9ERT
 
 | | Mouse CNPase catalytic domain with nano body 5E | | Descriptor: | 2',3'-cyclic-nucleotide 3'-phosphodiesterase, Chains: B, PHOSPHATE ION | | Authors: | Markusson, S, Raasakka, A, Opazo, F, Kursula, P. | | Deposit date: | 2024-03-25 | | Release date: | 2024-12-11 | | Method: | X-RAY DIFFRACTION (2.75 Å) | | Cite: | Mouse CNPase catalytic domain with nano body 5E
To Be Published
|
|
9ERU
 
 | | Mouse CNPase catalytic domain with nanobody 7E | | Descriptor: | 2',3'-cyclic-nucleotide 3'-phosphodiesterase, Chains: G,C,E,H | | Authors: | Markusson, S, Raasakka, A, Opazo, F, Kursula, P. | | Deposit date: | 2024-03-25 | | Release date: | 2024-12-25 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Mouse CNPase catalytic domain with nanobody 7E
To Be Published
|
|
9ERV
 
 | | Structure of Salmonella CapRel bound to Bas11 Gp54 | | Descriptor: | ADENOSINE-5'-TRIPHOSPHATE, GLYCEROL, GP54, ... | | Authors: | Garcia-Pino, A. | | Deposit date: | 2024-03-25 | | Release date: | 2024-11-06 | | Last modified: | 2024-12-04 | | Method: | X-RAY DIFFRACTION (2.253 Å) | | Cite: | A bacterial immunity protein directly senses two disparate phage proteins.
Nature, 635, 2024
|
|
9ERW
 
 | |
9ERX
 
 | | Structural basis of D9-THC analog activity at the Cannabinoid 1 receptor | | Descriptor: | (6aR,10aR)-9-(hydroxymethyl)-6,6-dimethyl-3-(2-methyloctan-2-yl)-6a,7,10,10a-tetrahydrobenzo[c]chromen-1-ol, Antibody ScFv16 Fab fragment, Cannabinoid receptor 1, ... | | Authors: | Thorsen, T.S, Kulkarni, Y, Boggild, A, Drace, T, Nissen, P, Gajhede, M, Boesen, T, Kastrup, J.S, Gloriam, D. | | Deposit date: | 2024-03-25 | | Release date: | 2024-06-26 | | Last modified: | 2025-07-02 | | Method: | ELECTRON MICROSCOPY (2.9 Å) | | Cite: | Structural basis of Delta 9 -THC analog activity at the Cannabinoid 1 receptor.
Res Sq, 2024
|
|
9ERY
 
 | | Co-crystal strucutre of PD-L1 with low molecular weight inhibitor | | Descriptor: | 5-[[5-[[2-[bis(fluoranyl)methyl]-3-(2,3-dihydro-1,4-benzodioxin-6-yl)phenyl]methoxy]-2-[(2-hydroxyethylamino)methyl]phenoxy]methyl]pyridine-3-carbonitrile, Programmed cell death 1 ligand 1, SULFATE ION | | Authors: | Plewka, J, Magiera-Mularz, K, Zhang, W. | | Deposit date: | 2024-03-25 | | Release date: | 2024-07-24 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Design, synthesis, and evaluation of antitumor activity of 2-arylmethoxy-4-(2-fluoromethyl-biphenyl-3-ylmethoxy) benzylamine derivatives as PD-1/PD-l1 inhibitors.
Eur.J.Med.Chem., 276, 2024
|
|
9ERZ
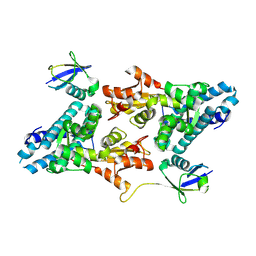 
 | | Structure of CBL-TKBD bound to Ubiquitin-fused CBLock peptide | | Descriptor: | CALCIUM ION, E3 ubiquitin-protein ligase CBL, Polyubiquitin-C,Ub-fused CBLock peptide | | Authors: | Ahmed, S.F, Huang, D.T. | | Deposit date: | 2024-03-25 | | Release date: | 2025-05-14 | | Last modified: | 2025-08-20 | | Method: | X-RAY DIFFRACTION (2.02 Å) | | Cite: | Locking CBL TKBD in its native conformation presents a novel therapeutic opportunity in mutant CBL-dependent leukemia.
Mol.Ther., 33, 2025
|
|
9ES0
 
 | |
9ES1
 | |
9ES2
 
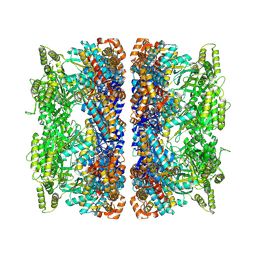 | |
9ES3
 | |
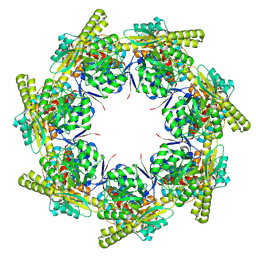
9ES4
 
