6HMB
 
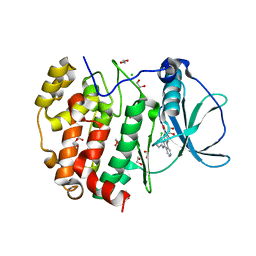 | | STRUCTURE OF PROTEIN KINASE CK2 CATALYTIC SUBUNIT (ISOFORM CK2ALPHA'; CSNK2A2 Gene product) IN COMPLEX WITH the inhibitor CX-4945 (Silmitasertib) | | Descriptor: | 1,2-ETHANEDIOL, 5-[(3-chlorophenyl)amino]benzo[c][2,6]naphthyridine-8-carboxylic acid, CHLORIDE ION, ... | | Authors: | Niefind, K, Lindenblatt, D, Applegate, V.M, Jose, J, Le Borgne, M. | | Deposit date: | 2018-09-12 | | Release date: | 2019-03-27 | | Last modified: | 2024-05-15 | | Method: | X-RAY DIFFRACTION (1.04 Å) | | Cite: | Diacritic Binding of an Indenoindole Inhibitor by CK2 alpha Paralogs Explored by a Reliable Path to Atomic Resolution CK2 alpha ' Structures.
Acs Omega, 4, 2019
|
|
4OB4
 
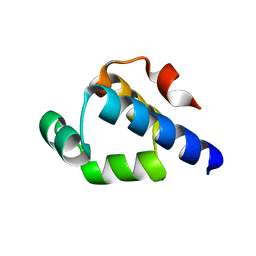 | | Structure of the S. venezulae BldD DNA-binding domain | | Descriptor: | Putative DNA-binding protein | | Authors: | schumacher, M.A, Tschowri, N, Buttner, M, Brennan, R. | | Deposit date: | 2014-01-06 | | Release date: | 2014-11-19 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Tetrameric c-di-GMP mediates effective transcription factor dimerization to control Streptomyces development.
Cell(Cambridge,Mass.), 158, 2014
|
|
2AIP
 
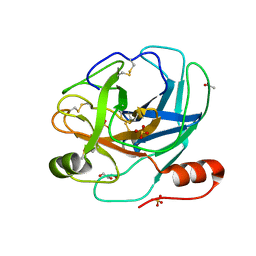 | |
3PRI
 
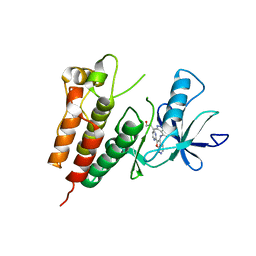 | | Crystal Structure of Human B-Raf Kinase in Complex with a Non-Oxime Furopyridine Inhibitor | | Descriptor: | 3-(4-{[2-(pyrimidin-2-yl)furo[2,3-c]pyridin-3-yl]amino}-1H-indazol-3-yl)propan-1-ol, Serine/threonine-protein kinase B-raf | | Authors: | Voegtli, W.C, Vigers, G.P.A, Morales, T, Brandhuber, B.J. | | Deposit date: | 2010-11-29 | | Release date: | 2011-02-02 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (3.5 Å) | | Cite: | Non-oxime inhibitors of B-Raf(V600E) kinase.
Bioorg.Med.Chem.Lett., 21, 2011
|
|
4JCG
 
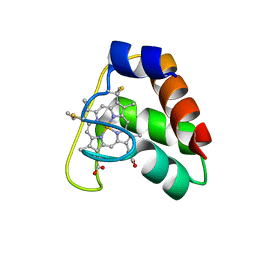 | | Recombinant wild type Nitrosomonas europaea cytochrome c552 | | Descriptor: | Cytochrome c-552, HEME C | | Authors: | Wedekind, J.E, Can, M, Krucinska, J, Bren, K.L. | | Deposit date: | 2013-02-21 | | Release date: | 2013-08-14 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.63 Å) | | Cite: | Structural Characterization of Nitrosomonas europaea Cytochrome c-552 Variants with Marked Differences in Electronic Structure.
Chembiochem, 14, 2013
|
|
2PNT
 
