9K75
 
 | | SARS-CoV-2 related bat coronavirus BANAL-103 spike in the closed state | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, Spike glycoprotein | | Authors: | Qingqing, L, Xiao, C, Xiaoning, L, Yibing, Z, Ru, L, Zirui, K, Didi, W, Jiaxu, W, Lili, L, Junxia, Y, Jianxiang, S, Shuiling, J, Ying, P, Na, Z, Yushun, W, Jian, S. | | Deposit date: | 2024-10-23 | | Release date: | 2025-07-16 | | Last modified: | 2026-02-04 | | Method: | ELECTRON MICROSCOPY (3.11 Å) | | Cite: | Structural and functional constraints on spike activation and host protease utilization limit cell entry of SARS-CoV-2-related bat coronaviruses.
J.Virol., 99, 2025
|
|
7T5O
 
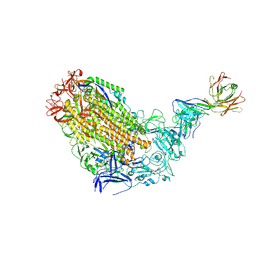 | | VFLIP Spike Trimer with GAR03 | | Descriptor: | GAR03 Fab heavy chain, GAR03 Fab light chain, Spike glycoprotein | | Authors: | Sobti, M, Stewart, A.G, Rouet, R, Langley, D.B. | | Deposit date: | 2021-12-12 | | Release date: | 2023-01-25 | | Last modified: | 2025-05-21 | | Method: | ELECTRON MICROSCOPY (3.39 Å) | | Cite: | Broadly neutralizing SARS-CoV-2 antibodies through epitope-based selection from convalescent patients.
Nat Commun, 14, 2023
|
|
7SL5
 
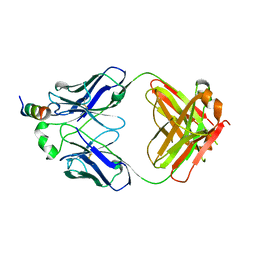 | |
7SKZ
 
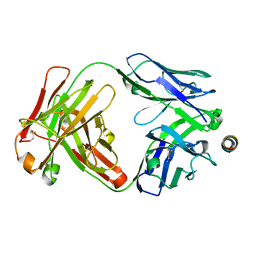 | |
7TGE
 
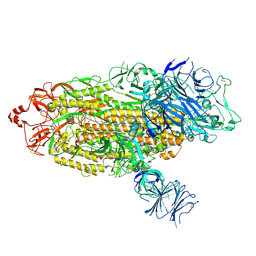 | |
8ZY2
 
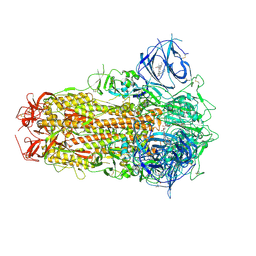 | | Sarbecovirus BANAL-20-52 Spike Trimer in a Locked Conformation | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, BILIVERDINE IX ALPHA, ... | | Authors: | Wang, J, Xiong, X. | | Deposit date: | 2024-06-16 | | Release date: | 2025-02-05 | | Last modified: | 2025-07-23 | | Method: | ELECTRON MICROSCOPY (2.9 Å) | | Cite: | SARS-related coronavirus S-protein structures reveal synergistic RBM interactions underpinning high-affinity human ACE2 binding.
Sci Adv, 11, 2025
|
|
8ZY3
 