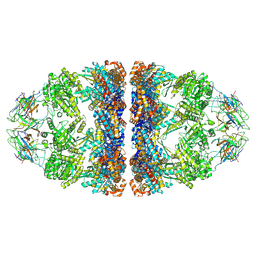 | | ADP:BeF3-bound human mitochondrial Hsp60-Hsp10 football complex | | Descriptor: | 10 kDa heat shock protein, mitochondrial, 60 kDa heat shock protein, ... | | Authors: | Lopez-Alonso, J.P, Tascon, I, Ubarretxena-Belandia, I, Shkolnisky, Y. | | Deposit date: | 2024-03-25 | | Release date: | 2025-04-09 | | Last modified: | 2025-12-10 | | Method: | ELECTRON MICROSCOPY (2.91 Å) | | Cite: | Structural basis for ATP-driven double-ring assembly of the human mitochondrial Hsp60 chaperonin.
Biorxiv, 2025
|
|
9ES5
 
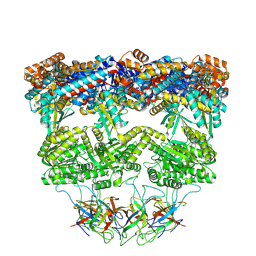 | | ADP:BeF3-bound human mitochondrial Hsp60-Hsp10 half-football complex | | Descriptor: | 10 kDa heat shock protein, mitochondrial, 60 kDa heat shock protein, ... | | Authors: | Lopez-Alonso, J.P, Tascon, I, Ubarretxena-Belandia, I, Shkolnisky, Y. | | Deposit date: | 2024-03-25 | | Release date: | 2025-04-09 | | Last modified: | 2025-12-10 | | Method: | ELECTRON MICROSCOPY (3.5 Å) | | Cite: | Structural basis for ATP-driven double-ring assembly of the human mitochondrial Hsp60 chaperonin.
Biorxiv, 2025
|
|
9ES6
 
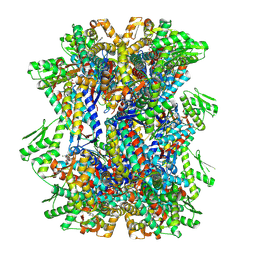 | | ADP:BeF3-bound human mitochondrial Hsp60 double-ring complex | | Descriptor: | 60 kDa heat shock protein, mitochondrial, ADENOSINE-5'-DIPHOSPHATE, ... | | Authors: | Lopez-Alonso, J.P, Tascon, I, Ubarretxena-Belandia, I, Shkolnisky, Y. | | Deposit date: | 2024-03-25 | | Release date: | 2025-04-09 | | Last modified: | 2025-12-10 | | Method: | ELECTRON MICROSCOPY (3 Å) | | Cite: | Structural basis for ATP-driven double-ring assembly of the human mitochondrial Hsp60 chaperonin.
Biorxiv, 2025
|
|
9ES7
 
