3O4A
 
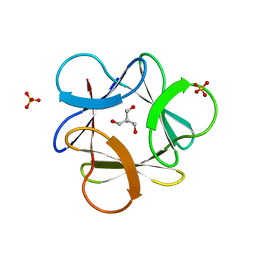 | |
8CYK
 
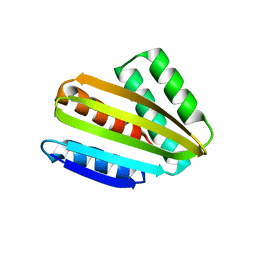 | | Crystal structure of hallucinated protein HALC1_878 | | Descriptor: | HALC1_878 | | Authors: | Ragotte, R.J, Bera, A.K, Milles, L.F, Wicky, B.I.M, Baker, D. | | Deposit date: | 2022-05-23 | | Release date: | 2022-09-28 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | Robust deep learning-based protein sequence design using ProteinMPNN.
Science, 378, 2022
|
|
7T2F
 
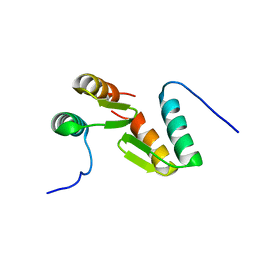 | | Solution structure of the model HEEH mini protein homodimer HEEH_TK_rd5_0341 | | Descriptor: | HEEH mini protein HEEH_TK_rd5_0341 | | Authors: | Lemak, A, Houliston, S, Kim, T.-E, Martel, C, Rocklin, G.J, Arrowsmith, C.H. | | Deposit date: | 2021-12-04 | | Release date: | 2022-10-05 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | Dissecting the stability determinants of a challenging de novo protein fold using massively parallel design and experimentation.
Proc.Natl.Acad.Sci.USA, 119, 2022
|
|
7UCP
 
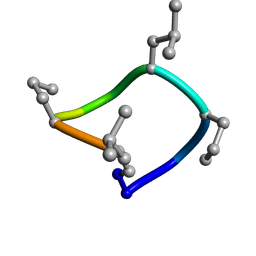 | | computationally designed macrocycle | | Descriptor: | computationally designed cyclic peptide D8.3.p2 | | Authors: | Bhardwaj, G, Baker, D, Rettie, S, Glynn, C, Sawaya, M. | | Deposit date: | 2022-03-17 | | Release date: | 2022-09-14 | | Last modified: | 2022-09-28 | | Method: | X-RAY DIFFRACTION (0.85 Å) | | Cite: | Accurate de novo design of membrane-traversing macrocycles.
Cell, 185, 2022
|
|
3O4D
 
