4QOY
 
 | |
3FLT
 
 | |
4QAM
 
 | | Crystal Structure of the RPGR RCC1-like domain in complex with the RPGR-interacting domain of RPGRIP1 | | Descriptor: | GLYCEROL, MAGNESIUM ION, X-linked retinitis pigmentosa GTPase regulator, ... | | Authors: | Remans, K, Buerger, M, Vetter, I.R, Wittinghofer, A. | | Deposit date: | 2014-05-05 | | Release date: | 2014-07-30 | | Last modified: | 2023-12-06 | | Method: | X-RAY DIFFRACTION (1.83 Å) | | Cite: | C2 domains as protein-protein interaction modules in the ciliary transition zone.
Cell Rep, 8, 2014
|
|
4QBB
 
 | | Structure of the foot-and-mouth disease virus leader proteinase in complex with inhibitor (N~2~-[(3S)-4-({(2R)-1-[(4-CARBAMIMIDAMIDOBUTYL)AMINO]-4-METHYL-1-OXOPENTAN-2-YL}AMINO)-3-HYDROXY-4-OXOBUTANOYL]-L-ARGINYL-L-PROLINAMIDE) | | Descriptor: | Leader protease, N~2~-[(3S)-4-({(2R)-1-[(4-carbamimidamidobutyl)amino]-4-methyl-1-oxopentan-2-yl}amino)-3-hydroxy-4-oxobutanoyl]-L-arginyl-L-prolinamide, PHOSPHATE ION, ... | | Authors: | Grishkovskaya, I, Steinberger, J, Cencic, R, Juliano, M.A, Juliano, L, Skern, T. | | Deposit date: | 2014-05-07 | | Release date: | 2014-11-05 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Foot-and-mouth disease virus leader proteinase: Structural insights into the mechanism of intermolecular cleavage.
Virology, 468-470C, 2014
|
|
3FVB
 
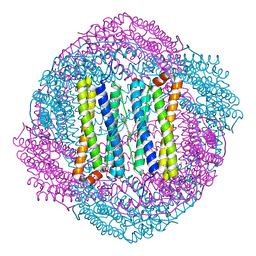 | |
3FWU
 
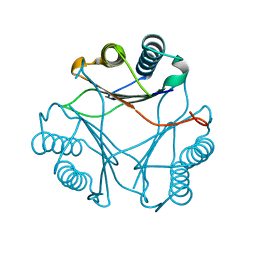 | | Crystal structure of Leishmania major MIF1 | | Descriptor: | Macrophage migration inhibitory factor-like protein | | Authors: | RIchardson, J.M, Walkinshaw, M.D. | | Deposit date: | 2009-01-19 | | Release date: | 2009-03-24 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Structures of Leishmania major orthologues of macrophage migration inhibitory factor
Biochem.Biophys.Res.Commun., 380, 2009
|
|
4S0U
 
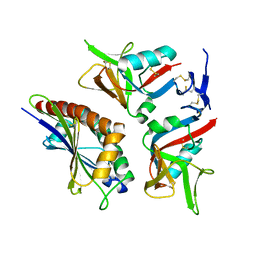 | | Crystal structure of NKG2D in complex with ULBP6 | | Descriptor: | NKG2-D type II integral membrane protein, Retinoic acid early transcript 1L protein | | Authors: | Mohammed, F, Willcox, B.E. | | Deposit date: | 2015-01-06 | | Release date: | 2016-04-20 | | Last modified: | 2017-06-14 | | Method: | X-RAY DIFFRACTION (2.35 Å) | | Cite: | A disease-linked ULBP6 polymorphism inhibits NKG2D-mediated target cell killing by enhancing the stability of NKG2D ligand binding.
Sci Signal, 10, 2017
|
|
4S1C
 
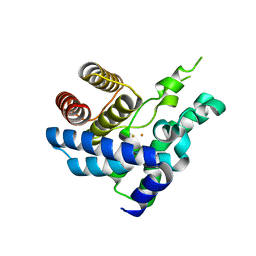 | |
3G10
 
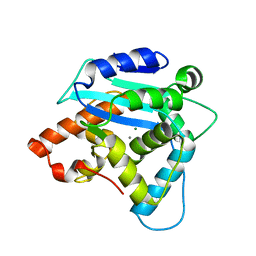 | | Structure of S. pombe Pop2p - Mg2+ and Mn2+ bound form | | Descriptor: | CCR4-Not complex subunit Caf1, MAGNESIUM ION, MANGANESE (II) ION | | Authors: | Andersen, K.R, Jonstrup, A.T, Van, L.B, Brodersen, D.E. | | Deposit date: | 2009-01-29 | | Release date: | 2009-03-31 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.597 Å) | | Cite: | The activity and selectivity of fission yeast Pop2p are affected by a high affinity for Zn2+ and Mn2+ in the active site
Rna, 15, 2009
|
|
3G2D
 
