4RJF
 
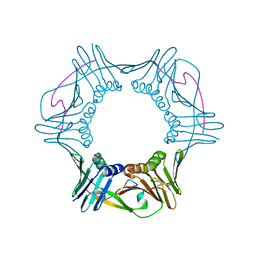 | |
4IQU
 
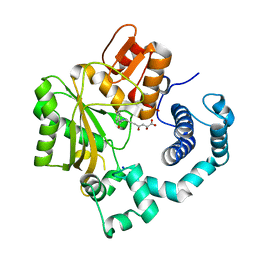 | | Tdt core in complex with inhibitor (2Z,5E)-6-[4-(4-fluorobenzoyl)-1H-pyrrol-2-yl]-2-hydroxy-4-oxohexa-2,5-dienoic acid | | Descriptor: | (2Z,5E)-6-[4-(4-fluorobenzoyl)-1H-pyrrol-2-yl]-2-hydroxy-4-oxohexa-2,5-dienoic acid, DNA nucleotidylexotransferase, SODIUM ION | | Authors: | Gouge, J, Delarue, M. | | Deposit date: | 2013-01-13 | | Release date: | 2013-09-04 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | New nucleotide-competitive non-nucleoside inhibitors of terminal deoxynucleotidyl transferase: discovery, characterization, and crystal structure in complex with the target.
J.Med.Chem., 56, 2013
|
|
8J4T
 
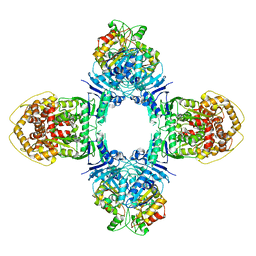 | | GajA-GajB complex | | Descriptor: | Endonuclease GajA, Gabija protein GajB | | Authors: | Wei, T, Ma, J, Huo, Y. | | Deposit date: | 2023-04-20 | | Release date: | 2024-02-21 | | Method: | ELECTRON MICROSCOPY (3.6 Å) | | Cite: | Structural and biochemical insights into the mechanism of the Gabija bacterial immunity system.
Nat Commun, 15, 2024
|
|
7QNQ
 
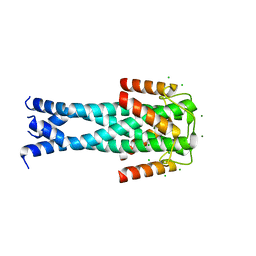 | | Structure of the Aux2 relaxosome protein of plasmid pLS20 | | Descriptor: | Auxiliary relaxosome protein, CHLORIDE ION, MAGNESIUM ION | | Authors: | Boer, D.R, Crespo, I. | | Deposit date: | 2021-12-22 | | Release date: | 2022-03-09 | | Last modified: | 2024-06-19 | | Method: | X-RAY DIFFRACTION (1.89 Å) | | Cite: | Structural and biochemical characterization of the relaxosome auxiliary proteins encoded on the Bacillus subtilis plasmid pLS20.
Comput Struct Biotechnol J, 20, 2022
|
|
8DSS
 
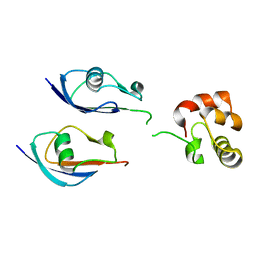 | |
7CCG
 
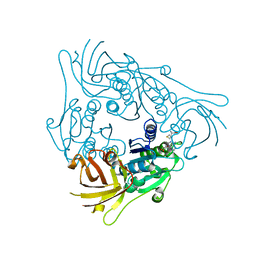 | |
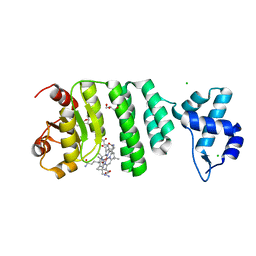
5C8F
 
 | |
5ZOD
 
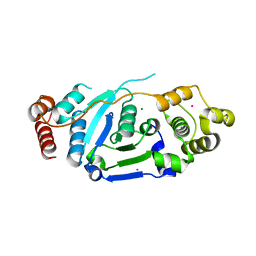 | | Crystal Structure of hFen1 in apo form | | Descriptor: | Flap endonuclease 1, MAGNESIUM ION, POTASSIUM ION | | Authors: | Han, W, Hua, Y, Zhao, Y. | | Deposit date: | 2018-04-13 | | Release date: | 2019-01-30 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Structural basis of 5' flap recognition and protein-protein interactions of human flap endonuclease 1.
Nucleic Acids Res., 46, 2018
|
|
7DG2
 