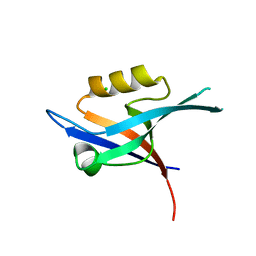 | | Crystal structure of the PDZ domain of human GRASP (GRP1) in complex with the C-terminal peptide of the metabotropic glutamate receptor type 1 | | Descriptor: | CHLORIDE ION, General receptor for phosphoinositides 1-associated scaffold protein | | Authors: | Elkins, J, Papagrigoriou, E, Cooper, C, Gileadi, C, Uppenberg, J, Bray, J, von Delft, F, Pike, A.C.W, Ugochukwu, E, Umeano, C, Gileadi, O, Edwards, A, Arrowsmith, C.H, Weigelt, J, Sundstrom, M, Doyle, D.A, Structural Genomics Consortium (SGC) | | Deposit date: | 2007-04-25 | | Release date: | 2007-05-08 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.148 Å) | | Cite: | Unusual binding interactions in PDZ domain crystal structures help explain binding mechanisms
Protein Sci., 19, 2010
|
|
1HT7
 
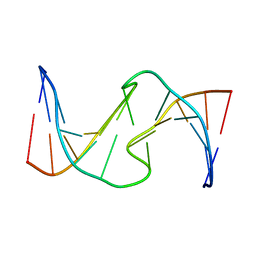 | |
5WUJ
 
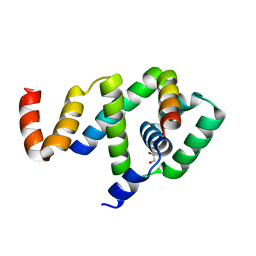 | | Crystal structure of FliF-FliG complex from H. pylori | | Descriptor: | Flagellar M-ring protein, Flagellar motor switch protein FliG, GLYCEROL | | Authors: | Au, S.W, Xue, C, Lam, K.H, Lee, S.H. | | Deposit date: | 2016-12-19 | | Release date: | 2017-12-20 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Crystal structure of the FliF-FliG complex from Helicobacter pylori yields insight into the assembly of the motor MS-C ring in the bacterial flagellum
J. Biol. Chem., 293, 2018
|
|
3LFP
 
 | |
1HT4
 
 | |
6KU8
 
 | | structure of HRV-C 3C protein with rupintrivir | | Descriptor: | 4-{2-(4-FLUORO-BENZYL)-6-METHYL-5-[(5-METHYL-ISOXAZOLE-3-CARBONYL)-AMINO]-4-OXO-HEPTANOYLAMINO}-5-(2-OXO-PYRROLIDIN-3-YL)-PENTANOIC ACID ETHYL ESTER, Genome polyprotein | | Authors: | Zhu, L, Yuan, S. | | Deposit date: | 2019-08-31 | | Release date: | 2020-03-25 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.05 Å) | | Cite: | Structure of the HRV-C 3C-Rupintrivir Complex Provides New Insights for Inhibitor Design.
Virol Sin, 35, 2020
|
|
3N1T
 
 | | Crystal structure of the H101A mutant ecHint GMP complex | | Descriptor: | GUANOSINE-5'-MONOPHOSPHATE, HIT-like protein hinT | | Authors: | Cody, V. | | Deposit date: | 2010-05-17 | | Release date: | 2010-10-20 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.716 Å) | | Cite: | Probing the Impact of the echinT C-Terminal Domain on Structure and Catalysis.
J.Mol.Biol., 404, 2010
|
|
6H6G
 
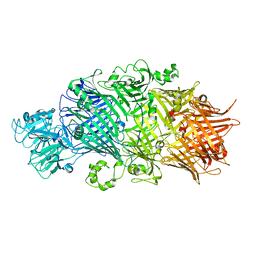 | | Crystal Structure of TcdB2-TccC3 without hypervariable C-terminal region | | Descriptor: | TcdB2,TccC3 | | Authors: | Gatsogiannis, C, Merino, F, Roderer, D, Balchin, D, Schubert, E, Kuhlee, A, Hayer-Hartl, M, Raunser, S. | | Deposit date: | 2018-07-27 | | Release date: | 2018-10-03 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (3.004 Å) | | Cite: | Tc toxin activation requires unfolding and refolding of a beta-propeller.
Nature, 563, 2018
|
|
3PJT
 
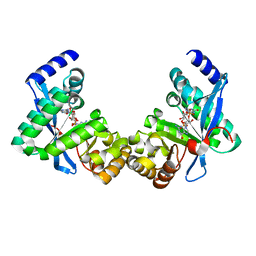 | | Structure of Pseudomonas fluorescence LapD EAL domain complexed with c-di-GMP, C2221 | | Descriptor: | 9,9'-[(2R,3R,3aS,5S,7aR,9R,10R,10aS,12S,14aR)-3,5,10,12-tetrahydroxy-5,12-dioxidooctahydro-2H,7H-difuro[3,2-d:3',2'-j][1,3,7,9,2,8]tetraoxadiphosphacyclododecine-2,9-diyl]bis(2-amino-1,9-dihydro-6H-purin-6-one), Cyclic dimeric GMP binding protein | | Authors: | Sondermann, H, Navarro, M.V.A.S, Krasteva, P. | | Deposit date: | 2010-11-10 | | Release date: | 2011-02-09 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.5154 Å) | | Cite: | Structural Basis for c-di-GMP-Mediated Inside-Out Signaling Controlling Periplasmic Proteolysis.
Plos Biol., 9, 2011
|
|
1J3S
 
