8P1Z
 
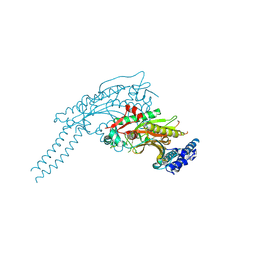 | |
8FZR
 
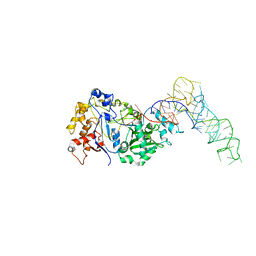 | |
4QEI
 
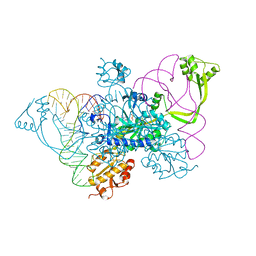 | | Two distinct conformational states of GlyRS captured in crystal lattice | | Descriptor: | ADENOSINE MONOPHOSPHATE, Glycine--tRNA ligase, tRNA-Gly-CCC-2-2 | | Authors: | Xie, W, Qin, X, Deng, X, Zhang, Q, Li, Q. | | Deposit date: | 2014-05-16 | | Release date: | 2015-05-20 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.875 Å) | | Cite: | Large Conformational Changes of Insertion 3 in Human Glycyl-tRNA Synthetase (hGlyRS) during Catalysis
J.Biol.Chem., 291, 2016
|
|
1F7G
 
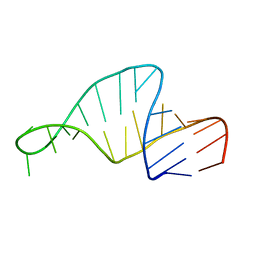 | |
1F6X
 
 | |
1F6Z
 
 | |
1F7F
 
 | |
6G6S
 
 | | Crystal structure of human Acinus RNA recognition motif domain | | Descriptor: | Apoptotic chromatin condensation inducer in the nucleus | | Authors: | Fernandes, H, Czapinska, H, Grudziaz, K, Bujnicki, J.M, Nowacka, M. | | Deposit date: | 2018-04-03 | | Release date: | 2018-06-27 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | Crystal structure of human Acinus RNA recognition motif domain.
PeerJ, 6, 2018
|
|
5H4V
 
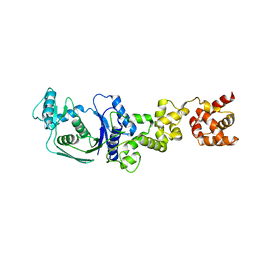 | |
2GJE
 
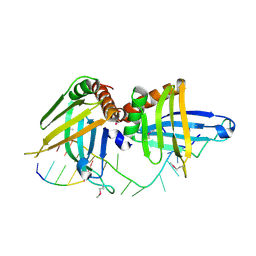 | | Structure of a guideRNA-binding protein complex bound to a gRNA | | Descriptor: | RNA tetramer, guide RNA 40-mer, mitochondrial RNA-binding protein 1, ... | | Authors: | Schumacher, M.A, Karamooz, E, Zikova, A, Trantirek, L, Lukes, J. | | Deposit date: | 2006-03-30 | | Release date: | 2006-09-05 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (3.37 Å) | | Cite: | Crystal Structures of T. brucei MRP1/MRP2 Guide-RNA Binding Complex Reveal RNA Matchmaking Mechanism.
Cell(Cambridge,Mass.), 126, 2006
|
|
4ZAJ
 
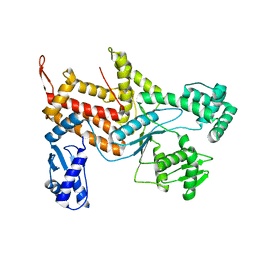 | |
5ZDK
 
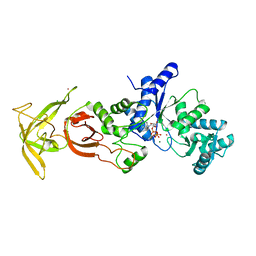 | | Crystal Structure Analysis of TtQRS in complex with ATP | | Descriptor: | ADENOSINE-5'-TRIPHOSPHATE, CESIUM ION, CHLORIDE ION, ... | | Authors: | Mutharasappan, N, Jain, V, Sharma, A, Manickam, Y, Jeyaraman, J. | | Deposit date: | 2018-02-23 | | Release date: | 2018-12-05 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.45 Å) | | Cite: | Structural and functional analysis of Glutaminyl-tRNA synthetase (TtGlnRS) from Thermus thermophilus HB8 and its complexes
Int. J. Biol. Macromol., 120, 2018
|
|
5ZDL
 