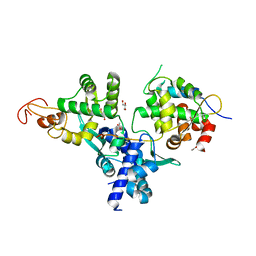 | | Nse1-Nse3-Nse4 complex | | Descriptor: | ACETATE ION, GLYCEROL, MAGE domain-containing protein, ... | | Authors: | Cho, Y, Jo, A. | | Deposit date: | 2020-11-10 | | Release date: | 2021-05-26 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Structure Basis for Shaping the Nse4 Protein by the Nse1 and Nse3 Dimer within the Smc5/6 Complex.
J.Mol.Biol., 433, 2021
|
|
1ES8
 
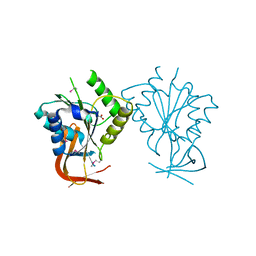 | | Crystal structure of free BglII | | Descriptor: | ACETIC ACID, RESTRICTION ENDONUCLEASE BGLII | | Authors: | Lukacs, C.M, Aggarwal, A.K. | | Deposit date: | 2000-04-07 | | Release date: | 2001-01-17 | | Last modified: | 2012-05-09 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Structure of free BglII reveals an unprecedented scissor-like motion for opening an endonuclease.
Nat.Struct.Biol., 8, 2001
|
|
1EO3
 
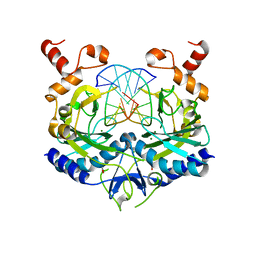 | | INHIBITION OF ECORV ENDONUCLEASE BY DEOXYRIBO-3'-S-PHOSPHOROTHIOLATES: A HIGH RESOLUTION X-RAY CRYSTALLOGRAPHIC STUDY | | Descriptor: | ACETIC ACID, DNA (5'-D(*CP*AP*AP*GP*AP*(TSP)P*AP*TP*CP*TP*T)-3'), MAGNESIUM ION, ... | | Authors: | Horton, N.C, Connolly, B.A, Perona, J.J. | | Deposit date: | 2000-03-21 | | Release date: | 2000-04-04 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Inhibition of EcoRV Endonuclease by Deoxyribo-3'-S-phosphorothiolates: A High-Resolution X-ray Crystallographic Study
J.Am.Chem.Soc., 122, 2000
|
|
5XJW
 
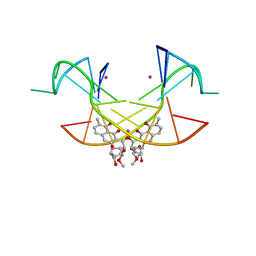 | | Crystal Structure of the [Co2+-(Chromomycin A3)2]-CCG repeats Complex | | Descriptor: | (1S)-5-deoxy-1-O-methyl-1-C-[(2R,3S)-3,5,7,10-tetrahydroxy-6-methyl-4-oxo-1,2,3,4-tetrahydroanthracen-2-yl]-D-xylulose, 2,6-dideoxy-4-O-methyl-alpha-D-galactopyranose-(1-3)-4-O-acetyl-2,6-dideoxy-beta-D-galactopyranose, 3-C-methyl-4-O-acetyl-alpha-L-Olivopyranose-(1-3)-beta-D-Olivopyranose-(1-3)-beta-D-Olivopyranose, ... | | Authors: | Tseng, W.H, Wu, P.C, Satange, R.B, Hou, M.H. | | Deposit date: | 2017-05-04 | | Release date: | 2018-05-16 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.097 Å) | | Cite: | Crystal Structure of the [Co2+-(Chromomycin A3)2]-CCG repeats Complex
To be published
|
|
1P4A
 
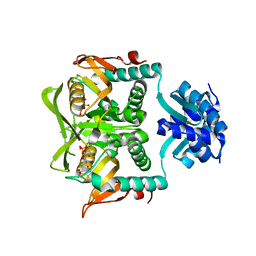 | | Crystal Structure of the PurR complexed with cPRPP | | Descriptor: | 1-ALPHA-PYROPHOSPHORYL-2-ALPHA,3-ALPHA-DIHYDROXY-4-BETA-CYCLOPENTANE-METHANOL-5-PHOSPHATE, Pur operon repressor | | Authors: | Bera, A.K, Zhu, J, Zalkin, H, Smith, J.L. | | Deposit date: | 2003-04-22 | | Release date: | 2003-12-09 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.22 Å) | | Cite: | Functional dissection of the Bacillus subtilis pur operator site.
J.Bacteriol., 185, 2003
|
|
3ERC
 