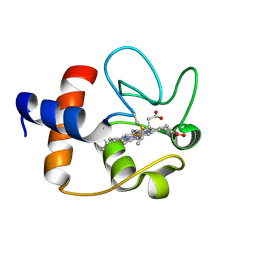 | |
3PJU
 
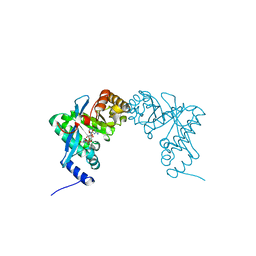 | | Structure of Pseudomonas fluorescence LapD EAL domain complexed with c-di-GMP, P6522 | | Descriptor: | 9,9'-[(2R,3R,3aS,5S,7aR,9R,10R,10aS,12S,14aR)-3,5,10,12-tetrahydroxy-5,12-dioxidooctahydro-2H,7H-difuro[3,2-d:3',2'-j][1,3,7,9,2,8]tetraoxadiphosphacyclododecine-2,9-diyl]bis(2-amino-1,9-dihydro-6H-purin-6-one), Cyclic dimeric GMP binding protein | | Authors: | Sondermann, H, Navarro, M.V.A.S, Krasteva, P. | | Deposit date: | 2010-11-10 | | Release date: | 2011-02-09 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.4991 Å) | | Cite: | Structural Basis for c-di-GMP-Mediated Inside-Out Signaling Controlling Periplasmic Proteolysis.
Plos Biol., 9, 2011
|
|
3BVD
 
 | | Structure of Surface-engineered Cytochrome ba3 Oxidase from Thermus thermophilus under Xenon Pressure, 100psi 5min | | Descriptor: | COPPER (II) ION, Cytochrome c oxidase polypeptide 2A, Cytochrome c oxidase subunit 1, ... | | Authors: | Luna, V.M, Chen, Y, Fee, J.A, Stout, C.D. | | Deposit date: | 2008-01-07 | | Release date: | 2008-05-20 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (3.37 Å) | | Cite: | Crystallographic Studies of Xe and Kr Binding within the Large Internal Cavity of Cytochrome ba3 from Thermus thermophilus: Structural Analysis and Role of Oxygen Transport Channels in the Heme-Cu Oxidases.
Biochemistry, 47, 2008
|
|
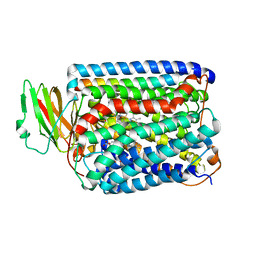
4BA6
 
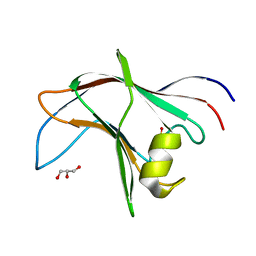 | | High Resolution structure of the C-terminal family 65 Carbohydrate Binding Module (CBM65B) of endoglucanase Cel5A from Eubacterium cellulosolvens | | Descriptor: | Endoglucanase cel5A, GLYCEROL | | Authors: | Venditto, I, Luis, A.S, Basle, A, Temple, M, Ferreira, L.M.A, Fontes, C.M.G.A, Gilbert, H.J, Najmudin, S. | | Deposit date: | 2012-09-11 | | Release date: | 2012-12-19 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.42 Å) | | Cite: | Understanding how noncatalytic carbohydrate binding modules can display specificity for xyloglucan.
J. Biol. Chem., 288, 2013
|
|
6JLB
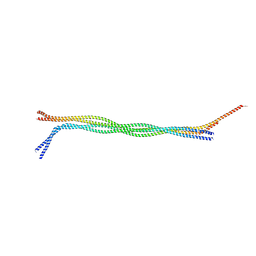 
 | |
2PX1
 
 | | crystal structure of the complex of bovine lactoferrin C-lobe with Ribose at 2.5 A resolution | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, CARBONATE ION, FE (III) ION, ... | | Authors: | Mir, R, Vikram, G, Sinha, M, Sharma, S, Kaur, P, Singh, T.P. | | Deposit date: | 2007-05-14 | | Release date: | 2007-05-29 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | crystal structure of the complex of bovine lactoferrin C-lobe with Ribose at 2.5 A resolution
To be Published
|
|
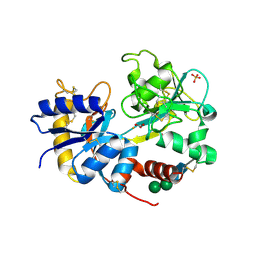
1JZC
 
