5QNT
 
 | |
5QNV
 
 | |
5QNL
 
 | |
5QNR
 
 | |
7S1F
 
 | | Crystal structure of E.coli DsbA in complex with compound MIPS-0001886 (compound 38) | | Descriptor: | 1-[(3-thiophen-3-ylphenyl)methyl]-3~{H}-pyrrol-2-one, COPPER (II) ION, GLYCEROL, ... | | Authors: | Heras, B, Scanlon, M.J, Martin, J.L, Caria, S. | | Deposit date: | 2021-09-02 | | Release date: | 2023-02-08 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (1.76 Å) | | Cite: | Fluoromethylketone-fragment conjugates designed as covalent modifiers of EcDsbA are atypical substrates
Chemrxiv, 2022
|
|
3GYK
 
 | | The crystal structure of a thioredoxin-like oxidoreductase from Silicibacter pomeroyi DSS-3 | | Descriptor: | 1,2-ETHANEDIOL, 27kDa outer membrane protein, SULFATE ION | | Authors: | Fan, Y, Marshall, N, Keigher, L, Joachimiak, A, Midwest Center for Structural Genomics (MCSG) | | Deposit date: | 2009-04-03 | | Release date: | 2009-06-09 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.76 Å) | | Cite: | The crystal structure of a thioredoxin-like oxidoreductase from Silicibacter pomeroyi DSS-3
To be Published
|
|
4OCE
 
 | |
5QO5
 
 | |
6POI
 
 | | Crystal Structure of EcDsbA in complex phenyl ether 25 | | Descriptor: | (6-phenoxy-1-benzofuran-3-yl)acetic acid, COPPER (II) ION, Thiol:disulfide interchange protein DsbA | | Authors: | Ilyichova, O.V, Scanlon, M.J. | | Deposit date: | 2019-07-04 | | Release date: | 2019-11-06 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (1.77 Å) | | Cite: | The Fragment-Based Development of a Benzofuran Hit as a New Class of Escherichia coli DsbA Inhibitors.
Molecules, 24, 2019
|
|
4ZIJ
 
 | | Crystal structure of E.Coli DsbA in complex with 2-(4-iodophenylsulfonamido) benzoic acid | | Descriptor: | 1,2-ETHANEDIOL, 2-{[(4-iodophenyl)sulfonyl]amino}benzoic acid, Thiol:disulfide interchange protein DsbA | | Authors: | Vazirani, M, Ilyichova, O.V, Scanlon, M.J. | | Deposit date: | 2015-04-28 | | Release date: | 2016-05-11 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (1.78 Å) | | Cite: | Determination of ligand binding modes in weak protein-ligand complexes using sparse NMR data.
J.Biomol.Nmr, 66, 2016
|
|
5QO4
 
 | |
5QNS
 
 | |
7LSM
 
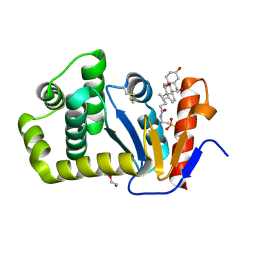 | | Crystal structure of E.coli DsbA in complex with bile salt taurocholate | | Descriptor: | DI(HYDROXYETHYL)ETHER, TAUROCHOLIC ACID, Thiol:disulfide interchange protein DsbA | | Authors: | Wang, G, Heras, B. | | Deposit date: | 2021-02-18 | | Release date: | 2021-12-29 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (1.786 Å) | | Cite: | Selective Binding of Small Molecules to Vibrio cholerae DsbA Offers a Starting Point for the Design of Novel Antibacterials.
Chemmedchem, 17, 2022
|
|
5QOG
 
 | |
5QNN
 
 | |
5QNZ
 
 | |
5QO6
 | |
7L7C
 
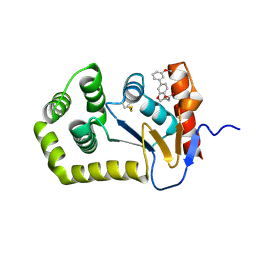 | |
8CZM
 
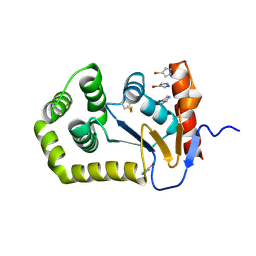 | |
8CXD
 
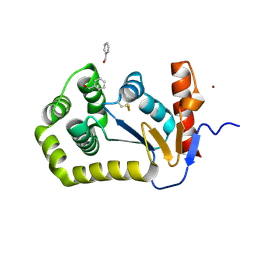 | |
6POQ
 
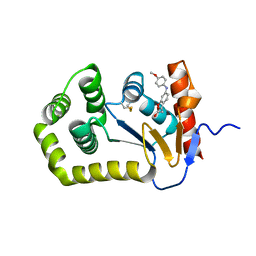 | | Crystal Structure of EcDsbA in complex with anisidine 16 | | Descriptor: | COPPER (II) ION, Thiol:disulfide interchange protein DsbA, {6-[(4-methoxyphenyl)amino]-1-benzofuran-3-yl}acetic acid | | Authors: | Ilyichova, O.V, Scanlon, M.J. | | Deposit date: | 2019-07-04 | | Release date: | 2019-11-06 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | The Fragment-Based Development of a Benzofuran Hit as a New Class of Escherichia coli DsbA Inhibitors.
Molecules, 24, 2019
|
|
3RPP
 
 | |
5HFI
 
 | |
5CP1
 
 | |
5COH
 
 | |
