3R9Y
 
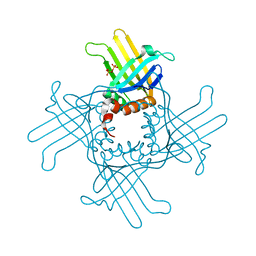 | | Crystal Structure of StWhy2 K67A (form I) | | Descriptor: | PHOSPHATE ION, Why2 protein | | Authors: | Cappadocia, L, Brisson, N, Sygusch, J. | | Deposit date: | 2011-03-26 | | Release date: | 2011-09-28 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (2.35 Å) | | Cite: | A conserved lysine residue of plant Whirly proteins is necessary for higher order protein assembly and protection against DNA damage.
Nucleic Acids Res., 40, 2012
|
|
3KHB
 
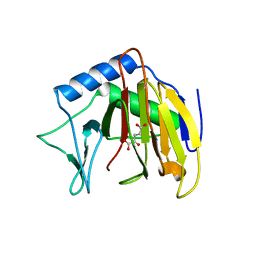 | | Crystal structure of Escherichia coli AlkB with Co(II) and 2-OG | | Descriptor: | 2-OXOGLUTARIC ACID, Alpha-ketoglutarate-dependent dioxygenase alkB, COBALT (II) ION | | Authors: | Hollis, T, Holland, P.J. | | Deposit date: | 2009-10-30 | | Release date: | 2010-01-12 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Structural and mutational analysis of Escherichia coli AlkB provides insight into substrate specificity and DNA damage searching.
Plos One, 5, 2010
|
|
3PJR
 
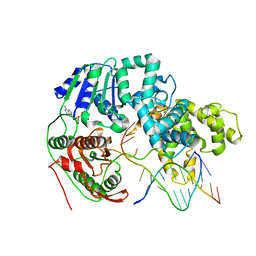 | | HELICASE SUBSTRATE COMPLEX | | Descriptor: | 5'-D(*CP*GP*AP*GP*CP*AP*CP*TP*GP*C)-3', 5'-D(*GP*CP*AP*GP*TP*GP*CP*TP*CP*GP*TP*TP*TP*TP*T)-3', ADENOSINE-5'-TRIPHOSPHATE, ... | | Authors: | Velankar, S.S, Soultanas, P, Dillingham, M.S, Subramanya, H.S, Wigley, D.B. | | Deposit date: | 1999-03-12 | | Release date: | 1999-04-08 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (3.3 Å) | | Cite: | Crystal structures of complexes of PcrA DNA helicase with a DNA substrate indicate an inchworm mechanism
Cell(Cambridge,Mass.), 97, 1999
|
|
1M6G
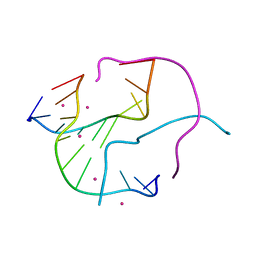 
 | | Structural Characterisation of the Holliday Junction TCGGTACCGA | | Descriptor: | 5'-D(*TP*CP*GP*GP*TP*AP*CP*CP*GP*A)-3', STRONTIUM ION | | Authors: | Thorpe, J.H, Gale, B.C, Teixeira, S.C.M, Cardin, C.J. | | Deposit date: | 2002-07-16 | | Release date: | 2003-05-06 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.652 Å) | | Cite: | Conformational and hydration effects of site-selective sodium, calcium and
strontium ion binding to the DNA Holliday junction structure
d(TCGGTACCGA)(4)
J.Mol.Biol., 327, 2003
|
|
3R4F
 
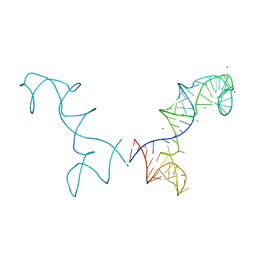 | | Prohead RNA | | Descriptor: | MAGNESIUM ION, pRNA | | Authors: | Ding, F, Lu, C, Zhano, W, Rajashankar, K.R, Anderson, D.L, Jardine, P.J, Grimes, S, Ke, A. | | Deposit date: | 2011-03-17 | | Release date: | 2011-04-20 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (3.5 Å) | | Cite: | Structure and assembly of the essential RNA ring component of a viral DNA packaging motor.
Proc.Natl.Acad.Sci.USA, 108, 2011
|
|
3U7E
 