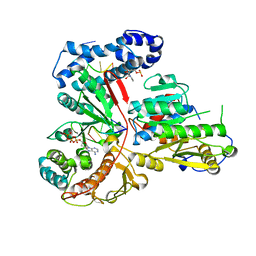 | | Crystal structure of the heterodimeric vaccinia virus mRNA polyadenylate polymerase with three fragments of RNA and 3'-deoxy ATP | | Descriptor: | 3'-DEOXYADENOSINE-5'-TRIPHOSPHATE, CALCIUM ION, Cap-specific mRNA (nucleoside-2'-O-)-methyltransferase, ... | | Authors: | Li, C, Li, H, Zhou, S, Poulos, T.L, Gershon, P.D. | | Deposit date: | 2008-10-01 | | Release date: | 2009-06-16 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (3.21 Å) | | Cite: | Polymerase Translocation with Respect to Single-Stranded Nucleic Acid: Looping or Wrapping of Primer around a Poly(A) Polymerase
Structure, 17, 2009
|
|
7SH1
 
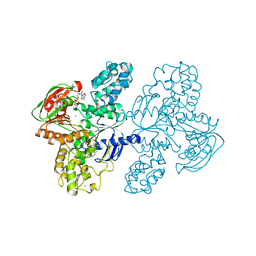 | | Class II UvrA protein - Ecm16 | | Descriptor: | ADENOSINE-5'-DIPHOSPHATE, CHLORIDE ION, Excinuclease ABC subunit UvrA, ... | | Authors: | Grade, P, Erlandson, A, Ullah, A, Mathews, I.I, Chen, X, Kim, C.-Y, Mera, P.E. | | Deposit date: | 2021-10-07 | | Release date: | 2022-10-26 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.04 Å) | | Cite: | Structural and functional analyses of the echinomycin resistance conferring protein Ecm16 from Streptomyces lasalocidi.
Sci Rep, 13, 2023
|
|
1TDH
 
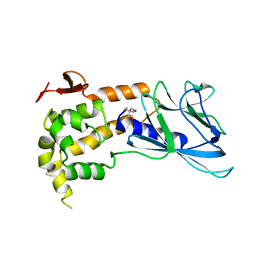 | | Crystal structure of human endonuclease VIII-like 1 (NEIL1) | | Descriptor: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, nei endonuclease VIII-like 1 | | Authors: | Doublie, S, Bandaru, V, Bond, J.P, Wallace, S.S. | | Deposit date: | 2004-05-22 | | Release date: | 2004-07-20 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | The crystal structure of human endonuclease VIII-like 1 (NEIL1) reveals a zincless finger motif required for glycosylase activity.
Proc.Natl.Acad.Sci.USA, 101, 2004
|
|
5TUP
 
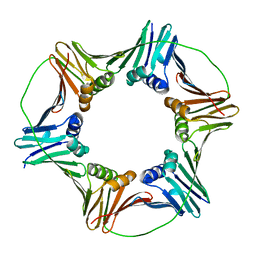 | | X-ray Crystal Structure of the Aspergillus fumigatus Sliding Clamp | | Descriptor: | Proliferating cell nuclear antigen | | Authors: | Bruning, J.B, Marshall, A.C, Kroker, A.J, Wegener, K.L, Rajapaksha, H. | | Deposit date: | 2016-11-06 | | Release date: | 2017-02-22 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (2.602 Å) | | Cite: | Structure of the sliding clamp from the fungal pathogen Aspergillus fumigatus (AfumPCNA) and interactions with Human p21.
FEBS J., 284, 2017
|
|
7BXT
 
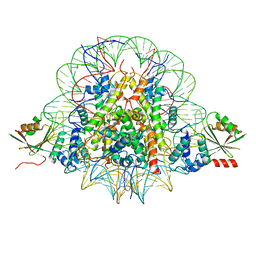 | | The cryo-EM structure of CENP-A nucleosome in complex with CENP-C peptide and CENP-N N-terminal domain | | Descriptor: | CENP-C, DNA (145-mer), Histone H2A type 1-B/E, ... | | Authors: | Ariyoshi, M, Makino, F, Fukagawa, T. | | Deposit date: | 2020-04-20 | | Release date: | 2021-02-10 | | Last modified: | 2024-03-27 | | Method: | ELECTRON MICROSCOPY (4.2 Å) | | Cite: | Cryo-EM structure of the CENP-A nucleosome in complex with phosphorylated CENP-C.
Embo J., 40, 2021
|
|
5WPA
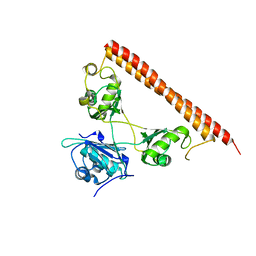 
 | | Structure of human SFPQ/PSPC1 heterodimer | | Descriptor: | Paraspeckle component 1, Splicing factor, proline- and glutamine-rich | | Authors: | Lee, M. | | Deposit date: | 2017-08-04 | | Release date: | 2018-03-21 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (2.29 Å) | | Cite: | Crystal structure of a SFPQ/PSPC1 heterodimer provides insights into preferential heterodimerization of human DBHS family proteins.
J. Biol. Chem., 293, 2018
|
|
4DRB
 