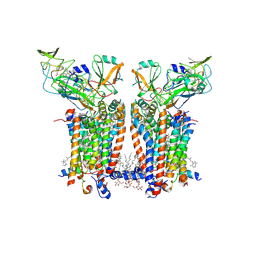 | | Cryo-EM structure of Spinacia oleracea cytochrome b6f complex with water molecules at 1.94 A resolution | | Descriptor: | 1,2-DI-O-ACYL-3-O-[6-DEOXY-6-SULFO-ALPHA-D-GLUCOPYRANOSYL]-SN-GLYCEROL, 1,2-DISTEAROYL-MONOGALACTOSYL-DIGLYCERIDE, BETA-CAROTENE, ... | | Authors: | Pietras, R, Pintscher, S, Mielecki, B, Szwalec, M, Wojcik-Augustyn, A, Indyka, P, Rawski, M, Koziej, L, Jaciuk, M, Wazny, G, Glatt, S, Osyczka, A. | | Deposit date: | 2024-03-25 | | Release date: | 2024-10-16 | | Last modified: | 2024-11-27 | | Method: | ELECTRON MICROSCOPY (1.94 Å) | | Cite: | Molecular basis of plastoquinone reduction in plant cytochrome b 6 f.
Nat.Plants, 10, 2024
|
|
9ES8
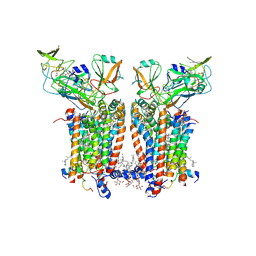 
 | | Cryo-EM structure of Spinacia oleracea cytochrome b6f with decylplastoquinone bound at plastoquionol reduction site | | Descriptor: | 1,2-DI-O-ACYL-3-O-[6-DEOXY-6-SULFO-ALPHA-D-GLUCOPYRANOSYL]-SN-GLYCEROL, 1,2-DISTEAROYL-MONOGALACTOSYL-DIGLYCERIDE, BETA-CAROTENE, ... | | Authors: | Pietras, R, Pintscher, S, Mielecki, B, Szwalec, M, Wojcik-Augustyn, A, Indyka, P, Rawski, M, Koziej, L, Jaciuk, M, Wazny, G, Glatt, S, Osyczka, A. | | Deposit date: | 2024-03-25 | | Release date: | 2024-10-16 | | Last modified: | 2024-11-27 | | Method: | ELECTRON MICROSCOPY (2.24 Å) | | Cite: | Molecular basis of plastoquinone reduction in plant cytochrome b 6 f.
Nat.Plants, 10, 2024
|
|
9ES9
 
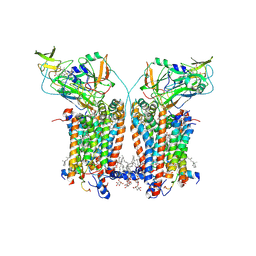 | | Cryo-EM structure of Spinacia oleracea cytochrome b6f complex with inhibitor DBMIB bound at plastoquinol oxidation site | | Descriptor: | 1,2-DI-O-ACYL-3-O-[6-DEOXY-6-SULFO-ALPHA-D-GLUCOPYRANOSYL]-SN-GLYCEROL, 1,2-DISTEAROYL-MONOGALACTOSYL-DIGLYCERIDE, 2,5-DIBROMO-3-ISOPROPYL-6-METHYLBENZO-1,4-QUINONE, ... | | Authors: | Pietras, R, Pintscher, S, Mielecki, B, Szwalec, M, Wojcik-Augustyn, A, Indyka, P, Rawski, M, Koziej, L, Jaciuk, M, Wazny, G, Glatt, S, Osyczka, A. | | Deposit date: | 2024-03-25 | | Release date: | 2024-10-16 | | Last modified: | 2024-11-27 | | Method: | ELECTRON MICROSCOPY (2.33 Å) | | Cite: | Molecular basis of plastoquinone reduction in plant cytochrome b 6 f.
Nat.Plants, 10, 2024
|
|
9ESA
 
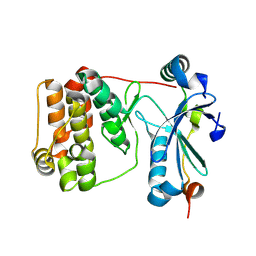 | | Aurora-C with SER mutation in complex with INCENP peptide | | Descriptor: | 1,2-ETHANEDIOL, Aurora kinase C, Inner centromere protein | | Authors: | Hillig, R.C. | | Deposit date: | 2024-03-26 | | Release date: | 2024-09-11 | | Last modified: | 2024-09-25 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Surface-mutagenesis strategies to enable structural biology crystallization platforms.
Acta Crystallogr D Struct Biol, 80, 2024
|
|
9ESB
 
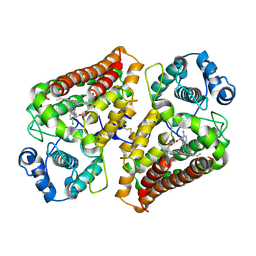 | | Holo IDO with a bound inhibitor | | Descriptor: | (~{R})-(6-chloranylimidazo[1,5-a]pyridin-5-yl)-(1-phenyl-1,2,3-triazol-4-yl)methanol, Indoleamine 2,3-dioxygenase 1, PROTOPORPHYRIN IX CONTAINING FE | | Authors: | Wicki, M, Mac Sweeney, A. | | Deposit date: | 2024-03-26 | | Release date: | 2025-03-05 | | Method: | X-RAY DIFFRACTION (2.249 Å) | | Cite: | SAR and cellular potency optimization of novel heme-binding IDO1 inhibitors
To Be Published
|
|
9ESC
 