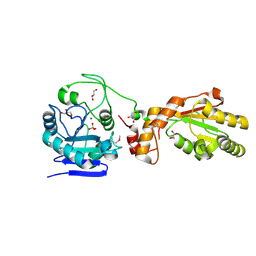 | | Crystal structure of mPNKP catalytic fragment (D170A) | | Descriptor: | Bifunctional polynucleotide phosphatase/kinase, GLYCEROL, MAGNESIUM ION, ... | | Authors: | Coquelle, N, Havali, Z, Bernstein, N, Green, R, Glover, J.N.M. | | Deposit date: | 2011-10-13 | | Release date: | 2011-12-14 | | Last modified: | 2023-12-06 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Structural basis for the phosphatase activity of polynucleotide kinase/phosphatase on single- and double-stranded DNA substrates.
Proc.Natl.Acad.Sci.USA, 108, 2011
|
|
1VDD
 
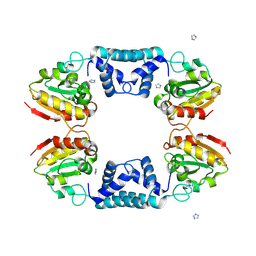 | |
3H1T
 
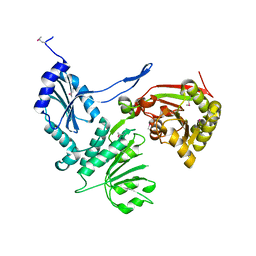 | |
3S4W
 
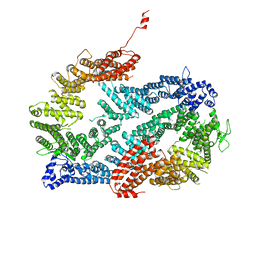 | | Structure of the FANCI-FANCD2 complex | | Descriptor: | Fanconi anemia group D2 protein homolog, Fanconi anemia group I protein homolog | | Authors: | Pavletich, N.P. | | Deposit date: | 2011-05-20 | | Release date: | 2011-07-27 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (3.408 Å) | | Cite: | Structure of the FANCI-FANCD2 complex: insights into the Fanconi anemia DNA repair pathway.
Science, 333, 2011
|
|
6J80
 
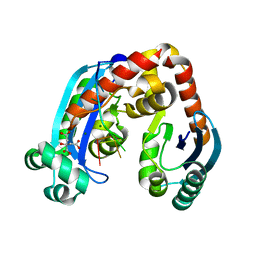 | | Human mitochondrial Oligoribonuclease in complex with poly-dT DNA | | Descriptor: | CITRIC ACID, DNA (5'-D(P*TP*TP*TP*TP*TP*TP*T)-3'), MAGNESIUM ION, ... | | Authors: | Chu, L.Y, Agrawal, S, Yuan, H.S. | | Deposit date: | 2019-01-18 | | Release date: | 2019-08-28 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (1.812 Å) | | Cite: | Structural insights into nanoRNA degradation by human Rexo2.
Rna, 25, 2019
|
|
3S51
 
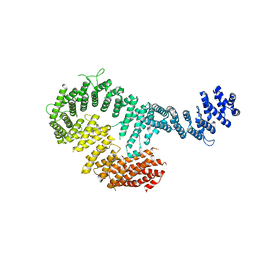 | | Structure of FANCI | | Descriptor: | Fanconi anemia group I protein homolog | | Authors: | Pavletich, N.P. | | Deposit date: | 2011-05-20 | | Release date: | 2011-07-27 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (3.3 Å) | | Cite: | Structure of the FANCI-FANCD2 complex: insights into the Fanconi anemia DNA repair pathway.
Science, 333, 2011
|
|
1K9G
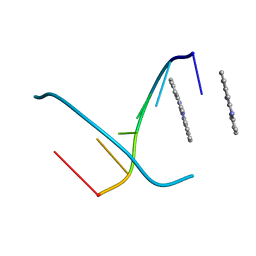 
 | | Crystal Structure of the Complex of Cryptolepine-d(CCTAGG)2 | | Descriptor: | 5'-D(*CP*CP*TP*AP*GP*G)-3', 5-METHYL-5H-INDOLO[3,2-B]QUINOLINE | | Authors: | Lisgarten, J.N, Coll, M, Portugal, J, Wright, C.W, Aymami, J. | | Deposit date: | 2001-10-29 | | Release date: | 2001-11-30 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | The antimalarial and cytotoxic drug cryptolepine intercalates into DNA at cytosine-cytosine sites.
Nat.Struct.Biol., 9, 2002
|
|
4ATH
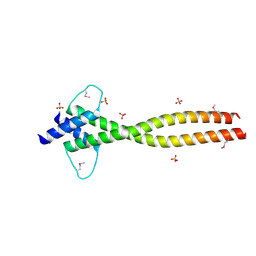 
 | | MITF apo structure | | Descriptor: | MICROPHTHALMIA-ASSOCIATED TRANSCRIPTION FACTOR, SULFATE ION | | Authors: | Pogenberg, V, Milewski, M, Wilmanns, M. | | Deposit date: | 2012-05-08 | | Release date: | 2012-12-12 | | Last modified: | 2019-07-17 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Restricted Leucine Zipper Dimerization and Specificity of DNA Recognition of the Melanocyte Master Regulator Mitf
Genes Dev., 26, 2012
|
|
7LIN
 
