4IYJ
 
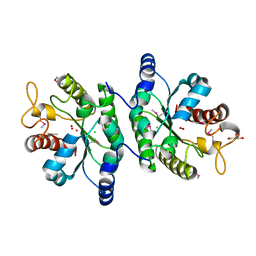 | |
4J6F
 | | Crystal structure of putative alcohol dehydrogenase from Sinorhizobium meliloti 1021, NYSGRC-Target 012230 | | Descriptor: | CHLORIDE ION, GLYCEROL, Putative alcohol dehydrogenase | | Authors: | Malashkevich, V.N, Bhosle, R, Toro, R, Hillerich, B, Gizzi, A, Garforth, S, Kar, A, Chan, M.K, Lafluer, J, Patel, H, Matikainen, B, Chamala, S, Lim, S, Celikgil, A, Villegas, G, Evans, B, Love, J, Fiser, A, Khafizov, K, Seidel, R, Bonanno, J.B, Almo, S.C, New York Structural Genomics Research Consortium (NYSGRC) | | Deposit date: | 2013-02-11 | | Release date: | 2013-03-20 | | Last modified: | 2023-12-06 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Crystal structure of putative alcohol dehydrogenase from Sinorhizobium meliloti 1021, NYSGRC-Target 012230
To be Published
|
|
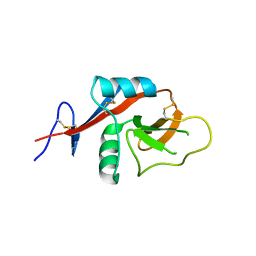
4J6K
 
 | |
4II1
 
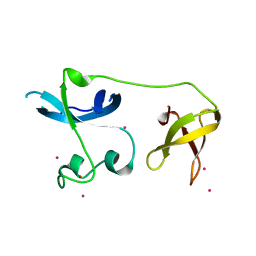 | | Crystal structure of the zinc finger of ZGPAT | | Descriptor: | UNKNOWN ATOM OR ION, ZINC ION, Zinc finger CCCH-type with G patch domain-containing protein | | Authors: | Bian, C, Tempel, W, Dong, A, Chao, X, Fu, M, Wernimont, A.K, Bountra, C, Weigelt, J, Arrowsmith, C.H, Edwards, A.M, Min, J, Structural Genomics Consortium (SGC) | | Deposit date: | 2012-12-19 | | Release date: | 2013-02-13 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.65 Å) | | Cite: | Crystal structure of the zinc finger of ZGPAT
to be published
|
|
4IIZ
 
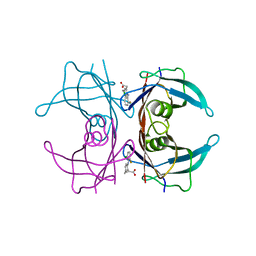 | |
3MCX
 
 | |
3MDO
 
 | |
4IKK
 
 | |
4IL3
 
 | | Crystal Structure of S. mikatae Ste24p | | Descriptor: | Ste24p, ZINC ION | | Authors: | Pryor Jr, E.E, Horanyi, P.S, Clark, K, Fedoriw, N, Connelly, S.M, Koszelak-Rosenblum, M, Zhu, G, Malkowski, M.G, Dumont, M.E, Wiener, M.C, Membrane Protein Structural Biology Consortium (MPSBC) | | Deposit date: | 2012-12-28 | | Release date: | 2013-02-20 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (3.102 Å) | | Cite: | Structure of the integral membrane protein CAAX protease Ste24p.
Science, 339, 2013
|
|
3N6Z
 
 | |
4FXV
 
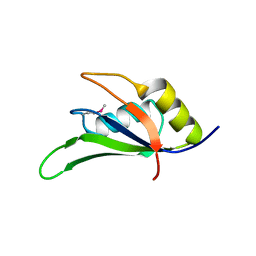 | |
4G8I
 
 | | Crystal Structure of HLA B2705-KK10-L6M | | Descriptor: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, Beta-2-microglobulin, Gag protein, ... | | Authors: | Gras, S, Wilmann, P.G, Rossjohn, J. | | Deposit date: | 2012-07-23 | | Release date: | 2013-03-20 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | A Molecular Basis for the Control of Preimmune Escape Variants by HIV-Specific CD8(+) T Cells.
Immunity, 38, 2013
|
|
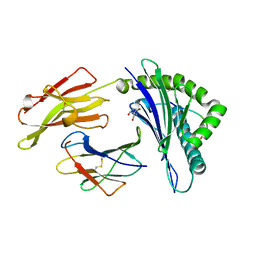
4GBN
 
 | | Crystal structure of aspart insulin at pH 6.5 | | Descriptor: | CHLORIDE ION, Insulin A chain, Insulin B chain, ... | | Authors: | Lima, L.M.T.R, Favero-Retto, M.P, Palmieri, L.C. | | Deposit date: | 2012-07-27 | | Release date: | 2013-06-12 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.872 Å) | | Cite: | A T3R3 hexamer of the human insulin variant B28Asp.
Biophys.Chem., 173, 2013
|
|
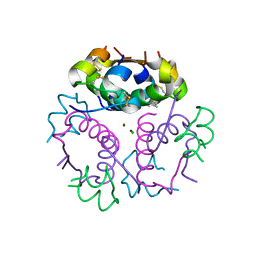
2A2E
 
