2O3K
 
 | |
2O3L
 
 | |
2O3M
 
 | | Monomeric G-DNA tetraplex from human C-kit promoter | | Descriptor: | 5'-D(*AP*GP*GP*GP*AP*GP*GP*GP*CP*GP*CP*TP*GP*GP*GP*AP*GP*GP*AP*GP*GP*G)-3' | | Authors: | Phan, A.T, Kuryavyi, V.V, Burge, S, Neidle, S, Patel, D.J. | | Deposit date: | 2006-12-01 | | Release date: | 2007-05-22 | | Last modified: | 2023-12-27 | | Method: | SOLUTION NMR | | Cite: | Structure of an unprecedented G-quadruplex scaffold in the human c-kit promoter.
J.Am.Chem.Soc., 129, 2007
|
|
2O3O
 | |
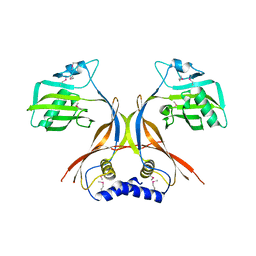
2O3P
 
 | | Crystal structure of Pim1 with Quercetin | | Descriptor: | 3,5,7,3',4'-PENTAHYDROXYFLAVONE, IMIDAZOLE, Proto-oncogene serine/threonine-protein kinase Pim-1 | | Authors: | Holder, S, Zemskova, M, Zhang, C, Tabrizizad, M, Bremer, R, Neidigh, J.W, Lilly, M.B. | | Deposit date: | 2006-12-01 | | Release date: | 2007-02-13 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (2.24 Å) | | Cite: | Characterization of a potent and selective small-molecule inhibitor of the PIM1 kinase.
Mol.Cancer Ther., 6, 2007
|
|
2O3Q
 
 | | Structural Basis for Formation and Hydrolysis of Calcium Messenger Cyclic ADP-ribose by Human CD38 | | Descriptor: | ADP-ribosyl cyclase 1, CYCLIC ADENOSINE DIPHOSPHATE-RIBOSE | | Authors: | Liu, Q, Kriksunov, I.A, Graeff, R, Lee, H.C, Hao, Q. | | Deposit date: | 2006-12-01 | | Release date: | 2006-12-12 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.98 Å) | | Cite: | Structural basis for formation and hydrolysis of the calcium messenger cyclic ADP-ribose by human CD38
J.Biol.Chem., 282, 2007
|
|
2O3R
 
 | | Structural Basis for Formation and Hydrolysis of Calcium Messenger Cyclic ADP-ribose by Human CD38 | | Descriptor: | ADP-ribosyl cyclase 1, CYCLIC ADENOSINE DIPHOSPHATE-RIBOSE | | Authors: | Liu, Q, Kriksunov, I.A, Graeff, R, Lee, H.C, Hao, Q. | | Deposit date: | 2006-12-01 | | Release date: | 2006-12-12 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Structural basis for formation and hydrolysis of the calcium messenger cyclic ADP-ribose by human CD38
J.Biol.Chem., 282, 2007
|
|
2O3S
 
 | | Structural Basis for Formation and Hydrolysis of Calcium Messenger Cyclic ADP-ribose by Human CD38 | | Descriptor: | ADP-ribosyl cyclase 1, CYCLIC ADENOSINE DIPHOSPHATE-RIBOSE | | Authors: | Liu, Q, Kriksunov, I.A, Graeff, R, Lee, H.C, Hao, Q. | | Deposit date: | 2006-12-01 | | Release date: | 2006-12-12 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Structural basis for formation and hydrolysis of the calcium messenger cyclic ADP-ribose by human CD38
J.Biol.Chem., 282, 2007
|
|
2O3T
 
 | | Structural Basis for Formation and Hydrolysis of Calcium Messenger Cyclic ADP-ribose by Human CD38 | | Descriptor: | ADP-ribosyl cyclase 1, CYCLIC GUANOSINE DIPHOSPHATE-RIBOSE | | Authors: | Liu, Q, Kriksunov, I.A, Graeff, R, Lee, H.C, Hao, Q. | | Deposit date: | 2006-12-01 | | Release date: | 2006-12-19 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.68 Å) | | Cite: | Structural basis for formation and hydrolysis of the calcium messenger cyclic ADP-ribose by human CD38
J.Biol.Chem., 282, 2007
|
|
2O3U
 