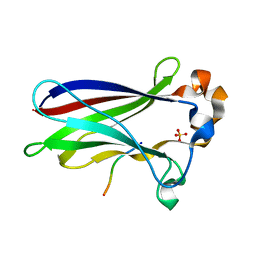 | |
7LIO
 
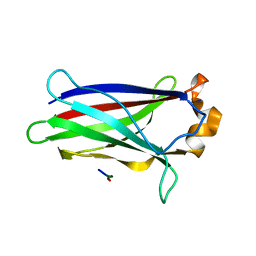 | |
7LIQ
 
 | |
7LIP
 
 | | X-ray structure of SPOP MATH domain (D140G) | | Descriptor: | SULFATE ION, Speckle-type POZ protein | | Authors: | Botuyan, M.V, Cui, G, Mer, G. | | Deposit date: | 2021-01-27 | | Release date: | 2021-04-14 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (1.48 Å) | | Cite: | ATM-phosphorylated SPOP contributes to 53BP1 exclusion from chromatin during DNA replication.
Sci Adv, 7, 2021
|
|
1UMU
 
 | |
1GE8
 
 | |
3L51
 
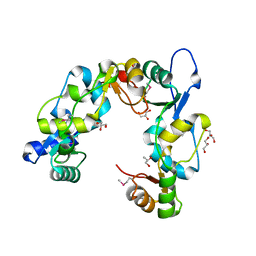 | | Crystal Structure of the Mouse Condensin Hinge Domain | | Descriptor: | GLYCEROL, Structural maintenance of chromosomes protein 2, Structural maintenance of chromosomes protein 4 | | Authors: | Griese, J.J, Hopfner, K.-P. | | Deposit date: | 2009-12-21 | | Release date: | 2010-02-16 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.506 Å) | | Cite: | Structure and DNA binding activity of the mouse condensin hinge domain highlight common and diverse features of SMC proteins
Nucleic Acids Res., 38, 2010
|
|
1V66
 
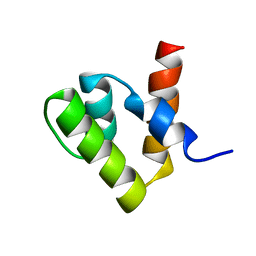 | | Solution structure of human p53 binding domain of PIAS-1 | | Descriptor: | Protein inhibitor of activated STAT protein 1 | | Authors: | Okubo, S, Hara, F, Tsuchida, Y, Shimotakahara, S, Suzuki, S, Hatanaka, H, Yokoyama, S, Tanaka, H, Yasuda, H, Shindo, H, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2003-11-27 | | Release date: | 2004-12-07 | | Last modified: | 2023-12-27 | | Method: | SOLUTION NMR | | Cite: | NMR structure of the N-terminal domain of SUMO ligase PIAS1 and its interaction with tumor suppressor p53 and A/T-rich DNA oligomers
J.Biol.Chem., 279, 2004
|
|
2LUN
 
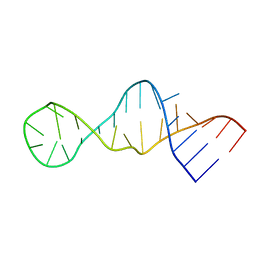 | |
5IU5
 
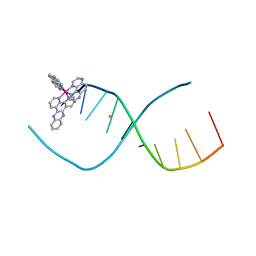 | | Lambda-[Ru(TAP)2(dppz)]2+ bound to d(TCGGCICCGA)2 | | Descriptor: | BARIUM ION, DNA (5'-D(*TP*CP*GP*GP*CP*IP*CP*CP*GP*A)-3'), Ru(tap)2(dppz) complex | | Authors: | Hall, J.P, Cardin, C.J. | | Deposit date: | 2016-03-17 | | Release date: | 2017-06-21 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Unexpected enhancement of sensitised photo-oxidation by a Ru(II) complex on replacement of guanine with inosine in DNA
To Be Published
|
|
6V3X
 
 | |
6V3Y
 
 | |