 | | Holo IDO with a bound inhibitor | | Descriptor: | (~{S})-[(7~{a}~{R})-2-cyclopentyl-5,6,7,7~{a}-tetrahydroimidazo[5,1-b][1,3]thiazol-3-yl]-cyclohexyl-methanol, Indoleamine 2,3-dioxygenase 1, PROTOPORPHYRIN IX CONTAINING FE | | Authors: | Wicki, M, Mac Sweeney, A. | | Deposit date: | 2024-03-26 | | Release date: | 2025-03-05 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | SAR and cellular potency optimization of novel heme-binding IDO1 inhibitors
To Be Published
|
|
9ESD
 
 | | Holo TDO with a bound inhibitor | | Descriptor: | (1~{R})-1-cyclohexyl-2-[(5~{S})-5~{H}-imidazo[1,5-b]isoindol-5-yl]ethanol, PROTOPORPHYRIN IX CONTAINING FE, Tryptophan 2,3-dioxygenase | | Authors: | Wicki, M, Mac Sweeney, A. | | Deposit date: | 2024-03-26 | | Release date: | 2025-03-05 | | Method: | X-RAY DIFFRACTION (2.103 Å) | | Cite: | SAR and cellular potency optimization of novel heme-binding IDO1 inhibitors
To Be Published
|
|
9ESE
 
 | | Holo IDO with a bound inhibitor | | Descriptor: | (~{R})-(2-cyclopropylimidazo[5,1-b][1,3]thiazol-3-yl)-[1-(1~{H}-indol-5-yl)-1,2,3-triazol-4-yl]methanol, Indoleamine 2,3-dioxygenase 1, PROTOPORPHYRIN IX CONTAINING FE | | Authors: | Wicki, M, Mac Sweeney, A. | | Deposit date: | 2024-03-26 | | Release date: | 2025-03-05 | | Method: | X-RAY DIFFRACTION (2.539 Å) | | Cite: | SAR and cellular potency optimization of novel heme-binding IDO1 inhibitors
To Be Published
|
|
9ESF
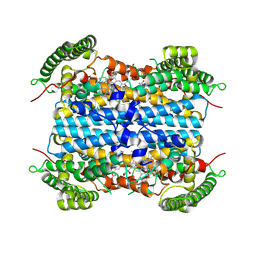 
 | | Holo IDO with a bound inhibitor | | Descriptor: | (~{S})-cyclohexyl(imidazo[1,5-a]pyridin-5-yl)methanol, PROTOPORPHYRIN IX CONTAINING FE, Tryptophan 2,3-dioxygenase | | Authors: | Wicki, M, Mac Sweeney, A. | | Deposit date: | 2024-03-26 | | Release date: | 2025-03-05 | | Method: | X-RAY DIFFRACTION (2.501 Å) | | Cite: | SAR and cellular potency optimization of novel heme-binding IDO1 inhibitors
To Be Published
|
|
9ESG
 
 | | Holo IDO with a bound inhibitor | | Descriptor: | (~{R})-(2-cyclopropylimidazo[5,1-b][1,3]thiazol-3-yl)-(1-phenyl-1,2,3-triazol-4-yl)methanol, Indoleamine 2,3-dioxygenase 1, PROTOPORPHYRIN IX CONTAINING FE | | Authors: | Wicki, M, Mac Sweeney, A. | | Deposit date: | 2024-03-26 | | Release date: | 2025-03-05 | | Method: | X-RAY DIFFRACTION (2.497 Å) | | Cite: | SAR and cellular potency optimization of novel heme-binding IDO1 inhibitors
To Be Published
|
|
9ESH
 
 | | Structure of a B-state intermediate committed to discard (Bd-I state) | | Descriptor: | G-patch domain-containing protein C1486.03, GUANOSINE-5'-TRIPHOSPHATE, INOSITOL HEXAKISPHOSPHATE, ... | | Authors: | Soni, K, Wild, K, Sinning, I. | | Deposit date: | 2024-03-26 | | Release date: | 2024-12-25 | | Last modified: | 2025-05-28 | | Method: | ELECTRON MICROSCOPY (3.2 Å) | | Cite: | Structures of aberrant spliceosome intermediates on their way to disassembly.
Nat.Struct.Mol.Biol., 32, 2025
|
|