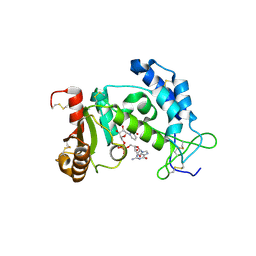 | | Structural Basis for Formation and Hydrolysis of Calcium Messenger Cyclic ADP-ribose by Human CD38 | | Descriptor: | 3-(AMINOCARBONYL)-1-[(2R,3R,4S,5R)-5-({[(S)-{[(S)-{[(2R,3S,4R,5R)-5-(2-AMINO-6-OXO-1,6-DIHYDRO-9H-PURIN-9-YL)-3,4-DIHYD ROXYTETRAHYDROFURAN-2-YL]METHOXY}(HYDROXY)PHOSPHORYL]OXY}(HYDROXY)PHOSPHORYL]OXY}METHYL)-3,4-DIHYDROXYTETRAHYDROFURAN-2- YL]PYRIDINIUM, ADP-ribosyl cyclase 1 | | Authors: | Liu, Q, Kriksunov, I.A, Graeff, R, Lee, H.C, Hao, Q. | | Deposit date: | 2006-12-01 | | Release date: | 2006-12-19 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.11 Å) | | Cite: | Structural basis for formation and hydrolysis of the calcium messenger cyclic ADP-ribose by human CD38
J.Biol.Chem., 282, 2007
|
|
2O3V
 
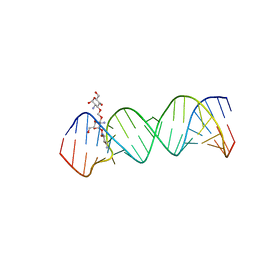 | | Crystal Structure of the Homo sapiens Cytoplasmic Ribosomal Decoding Site complexed with paromamine derivative NB33 | | Descriptor: | (2S,3R,4R,5S,6R)-3-AMINO-4-({[(2S,3R,4R,5S,6R)-3-AMINO-2-{[(1R,2R,3S,4R,6S)-4,6-DIAMINO-2,3-DIHYDROXYCYCLOHEXYL]OXY}-5-HYDROXY-6-(HYDROXYMETHYL)TETRAHYDRO-2H-PYRAN-4-YL]OXY}METHOXY)-6-(HYDROXYMETHYL)TETRAHYDRO-2H-PYRAN-2,5-DIOL, RNA (5'-R(*UP*UP*GP*CP*GP*UP*CP*GP*CP*UP*CP*CP*GP*GP*AP*AP*AP*AP*GP*UP*CP*GP*C)-3') | | Authors: | Kondo, J, Hainrichson, M, Nudelman, I, Shallom-Shezifi, D, Baasov, T, Westhof, E. | | Deposit date: | 2006-12-02 | | Release date: | 2007-11-06 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Differential Selectivity of Natural and Synthetic Aminoglycosides towards the Eukaryotic and Prokaryotic Decoding A Sites.
Chembiochem, 8, 2007
|
|
2O3W
 
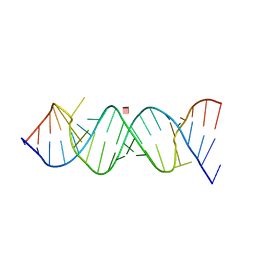 | | Crystal Structure of the Homo sapiens Cytoplasmic Ribosomal Decoding Site in presence of paromomycin | | Descriptor: | PAROMOMYCIN, RNA (5'-R(*UP*UP*GP*CP*GP*UP*CP*GP*CP*UP*CP*CP*GP*GP*AP*AP*AP*AP*GP*UP*CP*GP*C)-3') | | Authors: | Kondo, J, Hainrichson, M, Nudelman, I, Shallom-Shezifi, D, Baasov, T, Westhof, E. | | Deposit date: | 2006-12-02 | | Release date: | 2007-11-06 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Differential Selectivity of Natural and Synthetic Aminoglycosides towards the Eukaryotic and Prokaryotic Decoding A Sites.
Chembiochem, 8, 2007
|
|
2O3X
 