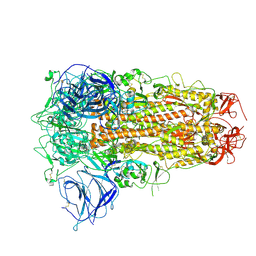 | | Sarbecovirus BANAL-20-236 Spike Trimer in a Locked Conformation | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, Spike glycoprotein | | Authors: | Wang, J, Xiong, X. | | Deposit date: | 2024-06-16 | | Release date: | 2025-02-05 | | Last modified: | 2025-07-23 | | Method: | ELECTRON MICROSCOPY (3.3 Å) | | Cite: | SARS-related coronavirus S-protein structures reveal synergistic RBM interactions underpinning high-affinity human ACE2 binding.
Sci Adv, 11, 2025
|
|
6LVN
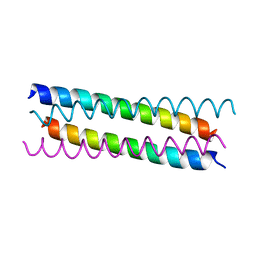 
 | | Structure of the 2019-nCoV HR2 Domain | | Descriptor: | Spike protein S2' | | Authors: | Zhu, Y, Sun, F. | | Deposit date: | 2020-02-04 | | Release date: | 2020-02-26 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.47 Å) | | Cite: | Crystal structure of HR2 domain of 2019-nCoV S2 subunit
To Be Published
|
|
6LXT
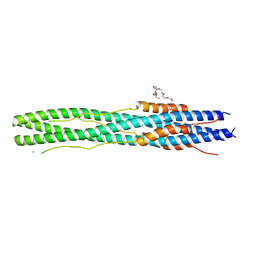 
 | | Structure of post fusion core of 2019-nCoV S2 subunit | | Descriptor: | Spike protein S2, TETRAETHYLENE GLYCOL, ZINC ION | | Authors: | Zhu, Y, Sun, F. | | Deposit date: | 2020-02-11 | | Release date: | 2020-02-26 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Inhibition of SARS-CoV-2 (previously 2019-nCoV) infection by a highly potent pan-coronavirus fusion inhibitor targeting its spike protein that harbors a high capacity to mediate membrane fusion.
Cell Res., 30, 2020
|
|
6M1V
 
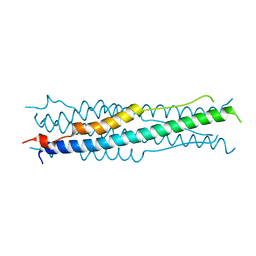 | |
9NXY
 
 | | Cryo-EM structure of SARS-CoV-2 spike S2' trimer | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, Spike protein S2, ... | | Authors: | Shi, W, Jonaid, G, Chen, B. | | Deposit date: | 2025-03-26 | | Release date: | 2025-12-31 | | Last modified: | 2026-01-14 | | Method: | ELECTRON MICROSCOPY (3.08 Å) | | Cite: | Effect of the S2' site cleavage on SARS-CoV-2 spike.
Nat Commun, 16, 2025
|
|
9NVG
 
 | | Structure of SARS-CoV-2 BA.1 spike RBD bound to COV2-3835 Fab | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, COV2-3835 Fab Heavy Chain, COV2-3835 Fab Light Chain, ... | | Authors: | Ramamohan, A.R, Johnson, N.V, McLellan, J.S. | | Deposit date: | 2025-03-20 | | Release date: | 2026-02-25 | | Last modified: | 2026-04-29 | | Method: | ELECTRON MICROSCOPY (3 Å) | | Cite: | Epitope-focused discovery of SARS-CoV-2 antibodies that potently neutralize Omicron variants.
Nat Microbiol, 11, 2026
|
|
9OG5
 
 | | SARS-COV-2-6P-MUT7 S PROTEIN-DY-III-281 complex 1 RBD up conformation | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, Spike glycoprotein, ... | | Authors: | Chandravanshi, M, Niu, L, Tolbert, W.D, Pazgier, M. | | Deposit date: | 2025-04-30 | | Release date: | 2025-10-01 | | Last modified: | 2025-12-10 | | Method: | ELECTRON MICROSCOPY (3.3 Å) | | Cite: | Optimization of VE607 to generate analogs with improved neutralization activities against SARS-CoV-2 variants.
J.Virol., 99, 2025
|
|
9OG6
 
 | | Apo SARS-COV-2-6P-MUT7 S PROTEIN closed conformation | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, Spike glycoprotein, ... | | Authors: | Niu, L, Chandravanshi, M, Tolbert, W.D, Pazgier, M. | | Deposit date: | 2025-04-30 | | Release date: | 2025-10-01 | | Last modified: | 2025-12-10 | | Method: | ELECTRON MICROSCOPY (3.14 Å) | | Cite: | Optimization of VE607 to generate analogs with improved neutralization activities against SARS-CoV-2 variants.
J.Virol., 99, 2025
|
|
9OG7
 