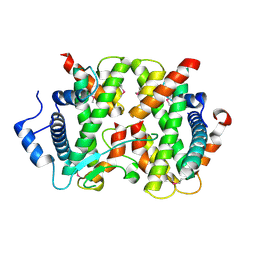 | | The crystal structure of FANCM bound MHF complex | | Descriptor: | Centromere protein S, Centromere protein X, Fanconi anemia group M protein | | Authors: | Tao, Y, Niu, L, Teng, M. | | Deposit date: | 2012-02-17 | | Release date: | 2012-05-16 | | Method: | X-RAY DIFFRACTION (2.634 Å) | | Cite: | The structure of the FANCM-MHF complex reveals physical features for functional assembly
Nat Commun, 3, 2012
|
|
4YKE
 
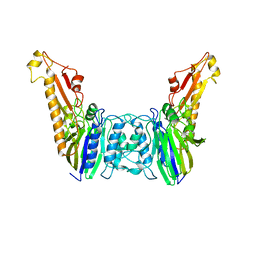 | |
7U65
 
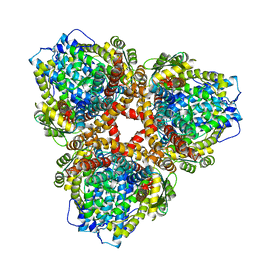 | | Structure of E. coli dGTPase bound to T7 bacteriophage protein Gp1.2 | | Descriptor: | Deoxyguanosinetriphosphate triphosphohydrolase, Inhibitor of dGTPase | | Authors: | Klemm, B.P, Hsu, A.L, Borgnia, M.J, Schaaper, R.M. | | Deposit date: | 2022-03-03 | | Release date: | 2022-08-31 | | Last modified: | 2024-06-12 | | Method: | ELECTRON MICROSCOPY (2.8 Å) | | Cite: | Mechanism by which T7 bacteriophage protein Gp1.2 inhibits Escherichia coli dGTPase.
Proc.Natl.Acad.Sci.USA, 119, 2022
|
|
7U67
 
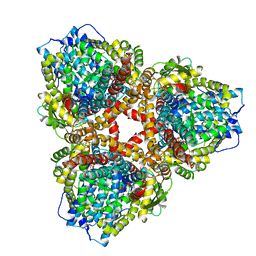 | | Structure of E. coli dGTPase bound to T7 bacteriophage protein Gp1.2 and GTP | | Descriptor: | Deoxyguanosinetriphosphate triphosphohydrolase, GUANOSINE-5'-TRIPHOSPHATE, Inhibitor of dGTPase, ... | | Authors: | Klemm, B.P, Hsu, A.L, Borgnia, M.J, Schaaper, R.M. | | Deposit date: | 2022-03-03 | | Release date: | 2022-08-31 | | Last modified: | 2024-06-12 | | Method: | ELECTRON MICROSCOPY (2.5 Å) | | Cite: | Mechanism by which T7 bacteriophage protein Gp1.2 inhibits Escherichia coli dGTPase.
Proc.Natl.Acad.Sci.USA, 119, 2022
|
|
7U66
 
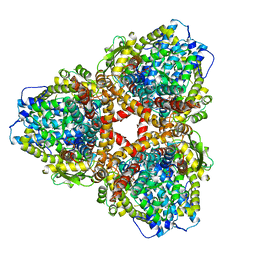 | | Structure of E. coli dGTPase bound to T7 bacteriophage protein Gp1.2 and dGTP | | Descriptor: | 2'-DEOXYGUANOSINE-5'-TRIPHOSPHATE, Deoxyguanosinetriphosphate triphosphohydrolase, Inhibitor of dGTPase, ... | | Authors: | Klemm, B.P, Dillard, L.B, Borgnia, M.J, Schaaper, R.M. | | Deposit date: | 2022-03-03 | | Release date: | 2022-08-31 | | Last modified: | 2024-06-12 | | Method: | ELECTRON MICROSCOPY (3.1 Å) | | Cite: | Mechanism by which T7 bacteriophage protein Gp1.2 inhibits Escherichia coli dGTPase.
Proc.Natl.Acad.Sci.USA, 119, 2022
|
|
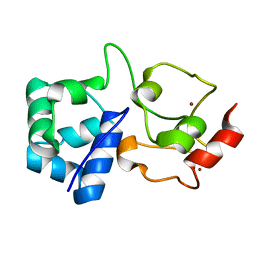
7OXD
 
 | |