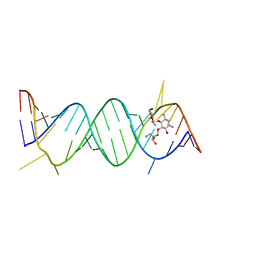 | | Crystal Structure of the Prokaryotic Ribosomal Decoding Site Complexed with Paromamine Derivative NB30 | | Descriptor: | (1R,2R,3S,4R,6S)-4,6-DIAMINO-2-[(5-AMINO-5-DEOXY-BETA-D-RIBOFURANOSYL)OXY]-3-HYDROXYCYCLOHEXYL 2-AMINO-2-DEOXY-ALPHA-D-GLUCOPYRANOSIDE, RNA (5'-R(*UP*UP*GP*CP*GP*UP*CP*AP*CP*AP*CP*CP*GP*GP*UP*GP*AP*AP*GP*UP*CP*GP*C)-3') | | Authors: | Kondo, J, Hainrichson, M, Nudelman, I, Shallom-Shezifi, D, Baasov, T, Westhof, E. | | Deposit date: | 2006-12-02 | | Release date: | 2007-11-06 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Differential Selectivity of Natural and Synthetic Aminoglycosides towards the Eukaryotic and Prokaryotic Decoding A Sites.
Chembiochem, 8, 2007
|
|
2O3Y
 
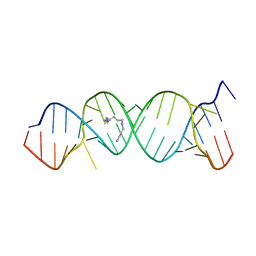 | | Crystal Structure of the Homo sapiens Cytoplasmic Ribosomal Decoding Site in Presence of Paromamine Derivative NB30 | | Descriptor: | RNA (5'-R(*UP*UP*GP*CP*GP*UP*CP*GP*CP*UP*CP*CP*GP*GP*AP*AP*AP*AP*GP*UP*CP*GP*C)-3'), SPERMINE | | Authors: | Kondo, J, Hainrichson, M, Nudelman, I, Shallom-Shezifi, D, Baasov, T, Westhof, E. | | Deposit date: | 2006-12-02 | | Release date: | 2007-11-06 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Differential Selectivity of Natural and Synthetic Aminoglycosides towards the Eukaryotic and Prokaryotic Decoding A Sites.
Chembiochem, 8, 2007
|
|
2O3Z
 
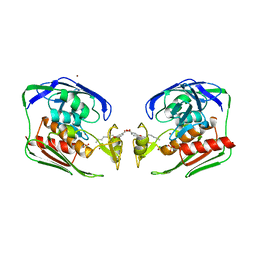 | | X-ray crystal structure of LpxC complexed with 3-heptyloxybenzoate | | Descriptor: | 3-(heptyloxy)benzoic acid, CHLORIDE ION, SULFATE ION, ... | | Authors: | Gennadios, H.A, Whittington, D.A, Christianson, D.W. | | Deposit date: | 2006-12-02 | | Release date: | 2007-02-27 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.25 Å) | | Cite: | Amphipathic benzoic acid derivatives: synthesis and binding in the hydrophobic tunnel of the zinc deacetylase LpxC.
Bioorg.Med.Chem., 15, 2007
|
|
2O40
 
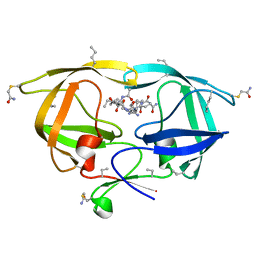 | |
2O42
 
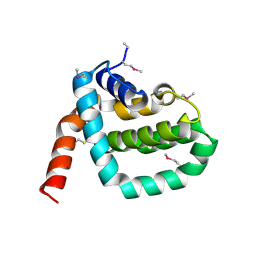 | |
2O43
 
 | | Structure of 23S rRNA of the large ribosomal subunit from Deinococcus radiodurans in complex with the macrolide erythromycylamine | | Descriptor: | (3R,4S,5S,6R,7R,9R,10S,11S,12R,13S,14R)-10-AMINO-6-{[(2S,3R,4S,6R)-4-(DIMETHYLAMINO)-3-HYDROXY-6-METHYLTETRAHYDRO-2H-PY RAN-2-YL]OXY}-14-ETHYL-7,12,13-TRIHYDROXY-4-{[(2R,4R,5S,6S)-5-HYDROXY-4-METHOXY-4,6-DIMETHYLTETRAHYDRO-2H-PYRAN-2-YL]OXY }-3,5,7,9,11,13-HEXAMETHYLOXACYCLOTETRADECAN-2-ONE, 23S rRNA | | Authors: | Pyetan, E, Baram, D, Auerbach-Nevo, T, Yonath, A. | | Deposit date: | 2006-12-03 | | Release date: | 2007-12-04 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (3.6 Å) | | Cite: | Chemical parameters influencing fine tuning in the binding of Macrolide antibiotics to the ribosomal tunnel
TO BE PUBLISHED
|
|
2O44
 