 | | APO SARS-COV-2-6P-MUT7 S PROTEIN 1 RBD UP CONFORMATION | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, Spike glycoprotein, ... | | Authors: | Niu, L, Chandravanshi, M, Tolbert, W.D, Pazgier, M. | | Deposit date: | 2025-04-30 | | Release date: | 2025-10-01 | | Last modified: | 2025-12-10 | | Method: | ELECTRON MICROSCOPY (3.09 Å) | | Cite: | Optimization of VE607 to generate analogs with improved neutralization activities against SARS-CoV-2 variants.
J.Virol., 99, 2025
|
|
9OG4
 
 | | SARS-COV-2-6P-MUT7 S PROTEIN-DY-III-281 complex closed conformation | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, Spike glycoprotein, ... | | Authors: | Chandravanshi, M, Niu, L, Tolbert, W.D, Pazgier, M. | | Deposit date: | 2025-04-30 | | Release date: | 2025-10-01 | | Last modified: | 2025-12-10 | | Method: | ELECTRON MICROSCOPY (3.56 Å) | | Cite: | Optimization of VE607 to generate analogs with improved neutralization activities against SARS-CoV-2 variants.
J.Virol., 99, 2025
|
|
9O5V
 
 | | Crystal structure of chimeric BANAL-52 RBD complexed with chimeric Rhinolophus sinicus ACE2 | | Descriptor: | 1,2-ETHANEDIOL, 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Hsueh, F.-C, Shi, K, Aihara, H, Li, F. | | Deposit date: | 2025-04-10 | | Release date: | 2026-01-28 | | Last modified: | 2026-02-18 | | Method: | X-RAY DIFFRACTION (3.35 Å) | | Cite: | Crystal structure of chimeric BANAL-52 RBD complexed with chimeric Rhinolophus sinicus ACE2
To Be Published
|
|
9O5T
 
 | | Crystal structure of chimeric SARS-CoV-2 RBD complexed with chimeric Rhinolophus sinicus ACE2 | | Descriptor: | 1,2-ETHANEDIOL, 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Hsueh, F.-C, Shi, K, Aihara, H, Li, F. | | Deposit date: | 2025-04-10 | | Release date: | 2026-01-28 | | Last modified: | 2026-02-18 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Crystal structure of chimeric SARS-CoV-2 RBD complexed with chimeric Rhinolophus sinicus ACE2
To Be Published
|
|
9PW4
 
 | | Structure of V30V4 in complex with SARS-CoV-2 spike | | Descriptor: | SP1-77 FAB HEAVY CHAIN, SP1-77 FAB LIGHT CHAIN, Spike protein S1, ... | | Authors: | Wang, Y.J, Kibria, G, Wesemann, D, Chen, B. | | Deposit date: | 2025-08-04 | | Release date: | 2025-11-19 | | Last modified: | 2026-01-14 | | Method: | ELECTRON MICROSCOPY (2.85 Å) | | Cite: | Affinity maturation and light-chain-mediated paratope diversification anticipates viral evolution.
Cell Rep, 45, 2025
|
|
9PSP
 
 | |
9T74
 
 | |
9UXD
 
 | | SARS-CoV2 Spike protein with Fab fragment antibody KXD355,state1 | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, Antibody KXD355, ... | | Authors: | Wang, H. | | Deposit date: | 2025-05-13 | | Release date: | 2025-07-16 | | Method: | ELECTRON MICROSCOPY (3.03 Å) | | Cite: | A rare B cell clonotype imprinted by ancestral SARS-CoV-2 develops cross-sarbecovirus neutralization in immune recalls.
Cell Rep, 44, 2025
|
|
9UXE
 
 | | SARS-CoV2 Spike protein with Fab fragment antibody KXD355,state2 | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, Antibody KXD355, ... | | Authors: | Wang, H. | | Deposit date: | 2025-05-13 | | Release date: | 2025-07-16 | | Method: | ELECTRON MICROSCOPY (3.17 Å) | | Cite: | A rare B cell clonotype imprinted by ancestral SARS-CoV-2 develops cross-sarbecovirus neutralization in immune recalls.
Cell Rep, 44, 2025
|
|
9PSN
 
 | |
9PSO
 
 | |
