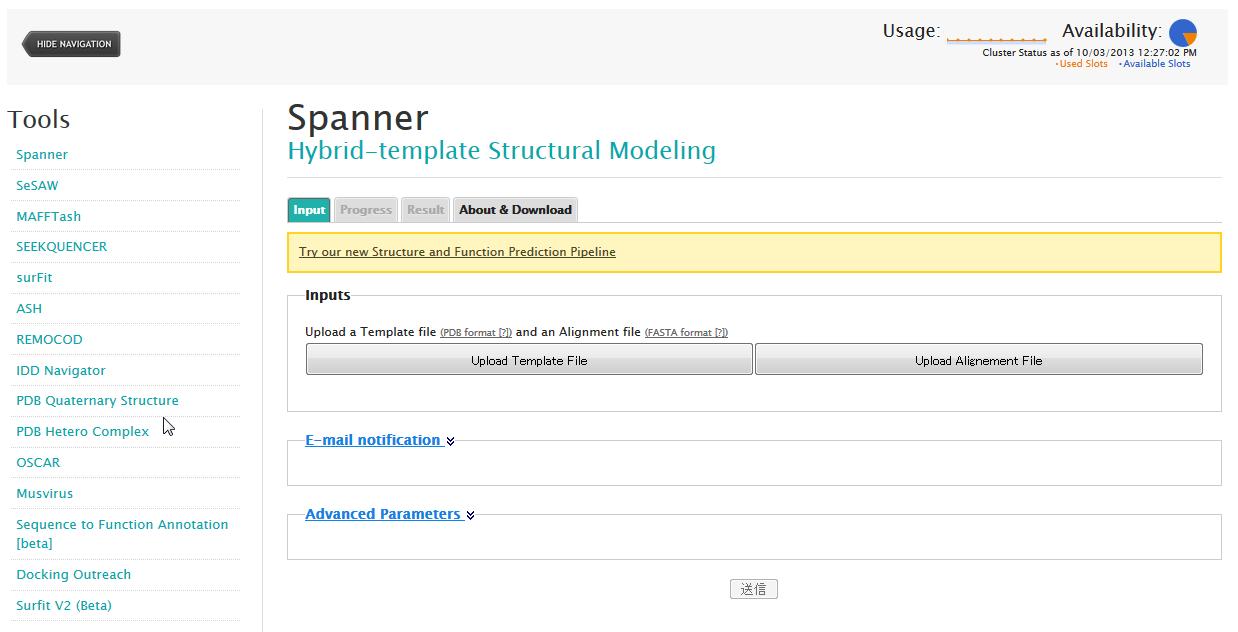
Spanner is a structural homology modeling program—that is, it threads a specific amino-acid sequence onto a specific PDB structure, patching up the gaps as best it can.
SFAS
SFAS (Sequence to Function Annotation Server) is a web-based tool for predicting the structure of an amino acid sequence. SFAS runs several external programs for sequence alignment and structural modeling and organized their results. SFAS does not do any complicated calculations. There are currently several choices for alignment and one choice for structural modeling. In the future, the number of choices will be increased.
gmfit
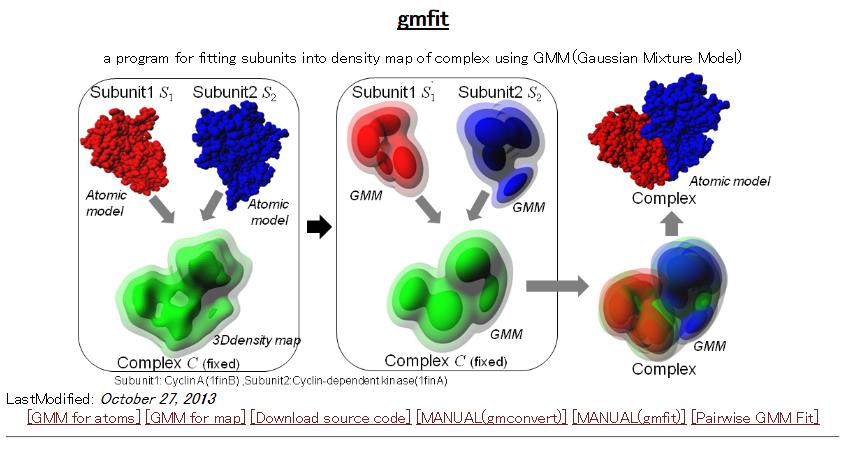
gmfit is a program for fast fitting of 3D objects using Gaussian mixture model. The fitting service between two EMDB density maps or PDB structures is available through the WEB.
HOMCOS
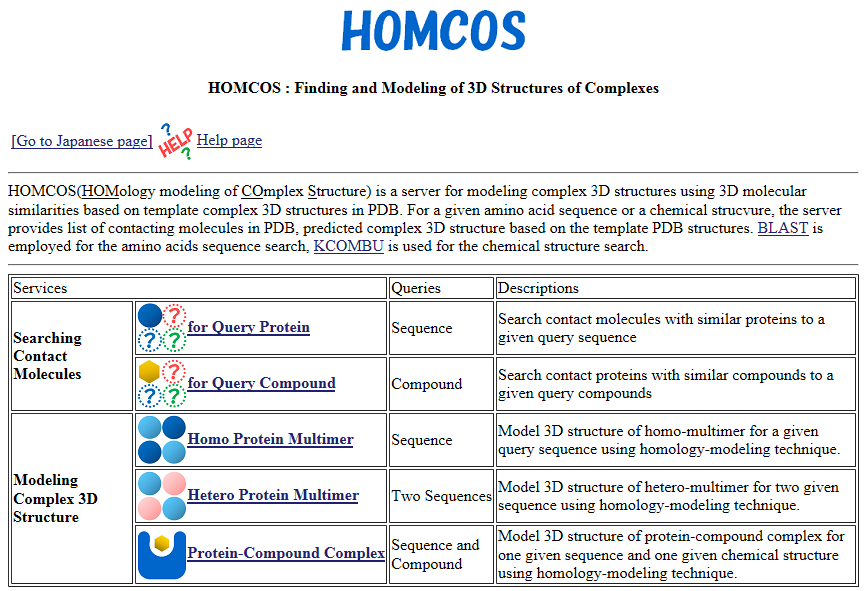

HOMCOS is a service for searching 3D complex structures in PDB from sequences or chemical structures, and for modeling 3D complex structures based on the found template structures.






