3F5N
 
 | |
5OM3
 
 | | Crystal structure of Alpha1-antichymotrypsin variant DBS-I5: a MMP14-cleavable drug-binding serpin for doxycycline | | Descriptor: | (4S,4AR,5S,5AR,6R,12AS)-4-(DIMETHYLAMINO)-3,5,10,12,12A-PENTAHYDROXY-6-METHYL-1,11-DIOXO-1,4,4A,5,5A,6,11,12A-OCTAHYDROTETRACENE-2-CARBOXAMIDE, Alpha-1-antichymotrypsin, DI(HYDROXYETHYL)ETHER | | Authors: | Schmidt, K, Muller, Y.A. | | Deposit date: | 2017-07-28 | | Release date: | 2018-05-23 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Design of an allosterically modulated doxycycline and doxorubicin drug-binding protein.
Proc. Natl. Acad. Sci. U.S.A., 115, 2018
|
|
3FGQ
 
 | | Crystal structure of native human neuroserpin | | Descriptor: | GLYCEROL, Neuroserpin | | Authors: | Takehara, S, Yang, X, Mikami, B, Onda, M. | | Deposit date: | 2008-12-08 | | Release date: | 2009-04-28 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.09 Å) | | Cite: | The 2.1-A crystal structure of native neuroserpin reveals unique structural elements that contribute to conformational instability
J.Mol.Biol., 388, 2009
|
|
3ZHA
 
 | |
4AQH
 
 | | Plasminogen activator inhibitor type-1 in complex with the inhibitor AZ3976 | | Descriptor: | PLASMINOGEN ACTIVATOR INHIBITOR 1, TERT-BUTYL 3-[(4-OXO-3H-PYRIDO[2,3-D]PYRIMIDIN-2-YL)AMINO]AZETIDINE-1-CARBOXYLATE | | Authors: | Fjellstrom, O, Deinum, J, Sjogren, T, Johansson, C, Geschwindner, S, Nerme, V, Legnehed, A, McPheat, J, Olsson, K, Bodin, C, Gustafsson, D. | | Deposit date: | 2012-04-17 | | Release date: | 2012-11-28 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Characterization of a Small Molecule Inhibitor of Plasminogen Activator Inhibitor Type 1 that Accelerates the Transition Into the Latent Conformation
J.Biol.Chem., 288, 2013
|
|
4AJT
 | |
4AJU
 
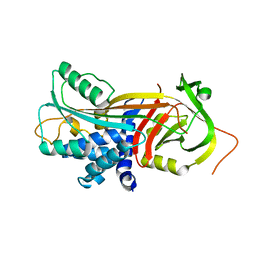
 | | Crystal structure of the reactive loop cleaved ZPI in P41 space group | | Descriptor: | 1,2-ETHANEDIOL, PROTEIN Z-DEPENDENT PROTEASE INHIBITOR, SULFATE ION | | Authors: | Zhou, A, Yan, Y, Wei, Z. | | Deposit date: | 2012-02-19 | | Release date: | 2013-01-09 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.65 Å) | | Cite: | Structural Basis for Catalytic Activation of Protein Z-Dependent Protease Inhibitor (Zpi) by Protein Z.
Blood, 120, 2012
|
|
4AFX
 
 | | Crystal structure of the reactive loop cleaved ZPI in I2 space group | | Descriptor: | CALCIUM ION, PHOSPHATE ION, PROTEIN Z DEPENDENT PROTEASE INHIBITOR, ... | | Authors: | Zhou, A, Yan, Y, Wei, Z. | | Deposit date: | 2012-01-23 | | Release date: | 2013-01-09 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.09 Å) | | Cite: | Structural Basis for Catalytic Activation of Protein Z-Dependent Protease Inhibitor (Zpi) by Protein Z.
Blood, 120, 2012
|
|
4AU4
 
 | | Crystal Structure of Hsp47 | | Descriptor: | SERPIN PEPTIDASE INHIBITOR, CLADE H (HEAT SHOCK PROTEIN 47), MEMBER 1, ... | | Authors: | Widmer, C, Gebauer, J.M, Brunstein, E, Rodenbaum, S, Zaucke, F, Drogemuller, C, Leeb, T, Baumann, U. | | Deposit date: | 2012-05-14 | | Release date: | 2012-08-15 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2.97 Å) | | Cite: | Molecular Basis for the Action of the Collagen-Specific Chaperone Hsp47/Serpinh1 and its Structure-Specific Client Recognition.
Proc.Natl.Acad.Sci.USA, 109, 2012
|
|
4AU2
 