 | | Structure of 23S rRNA of the large ribosomal subunit from Deinococcus radiodurans in complex with the macrolide josamycin | | Descriptor: | (2S,3S,4R,6S)-6-{[(2R,3S,4R,5R,6S)-6-{[(4R,5S,6S,7R,9R,10S,12E,14Z,16R)-4-(ACETYLOXY)-10-HYDROXY-5-METHOXY-9,16-DIMETHYL-2-OXO-7-(2-OXOETHYL)OXACYCLOHEXADECA-12,14-DIEN-6-YL]OXY}-4-(DIMETHYLAMINO)-5-HYDROXY-2-METHYLTETRAHYDRO-2H-PYRAN-3-YL]OXY}-4-HYDROXY-2,4-DIMETHYLTETRAHYDRO-2H-PYRAN-3-YL 3-METHYLBUTANOATE, 23S rRNA | | Authors: | Pyetan, E, Daram, D, Auerbach-Nevo, T, Yonath, A. | | Deposit date: | 2006-12-03 | | Release date: | 2007-12-04 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (3.3 Å) | | Cite: | Chemical parameters influencing fine tuning in the binding of Macrolide antibiotics to the ribosomal tunnel
TO BE PUBLISHED
|
|
2O45
 
 | | Structure of the 23S rRNA of the large ribosomal subunit from Deinococcus Radiodurans in complex with the macrolide RU-69874 | | Descriptor: | (3AS,4R,7R,8S,9S,10R,11R,13R,15R,15AR)-4-ETHYL-11-METHOXY-3A,7,9,11,13,15-HEXAMETHYL-2,6,14-TRIOXO-1-[4-(4-PYRIDIN-3-YL-1H-IMIDAZOL-1-YL)BUTYL]-10-{[3,4,6-TRIDEOXY-3-(DIMETHYLAMINO)-BETA-D-XYLO-HEXOPYRANOSYL]OXY}TETRADECAHYDRO-2H-OXACYCLOTETRADECINO[4,3-D][1,3]OXAZOL-8-YL 2,6-DIDEOXY-3-C-METHYL-3-O-METHYL-ALPHA-L-RIBO-HEXOPYRANOSIDE, 23S rRNA | | Authors: | Baram, D, Pyetan, E, Auerbach-Nevo, T, Yonath, A. | | Deposit date: | 2006-12-03 | | Release date: | 2007-12-04 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (3.6 Å) | | Cite: | Chemical Parameters Influencing Fine Tuning in the Binding of Macrolide Antibiotics to the Ribosomal Tunnel
TO BE PUBLISHED
|
|
2O48
 
 | | Crystal structure of Mammalian Dimeric Dihydrodiol Dehydrogenase | | Descriptor: | BETA-MERCAPTOETHANOL, Dimeric dihydrodiol dehydrogenase, P-HYDROXYACETOPHENONE, ... | | Authors: | Carbone, V, El-Kabbani, O. | | Deposit date: | 2006-12-04 | | Release date: | 2007-07-31 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (2.59 Å) | | Cite: | Structures of dimeric dihydrodiol dehydrogenase apoenzyme and inhibitor complex: probing the subunit interface with site-directed mutagenesis.
Proteins, 70, 2008
|
|
2O49
 
 | |
2O4A
 
 | |
2O4C
 
 | | Crystal Structure of D-Erythronate-4-phosphate Dehydrogenase Complexed with NAD | | Descriptor: | Erythronate-4-phosphate dehydrogenase, GLYCEROL, L(+)-TARTARIC ACID, ... | | Authors: | Ha, J.Y, Lee, J.H, Kim, K.H, Kim, D.J, Lee, H.H, Kim, H.K, Yoon, H.J, Suh, S.W. | | Deposit date: | 2006-12-04 | | Release date: | 2007-02-20 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Crystal Structure of d-Erythronate-4-phosphate Dehydrogenase Complexed with NAD
J.Mol.Biol., 366, 2007
|
|
2O4D
 
 | | Crystal Structure of a hypothetical protein from Pseudomonas aeruginosa | | Descriptor: | Hypothetical protein PA0269 | | Authors: | McGrath, T.E, Battaile, K, Kisselman, G, Romanov, V, Wu-Brown, J, Virag, C, Ng, I, Kimber, M, Edwards, A.M, Pai, E.F, Chirgadze, N.Y. | | Deposit date: | 2006-12-04 | | Release date: | 2007-01-30 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Crystal Structure of a hypothetical protein from Pseudomonas aeruginosa
To be Published
|
|