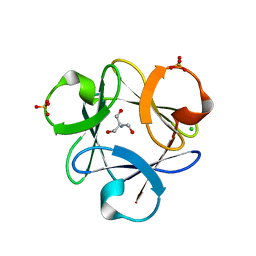 | |
3O49
 
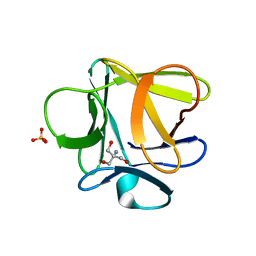 | |
3OGF
 
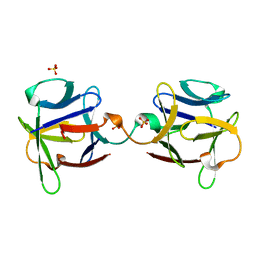 | |
3OL0
 
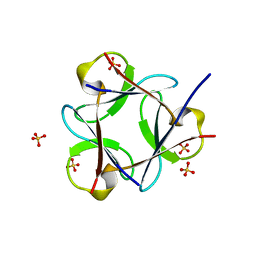 | |
5KAY
 
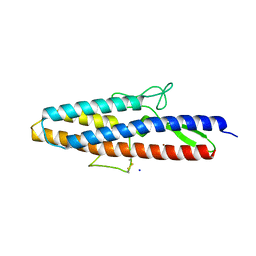 | | Structure of Spelter bound to Zn2+ | | Descriptor: | SODIUM ION, Spelter, ZINC ION | | Authors: | Guffy, S.L, Der, B.S, Kuhlman, B. | | Deposit date: | 2016-06-02 | | Release date: | 2016-08-03 | | Last modified: | 2019-11-27 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Probing the minimal determinants of zinc binding with computational protein design.
Protein Eng.Des.Sel., 29, 2016
|
|
7BG1
 
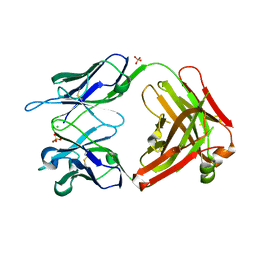 | |
8T6E
 
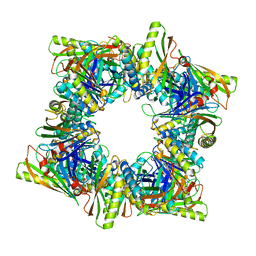 | | Crystal structure of T33-28.3: Deep-learning sequence design of co-assembling tetrahedron protein nanoparticles | | Descriptor: | T33-28.3: A, T33-28.3: B | | Authors: | Bera, A.K, de Haas, R.J, Kang, A, Sankaran, B, King, N.P. | | Deposit date: | 2023-06-15 | | Release date: | 2024-04-24 | | Method: | X-RAY DIFFRACTION (2.48 Å) | | Cite: | Rapid and automated design of two-component protein nanomaterials using ProteinMPNN.
Proc.Natl.Acad.Sci.USA, 121, 2024
|
|
8T6C
 
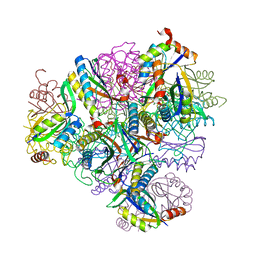 | | Crystal structure of T33-18.2: Deep-learning sequence design of co-assembling tetrahedron protein nanoparticles | | Descriptor: | T33-18.2 : A, T33-18.2 : B | | Authors: | Bera, A.K, de Haas, R.J, Kang, A, Sankaran, B, King, N.P. | | Deposit date: | 2023-06-15 | | Release date: | 2024-04-24 | | Method: | X-RAY DIFFRACTION (1.92 Å) | | Cite: | Rapid and automated design of two-component protein nanomaterials using ProteinMPNN.
Proc.Natl.Acad.Sci.USA, 121, 2024
|
|
8T6N
 
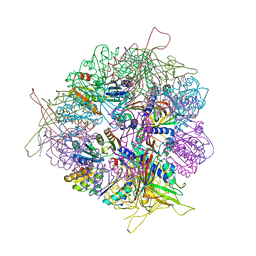 | | Crystal structure of T33-27.1: Deep-learning sequence design of co-assembling tetrahedron protein nanoparticles | | Descriptor: | T33-27.1 : A, T33-27.1 : B | | Authors: | Bera, A.K, de Haas, R.J, Kang, A, Sankaran, B, King, N.P. | | Deposit date: | 2023-06-16 | | Release date: | 2024-04-24 | | Method: | X-RAY DIFFRACTION (3.63 Å) | | Cite: | Rapid and automated design of two-component protein nanomaterials using ProteinMPNN.
Proc.Natl.Acad.Sci.USA, 121, 2024
|
|
4PN9
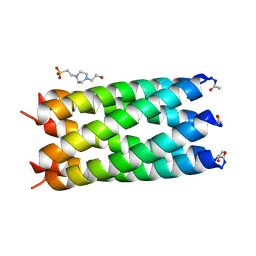 
 | | A de novo designed hexameric coiled coil CC-Hex2 | | Descriptor: | 4-(2-HYDROXYETHYL)-1-PIPERAZINE ETHANESULFONIC ACID, CC-Hex2 | | Authors: | Wood, C.W, Burton, A.J, Thomson, A.R, Brady, R.L, Woolfson, D.N. | | Deposit date: | 2014-05-23 | | Release date: | 2014-10-22 | | Last modified: | 2024-05-01 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Computational design of water-soluble alpha-helical barrels.
Science, 346, 2014
|
|
4PNA
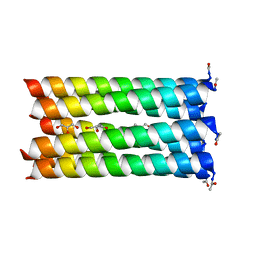 
 | | A de novo designed heptameric coiled coil CC-Hept | | Descriptor: | CC-Hept, GLYCEROL | | Authors: | Burton, A.J, Wood, C.W, Thomson, A.R, Brady, R.L, Woolfson, D.N. | | Deposit date: | 2014-05-23 | | Release date: | 2014-10-22 | | Last modified: | 2017-09-13 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Computational design of water-soluble alpha-helical barrels.
Science, 346, 2014
|
|
4PN8
 