 | | Crystal Structure of a Hsp47-collagen complex | | Descriptor: | 15ER COLLAGEN MODEL PEPTIDE 15-R8, SERPIN PEPTIDASE INHIBITOR, CLADE H (HEAT SHOCK PROTEIN 47), ... | | Authors: | Widmer, C, Gebauer, J.M, Brunstein, E, Drogemuller, C, Leeb, T, Baumann, U. | | Deposit date: | 2012-05-13 | | Release date: | 2012-08-15 | | Last modified: | 2015-04-01 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Molecular Basis for the Action of the Collagen-Specific Chaperone Hsp47/Serpinh1 and its Structure-Specific Client Recognition.
Proc.Natl.Acad.Sci.USA, 109, 2012
|
|
4BB2
 
 | | Crystal structure of cleaved corticosteroid-binding globulin in complex with progesterone | | Descriptor: | 1,2-ETHANEDIOL, CORTICOSTEROID-BINDING GLOBULIN, CYSTEINE, ... | | Authors: | Gardill, B.R, Vogl, M.R, Lin, H, Hammond, G.L, Muller, Y.A. | | Deposit date: | 2012-09-19 | | Release date: | 2012-12-26 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.48 Å) | | Cite: | Corticosteroid-Binding Globulin: Structure-Function Implications from Species Differences
Plos One, 7, 2012
|
|
4C49
 
 | | Reactive loop cleaved human CBG in complex with cortisol | | Descriptor: | (11alpha,14beta)-11,17,21-trihydroxypregn-4-ene-3,20-dione, CORTICOSTEROID-BINDING GLOBULIN | | Authors: | Chan, W.L, Zhou, A, Read, R.J. | | Deposit date: | 2013-09-02 | | Release date: | 2014-10-01 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Role of Sa-Hd Loop Movement in Cortisol Release Mechanism of Cbg
To be Published
|
|
4C41
 
 | |
4CAA
 
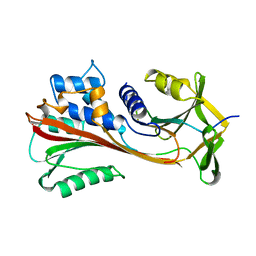 | | CLEAVED ANTICHYMOTRYPSIN T345R | | Descriptor: | ANTICHYMOTRYPSIN | | Authors: | Lukacs, C.M, Christianson, D.W. | | Deposit date: | 1997-08-14 | | Release date: | 1998-02-25 | | Last modified: | 2024-05-22 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Engineering an anion-binding cavity in antichymotrypsin modulates the "spring-loaded" serpin-protease interaction.
Biochemistry, 37, 1998
|
|
1WZ9
 
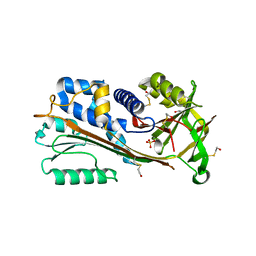 | | The 2.1 A structure of a tumour suppressing serpin | | Descriptor: | Maspin precursor, SULFATE ION | | Authors: | Law, R.H, Irving, J.A, Buckle, A.M, Ruzyla, K, Buzza, M, Bashtannyk-Puhalovich, T.A, Beddoe, T.C, Kim, N, Worrall, D.M, Bottomley, S.P, Bird, P.I, Rossjohn, J, Whisstock, J.C. | | Deposit date: | 2005-03-03 | | Release date: | 2005-03-15 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | The high resolution crystal structure of the human tumor suppressor maspin reveals a novel conformational switch in the G-helix.
J.Biol.Chem., 280, 2005
|
|
1LQ8
 
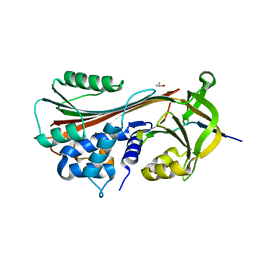 | | Crystal structure of cleaved protein C inhibitor | | Descriptor: | 2-acetamido-2-deoxy-alpha-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Huntington, J.A, Kjellberg, M, Stenflo, J. | | Deposit date: | 2002-05-09 | | Release date: | 2003-02-11 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Crystal Structure of Protein C Inhibitor Provides Insights into Hormone Binding and Heparin Activation
Structure, 11, 2003
|
|
1LK6
 