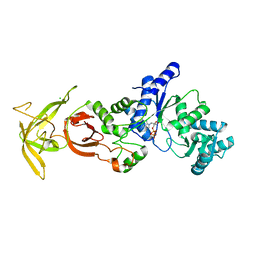 | | Crystal Structure Analysis of TtQRS in co-crystallised with ATP | | Descriptor: | ADENOSINE-5'-TRIPHOSPHATE, CHLORIDE ION, Glutamine--tRNA ligase | | Authors: | Mutharasappan, N, Jain, V, Sharma, A, Manickam, Y, Jeyaraman, J. | | Deposit date: | 2018-02-23 | | Release date: | 2018-12-05 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Structural and functional analysis of Glutaminyl-tRNA synthetase (TtGlnRS) from Thermus thermophilus HB8 and its complexes
Int. J. Biol. Macromol., 120, 2018
|
|
5ZDO
 
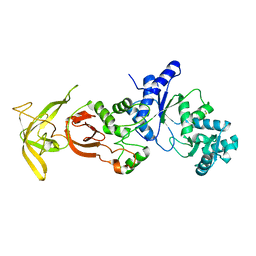 | | Crystal Structure Analysis of TtQRS in co-crystallised with ATP | | Descriptor: | CHLORIDE ION, Glutamine-tRNA ligase | | Authors: | Mutharasappan, N, Jain, V, Sharma, A, Manickam, Y, Jeyaraman, J. | | Deposit date: | 2018-02-23 | | Release date: | 2018-12-05 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Structural and functional analysis of Glutaminyl-tRNA synthetase (TtGlnRS) from Thermus thermophilus HB8 and its complexes
Int. J. Biol. Macromol., 120, 2018
|
|
2LHN
 
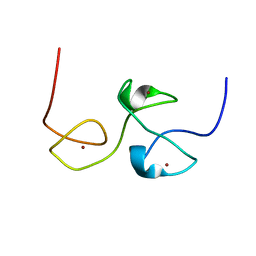 | | RNA-binding zinc finger protein | | Descriptor: | Nuclear polyadenylated RNA-binding protein NAB2, ZINC ION | | Authors: | Brockmann, C, Neuhaus, D, Stewart, M. | | Deposit date: | 2011-08-12 | | Release date: | 2012-06-27 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Structural Basis for Polyadenosine-RNA Binding by Nab2 Zn Fingers and Its Function in mRNA Nuclear Export.
Structure, 20, 2012
|
|
2YTC
 
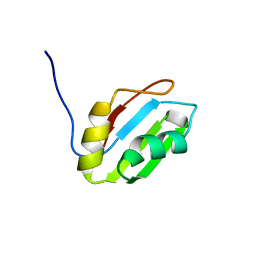 | | Solution structure of RNA binding domain in Pre-mRNA-splicing factor RBM22 | | Descriptor: | Pre-mRNA-splicing factor RBM22 | | Authors: | Kasahara, N, Tsuda, K, Muto, Y, Inoue, M, Kigawa, T, Terada, T, Shirouzu, M, Yokoyama, S, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2007-04-05 | | Release date: | 2007-10-09 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Solution structure of RNA binding domain in Pre-mRNA-splicing factor RBM22
To be Published
|
|
1E4P
 