 | | A de novo designed pentameric coiled coil CC-Pent. | | Descriptor: | CC-Pent | | Authors: | Wood, C.W, Burton, A.J, Thomson, A.R, Brady, R.L, Woolfson, D.N. | | Deposit date: | 2014-05-23 | | Release date: | 2014-10-22 | | Last modified: | 2017-09-13 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Computational design of water-soluble alpha-helical barrels.
Science, 346, 2014
|
|
4PNB
 
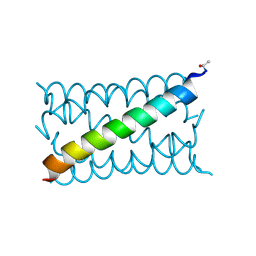 | | A de novo designed hexameric coiled coil CC-Hex3. | | Descriptor: | CC-Hex3 | | Authors: | Wood, C.W, Burton, A.J, Thomson, A.R, Brady, R.L, Woolfson, D.N. | | Deposit date: | 2014-05-23 | | Release date: | 2014-10-22 | | Last modified: | 2019-12-11 | | Method: | X-RAY DIFFRACTION (2.052 Å) | | Cite: | Computational design of water-soluble alpha-helical barrels.
Science, 346, 2014
|
|
4Q4Z
 
 | | Thermus thermophilus RNA polymerase de novo transcription initiation complex | | Descriptor: | 5'-O-[(S)-hydroxy{[(S)-hydroxy(phosphonooxy)phosphoryl]methyl}phosphoryl]cytidine, ADENOSINE-5'-TRIPHOSPHATE, DNA (25-MER), ... | | Authors: | Murakami, K.S. | | Deposit date: | 2014-04-15 | | Release date: | 2014-07-30 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Structural basis of transcription initiation by bacterial RNA polymerase holoenzyme.
J.Biol.Chem., 289, 2014
|
|
4PND
 
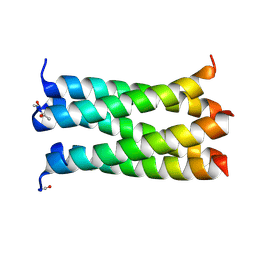 | | A de novo designed pentameric coiled coil CC-Pent_Variant | | Descriptor: | CC-Pent_Variant | | Authors: | Wood, C.W, Burton, A.J, Thomson, A.R, Brady, R.L, Woolfson, D.N. | | Deposit date: | 2014-05-23 | | Release date: | 2014-10-22 | | Last modified: | 2017-09-13 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Computational design of water-soluble alpha-helical barrels.
Science, 346, 2014
|
|
3R2X
 