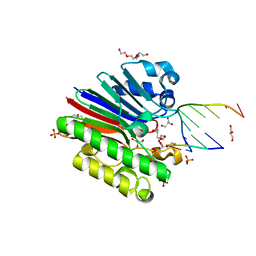 | | Complex of Mth0212 and a 4 bp dsDNA with 3'-overhang | | Descriptor: | 5'-D(*CP*CP*TP*GP*UP*GP*CP*GP*AP*T)-3', 5'-D(*CP*GP*CP*G*CP*AP*GP*GP*C)-3', Exodeoxyribonuclease, ... | | Authors: | Lakomek, K, Dickmanns, A, Ficner, R. | | Deposit date: | 2009-01-31 | | Release date: | 2010-03-09 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Crystal Structure Analysis of DNA Uridine Endonuclease Mth212 Bound to DNA
J.Mol.Biol., 399, 2010
|
|
3G05
 
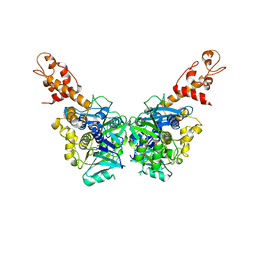 | | Crystal structure of N-terminal domain (2-550) of E.coli MnmG | | Descriptor: | SULFATE ION, tRNA uridine 5-carboxymethylaminomethyl modification enzyme mnmG | | Authors: | Shi, R, Matte, A, Cygler, M, Montreal-Kingston Bacterial Structural Genomics Initiative (BSGI) | | Deposit date: | 2009-01-27 | | Release date: | 2009-10-20 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (3.49 Å) | | Cite: | Structure-function analysis of Escherichia coli MnmG (GidA), a highly conserved tRNA-modifying enzyme.
J.Bacteriol., 191, 2009
|
|
3WC7
 
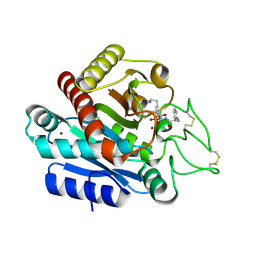 | | Carboxypeptidase B in complex with EF6265 | | Descriptor: | (2S)-7-amino-2-{[(R)-hydroxy{(1R)-2-methyl-1-[(3-phenylpropanoyl)amino]propyl}phosphoryl]methyl}heptanoic acid, Carboxypeptidase B, ZINC ION | | Authors: | Yoshimoto, N, Itoh, T, Inaba, Y, Yamamoto, K. | | Deposit date: | 2013-05-24 | | Release date: | 2013-10-02 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Structural basis for inhibition of carboxypeptidase B by selenium-containing inhibitor: selenium coordinates to zinc in enzyme.
J.Med.Chem., 56, 2013
|
|
3W9V
 
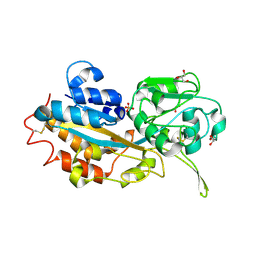 | | Crystal structure of refolded DING protein | | Descriptor: | GLYCEROL, PHOSPHATE ION, Phosphate-binding protein | | Authors: | Gai, Z.Q, Nakamura, A, Tanaka, Y, Hirano, N, Tanaka, I, Yao, M. | | Deposit date: | 2013-04-17 | | Release date: | 2013-10-30 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.031 Å) | | Cite: | Crystal structure analysis, overexpression and refolding behaviour of a DING protein with single mutation.
J.SYNCHROTRON RADIAT., 20, 2013
|
|
3WAB
 
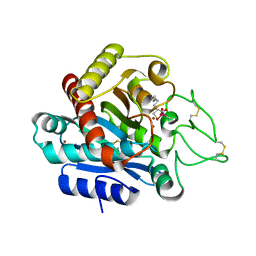 | | Carboxypeptidase B in complex with DD2 | | Descriptor: | (2R)-7-amino-2-(sulfanylmethyl)heptanoic acid, CACODYLATE ION, Carboxypeptidase B, ... | | Authors: | Yoshimoto, N, Itoh, T, Inaba, Y, Yamamoto, K. | | Deposit date: | 2013-05-01 | | Release date: | 2013-10-02 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.154 Å) | | Cite: | Structural basis for inhibition of carboxypeptidase B by selenium-containing inhibitor: selenium coordinates to zinc in enzyme.
J.Med.Chem., 56, 2013
|
|
4RLW
 