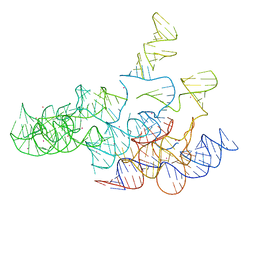 | | Crystal structure of the RNA subunit of Ribonuclease P. Bacterial A-type. | | Descriptor: | OSMIUM ION, RNA subunit of RNase P | | Authors: | Torres-Larios, A, Swinger, K.K, Krasilnikov, A.S, Pan, T, Mondragon, A. | | Deposit date: | 2005-06-22 | | Release date: | 2005-09-06 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (3.85 Å) | | Cite: | Crystal structure of the RNA component of bacterial ribonuclease P.
Nature, 437, 2005
|
|
4J6D
 
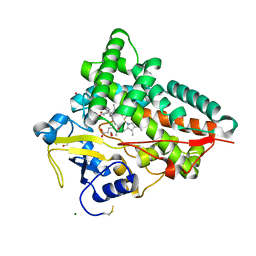 | |
4J6V
 
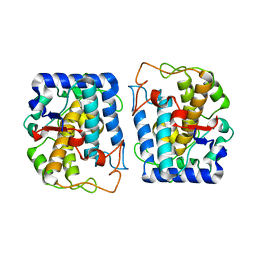 | | Crystal Structure of Tyrosinase from Bacillus megaterium N205D mutant | | Descriptor: | COPPER (II) ION, Tyrosinase | | Authors: | Kanteev, M, Goldfeder, M, Adir, N, Fishman, A. | | Deposit date: | 2013-02-12 | | Release date: | 2013-12-25 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | The mechanism of copper uptake by tyrosinase from Bacillus megaterium.
J.Biol.Inorg.Chem., 18, 2013
|
|
4J91
 
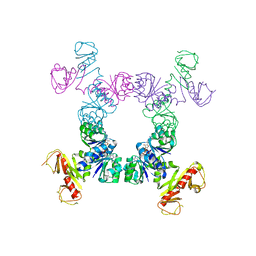 | |
4JQY
 
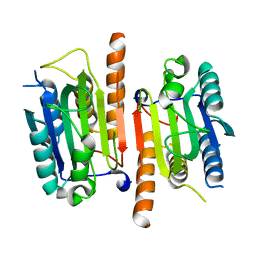 | | Human procaspase-3, crystal form 1 | | Descriptor: | Procaspase-3 | | Authors: | Thomsen, N.D, Wells, J.A. | | Deposit date: | 2013-03-20 | | Release date: | 2013-05-08 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.495 Å) | | Cite: | Structural snapshots reveal distinct mechanisms of procaspase-3 and -7 activation.
Proc.Natl.Acad.Sci.USA, 110, 2013
|
|
4J6T
 
 | | Crystal Structure of Tyrosinase from Bacillus megaterium F197A mutant | | Descriptor: | COPPER (II) ION, Tyrosinase | | Authors: | Kanteev, M, Goldfeder, M, Adir, N, Fishman, A. | | Deposit date: | 2013-02-12 | | Release date: | 2013-12-25 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.43 Å) | | Cite: | The mechanism of copper uptake by tyrosinase from Bacillus megaterium.
J.Biol.Inorg.Chem., 18, 2013
|
|
4J6B
 
 | |
4J7C
 
 | |
4JGL
 
 | |
4IVZ
 
 | |
2R61
 
 | |
2R7Q
 
 | | Crystal Structure of VP1 apoenzyme of Rotavirus SA11 (C-terminal hexahistidine-tagged) | | Descriptor: | RNA-dependent RNA polymerase | | Authors: | Lu, X, Harrison, S.C, Tao, Y.J, Patton, J.T, Nibert, M.L. | | Deposit date: | 2007-09-09 | | Release date: | 2008-07-29 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Mechanism for coordinated RNA packaging and genome replication by rotavirus polymerase VP1.
Structure, 16, 2008
|
|