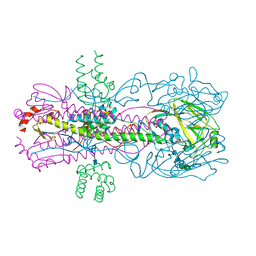 | |
1CHU
 
 | |
1DB3
 
 | | E.COLI GDP-MANNOSE 4,6-DEHYDRATASE | | Descriptor: | GDP-MANNOSE 4,6-DEHYDRATASE | | Authors: | Somoza, J.R, Menon, S, Somers, W.S, Sullivan, F.X. | | Deposit date: | 1999-11-02 | | Release date: | 1999-11-24 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Structural and kinetic analysis of Escherichia coli GDP-mannose 4,6 dehydratase provides insights into the enzyme's catalytic mechanism and regulation by GDP-fucose.
Structure Fold.Des., 8, 2000
|
|
5IEN
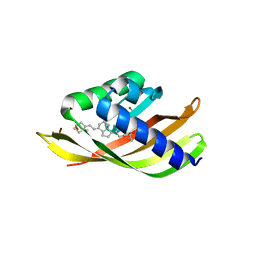 
 | | Structure of CDL2.2, a computationally designed Vitamin-D3 binder | | Descriptor: | 3-{2-[1-(5-HYDROXY-1,5-DIMETHYL-HEXYL)-7A-METHYL-OCTAHYDRO-INDEN-4-YLIDENE]-ETHYLIDENE}-4-METHYLENE-CYCLOHEXANOL, CDL2.2, GLYCEROL | | Authors: | Stoddard, B.L, Doyle, L.A. | | Deposit date: | 2016-02-25 | | Release date: | 2017-03-01 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (2.089 Å) | | Cite: | Unintended specificity of an engineered ligand-binding protein facilitated by unpredicted plasticity of the protein fold.
Protein Eng.Des.Sel., 31, 2018
|
|
5IEO
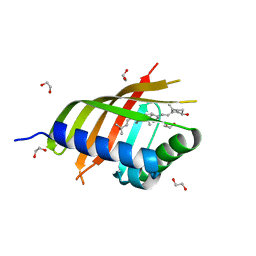 
 | | Structure of CDL2.3a, a computationally designed Vitamin-D3 binder | | Descriptor: | 1,2-ETHANEDIOL, 3-{2-[1-(5-HYDROXY-1,5-DIMETHYL-HEXYL)-7A-METHYL-OCTAHYDRO-INDEN-4-YLIDENE]-ETHYLIDENE}-4-METHYLENE-CYCLOHEXANOL, CDL2.3a | | Authors: | Stoddard, B.L, Doyle, L.A. | | Deposit date: | 2016-02-25 | | Release date: | 2017-03-01 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (1.851 Å) | | Cite: | Unintended specificity of an engineered ligand-binding protein facilitated by unpredicted plasticity of the protein fold.
Protein Eng.Des.Sel., 31, 2018
|
|
5IEP
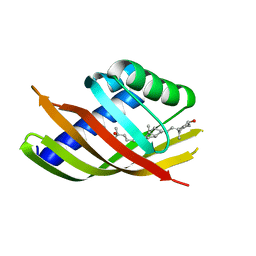 
 | | Structure of CDL2.3b, a computationally designed Vitamin-D3 binder | | Descriptor: | 3-{2-[1-(5-HYDROXY-1,5-DIMETHYL-HEXYL)-7A-METHYL-OCTAHYDRO-INDEN-4-YLIDENE]-ETHYLIDENE}-4-METHYLENE-CYCLOHEXANOL, CDL2.3b | | Authors: | Stoddard, B.L, Doyle, L.A. | | Deposit date: | 2016-02-25 | | Release date: | 2017-03-01 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (1.893 Å) | | Cite: | Unintended specificity of an engineered ligand-binding protein facilitated by unpredicted plasticity of the protein fold.
Protein Eng.Des.Sel., 31, 2018
|
|