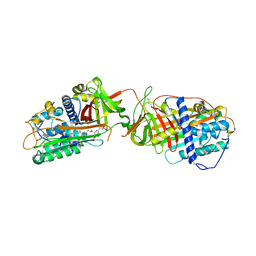 | | Structure of dimeric antithrombin complexed with a P14-P9 reactive loop peptide and an exogenous tripeptide | | Descriptor: | 2-acetamido-2-deoxy-alpha-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose, GLYCEROL, ... | | Authors: | Zhou, A, Huntington, J.A, Lomas, D.A, Carrell, R.W, Stein, P.E. | | Deposit date: | 2002-04-24 | | Release date: | 2003-06-03 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Serpin Polymerization Is Prevented by a Hydrogen Bond Network That Is Centered on His-334 and Stabilized by Glycerol
J.Biol.Chem., 278, 2003
|
|
1LJ5
 
 | |
1XU8
 
 | | The 2.8 A structure of a tumour suppressing serpin | | Descriptor: | Maspin, SULFATE ION | | Authors: | Irving, J.A, Law, R.H, Ruzyla, K, Bashtannyk-Puhalovich, T.A, Kim, N, Worrall, D.M, Rossjohn, J, Whisstock, J.C. | | Deposit date: | 2004-10-25 | | Release date: | 2005-03-15 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | The high resolution crystal structure of the human tumor suppressor maspin reveals a novel conformational switch in the G-helix.
J.Biol.Chem., 280, 2005
|
|
1XQG
 
 | |
1XQJ
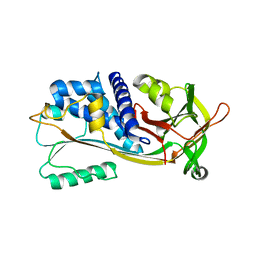 
 | |
1M93
 
 | |
1MTP
 
 | | The X-ray crystal structure of a serpin from a thermophilic prokaryote | | Descriptor: | Serine Proteinase Inhibitor (SERPIN), Chain A, Chain B | | Authors: | Irving, J.A, Cabrita, L.D, Rossjohn, J, Pike, R.N, Bottomley, S.P, Whisstock, J.C. | | Deposit date: | 2002-09-21 | | Release date: | 2003-04-15 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | The 1.5 A crystal structure of a prokaryote serpin: controlling conformational change in a heated environment
Structure, 11, 2003
|
|
1NQ9
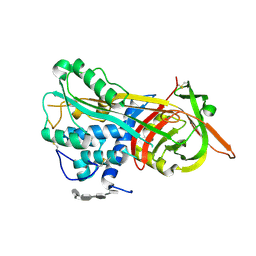 
 | | Crystal Structure of Antithrombin in the Pentasaccharide-Bound Intermediate State | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, 3,4-di-O-methyl-2,6-di-O-sulfo-alpha-D-glucopyranose-(1-4)-2,3-di-O-methyl-beta-D-glucopyranuronic acid-(1-4)-2,3,6-tri-O-sulfo-alpha-D-glucopyranose-(1-4)-3-O-methyl-2-O-sulfo-alpha-L-idopyranuronic acid-(1-4)-methyl 2,3,6-tri-O-sulfo-alpha-D-glucopyranoside, ... | | Authors: | Huntington, J.A, Johnson, D.J.D. | | Deposit date: | 2003-01-21 | | Release date: | 2003-09-30 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Crystal Structure of Antithrombin in a Heparin-Bound Intermediate State
Biochemistry, 42, 2003
|
|
1YXA
 
 | | Serpina3n, a murine orthologue of human antichymotrypsin | | Descriptor: | serine (or cysteine) proteinase inhibitor, clade A, member 3N | | Authors: | Horvath, A.J, Irving, J.A, Law, R.H, Rossjohn, J, Bottomley, S.P, Quinsey, N.S, Pike, R.N, Coughlin, P.B, Whisstock, J.C. | | Deposit date: | 2005-02-20 | | Release date: | 2005-09-06 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | The murine orthologue of human antichymotrypsin: a structural paradigm for clade A3 serpins.
J.Biol.Chem., 280, 2005
|
|