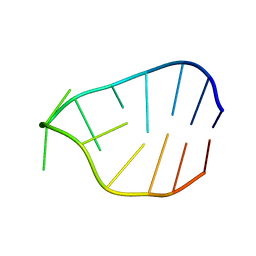 | |
1V9A
 
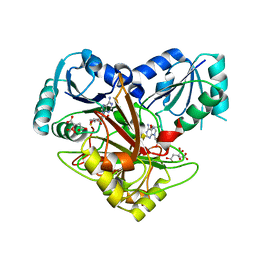 | | Crystal structure of Uroporphyrin-III C-methyl transferase from Thermus thermophilus complexed with S-adenyl homocysteine | | Descriptor: | CITRATE ANION, S-ADENOSYL-L-HOMOCYSTEINE, Uroporphyrin-III C-methyltransferase | | Authors: | Rehse, P.H, Kitao, T, Tahirov, T.H, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2004-01-23 | | Release date: | 2005-02-01 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structure of a closed-form uroporphyrinogen-III C-methyltransferase from Thermus thermophilus.
Acta Crystallogr.,Sect.D, 61, 2005
|
|
1VA0
 
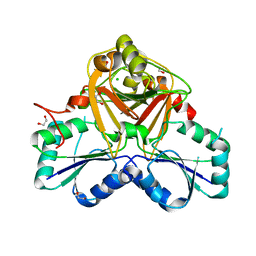 | | Crystal Structure of the Native Form of Uroporphyrin III C-methyl transferase from Thermus thermophilus | | Descriptor: | 1,2-ETHANEDIOL, CHLORIDE ION, Uroporphyrin-III C-methyltransferase | | Authors: | Rehse, P.H, Kitao, T, Tahirov, T.H, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2004-02-05 | | Release date: | 2005-02-15 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.97 Å) | | Cite: | Structure of a closed-form uroporphyrinogen-III C-methyltransferase from Thermus thermophilus.
Acta Crystallogr.,Sect.D, 61, 2005
|
|
2Q70
 
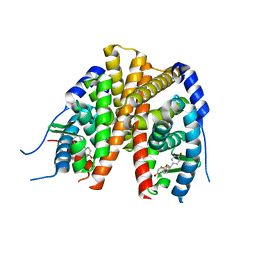 | | Estrogen receptor alpha ligand-binding domain complxed to a benzopyran ligand | | Descriptor: | (3AS,4R,9BR)-2,2-DIFLUORO-4-(4-HYDROXYPHENYL)-1,2,3,3A,4,9B-HEXAHYDROCYCLOPENTA[C]CHROMEN-8-OL, Estrogen receptor | | Authors: | Wang, Y. | | Deposit date: | 2007-06-05 | | Release date: | 2007-08-07 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Benzopyrans as selective estrogen receptor beta agonists (SERBAs). Part 3: synthesis of cyclopentanone and cyclohexanone intermediates for C-ring modification.
Bioorg.Med.Chem.Lett., 17, 2007
|
|
1B5K
 
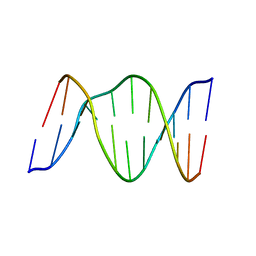 | | 3,N4-ETHENO-2'-DEOXYCYTIDINE OPPOSITE THYMIDINE IN AN 11-MER DUPLEX, SOLUTION STRUCTURE FROM NMR AND MOLECULAR DYNAMICS | | Descriptor: | DNA (5'-D(*CP*GP*TP*AP*CP*EDCP*CP*AP*TP*GP*C)-3'), DNA (5'-D(*GP*CP*AP*TP*GP*TP*GP*TP*AP*CP*G)-3') | | Authors: | Cullinan, D, Korobka, A, Grollman, A.P, Patel, D.J, Eisenberg, M, De Santos, C.L. | | Deposit date: | 1999-01-07 | | Release date: | 1999-01-13 | | Last modified: | 2023-12-27 | | Method: | SOLUTION NMR | | Cite: | NMR solution structure of an oligodeoxynucleotide duplex containing the exocyclic lesion 3,N4-etheno-2'-deoxycytidine opposite thymidine: comparison with the duplex containing deoxyadenosine opposite the adduct.
Biochemistry, 35, 1996
|
|