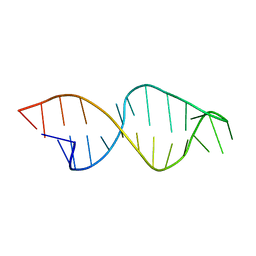 | |
8AMK
 
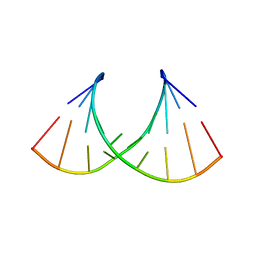 | |
8AMN
 
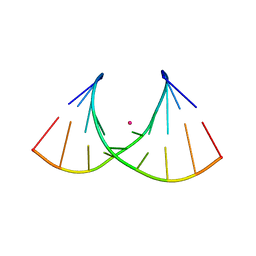 | | Crystal structure of AUGUGGCAU duplex with strontium ions | | Descriptor: | RNA (5'-R(*AP*UP*GP*UP*GP*GP*CP*AP*U)-3'), STRONTIUM ION | | Authors: | Kiliszek, A, Rypniewski, W. | | Deposit date: | 2022-08-03 | | Release date: | 2022-11-23 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.46 Å) | | Cite: | Structure and thermodynamics of a UGG motif interacting with Ba2+ and other metal ions: accommodating changes in the RNA structure and the presence of a G(syn)-G(syn) pair.
Rna, 29, 2022
|
|
1WZ2
 
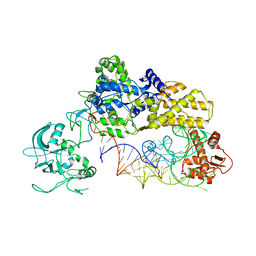 | |
3FNR
 
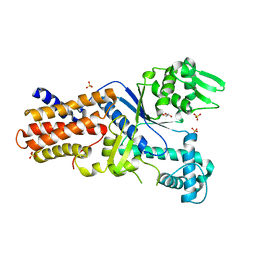 | | CRYSTAL STRUCTURE OF PUTATIVE ARGINYL T-RNA SYNTHETASE FROM Campylobacter jejuni; | | Descriptor: | Arginyl-tRNA synthetase, GLYCEROL, SULFATE ION | | Authors: | Patskovsky, Y, Ramagopal, U, Toro, R, Gilmore, M, Chang, S, Groshong, C, Sauder, J.M, Burley, S.K, Almo, S.C, New York SGX Research Center for Structural Genomics (NYSGXRC) | | Deposit date: | 2008-12-26 | | Release date: | 2009-01-27 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | CRYSTAL STRUCTURE OF A PUTATIVE ARGINYL T-RNA SYNTHETASE FROM Campylobacter jejuni
To be Published
|
|
2L2J
 
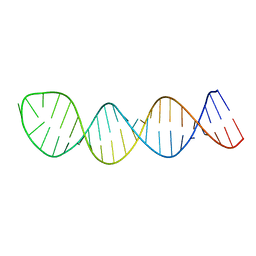 | |
5V2R
 
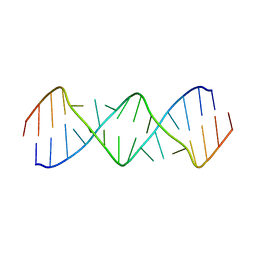 | | Structure of a GA Rich 8x8 Nucleotide RNA Internal Loop | | Descriptor: | RNA (5'-R(*CP*CP*AP*GP*AP*AP*AP*CP*GP*GP*AP*UP*GP*GP*A)-3') | | Authors: | Kauffmann, A.D, Kennedy, S.D, Turner, D.H. | | Deposit date: | 2017-03-06 | | Release date: | 2017-07-26 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Nuclear Magnetic Resonance Structure of an 8 8 Nucleotide RNA Internal Loop Flanked on Each Side by Three Watson-Crick Pairs and Comparison to Three-Dimensional Predictions.
Biochemistry, 56, 2017
|
|
2RPT
 
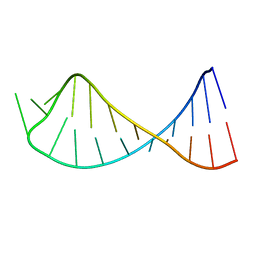 | |
1G2J
 
 | | RNA OCTAMER R(CCCP*GGGG) CONTAINING PHENYL RIBONUCLEOTIDE | | Descriptor: | 5'-R(*CP*CP*CP*(PYY)P*GP*GP*GP*G)-3', CALCIUM ION | | Authors: | Minasov, G, Matulic-Adamic, J, Wilds, C.J, Haeberli, P, Usman, N, Beigelman, L, Egli, M. | | Deposit date: | 2000-10-20 | | Release date: | 2000-12-06 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (1.97 Å) | | Cite: | Crystal structure of an RNA duplex containing phenyl-ribonucleotides, hydrophobic isosteres of the natural pyrimidines.
RNA, 6, 2000
|
|