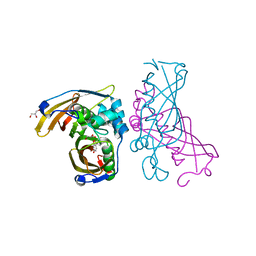 | | Crystal Structure of (3R)-hydroxyacyl-ACP dehydratase HadAB hetero-dimer from Mycobacterium tuberculosis complexed with Butein | | Descriptor: | (2E)-1-(2,4-dihydroxyphenyl)-3-(3,4-dihydroxyphenyl)prop-2-en-1-one, (3R)-hydroxyacyl-ACP dehydratase subunit HadA, (3R)-hydroxyacyl-ACP dehydratase subunit HadB, ... | | Authors: | Li, J, Dong, Y, Rao, Z.H. | | Deposit date: | 2014-10-18 | | Release date: | 2015-10-21 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.196 Å) | | Cite: | Molecular basis for the inhibition of beta-hydroxyacyl-ACP dehydratase HadAB complex from Mycobacterium tuberculosis by flavonoid inhibitors.
Protein Cell, 6, 2015
|
|
4RP4
 
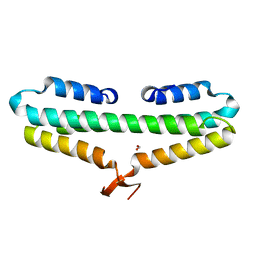 | |
3G7W
 
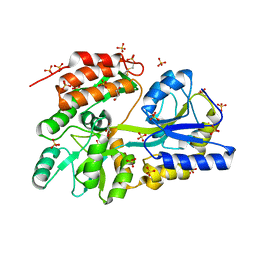 | | Islet Amyloid Polypeptide (IAPP or Amylin) Residues 1 to 22 fused to Maltose Binding Protein | | Descriptor: | GLYCEROL, Maltose-binding periplasmic protein, Islet amyloid polypeptide fusion protein, ... | | Authors: | Wiltzius, J.J.W, Sawaya, M.R, Eisenberg, D. | | Deposit date: | 2009-02-11 | | Release date: | 2009-06-23 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Atomic structures of IAPP (amylin) fusions suggest a mechanism for fibrillation and the role of insulin in the process
Protein Sci., 18, 2009
|
|
3FVA
 | | NNQNTF segment from elk prion | | Descriptor: | Major prion protein | | Authors: | Apostol, M.I, Eisenberg, D. | | Deposit date: | 2009-01-15 | | Release date: | 2009-06-30 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (1.458 Å) | | Cite: | Molecular mechanisms for protein-encoded inheritance.
Nat.Struct.Mol.Biol., 16, 2009
|
|
3WG1
 
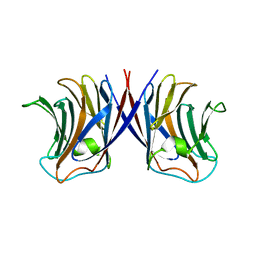 | | Crystal structure of Agrocybe cylindracea galectin with lactose | | Descriptor: | Galactoside-binding lectin, beta-D-galactopyranose-(1-4)-beta-D-glucopyranose | | Authors: | Kuwabara, N, Hu, D, Tateno, H, Makio, H, Hirabayashi, J, Kato, R. | | Deposit date: | 2013-07-25 | | Release date: | 2013-11-27 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Conformational change of a unique sequence in a fungal galectin from Agrocybe cylindracea controls glycan ligand-binding specificity.
Febs Lett., 587, 2013
|
|
3FVC
 
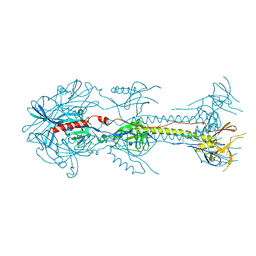 | |
3FVR
 
 | |
4RTC
 | | Crystal structure of the green fluorescent variant, nowGFP, of the cyan Cerulean at pH 9.0 | | Descriptor: | GLYCEROL, nowGFP | | Authors: | Pletnev, V.Z, Pletneva, N.V, Pletnev, S.V. | | Deposit date: | 2014-11-14 | | Release date: | 2015-09-02 | | Last modified: | 2023-12-06 | | Method: | X-RAY DIFFRACTION (1.35 Å) | | Cite: | Structure of the green fluorescent protein NowGFP with an anionic tryptophan-based chromophore.
Acta Crystallogr.,Sect.D, 71, 2015
|
|
3WFE
 
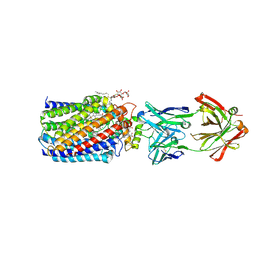 | | Reduced and cyanide-bound cytochrome c-dependent nitric oxide reductase (cNOR) from Pseudomonas aeruginosa in complex with antibody fragment | | Descriptor: | CALCIUM ION, CYANIDE ION, FE (III) ION, ... | | Authors: | Sato, N, Ishii, S, Hino, T, Sugimoto, H, Fukumori, Y, Shiro, Y, Tosha, T. | | Deposit date: | 2013-07-18 | | Release date: | 2014-05-28 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.49 Å) | | Cite: | Structures of reduced and ligand-bound nitric oxide reductase provide insights into functional differences in respiratory enzymes.
Proteins, 82, 2014
|
|
3FTK
 | |
3FYT
 
 | |
