4DQ4
 
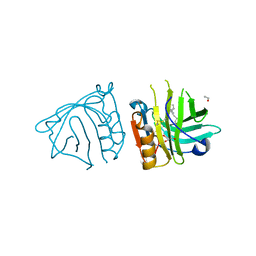 | | Bovine beta-lactoglobulin complex with linoleic acid | | Descriptor: | Beta-lactoglobulin, ETHANOL, GLYCEROL, ... | | Authors: | Loch, J.I, Lewinski, K. | | Deposit date: | 2012-02-15 | | Release date: | 2012-02-29 | | Last modified: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Binding of 18-carbon unsaturated fatty acids to bovine beta-lactoglobulin--structural and thermodynamic studies.
Int.J.Biol.Macromol., 57, 2013
|
|
4DQ3
 
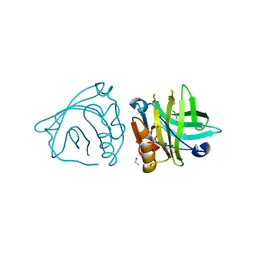 | | Bovine beta-lactoglobulin complex with oleic acid | | Descriptor: | Beta-lactoglobulin, ETHANOL, GLYCEROL, ... | | Authors: | Loch, J.I, Lewinski, K. | | Deposit date: | 2012-02-15 | | Release date: | 2012-02-29 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Binding of 18-carbon unsaturated fatty acids to bovine beta-lactoglobulin--structural and thermodynamic studies.
Int.J.Biol.Macromol., 57, 2013
|
|
2LB6
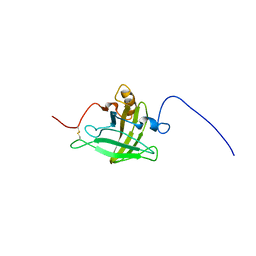 
 | | Structure of 18694Da MUP, typical to the major urinary protein family: MUP9, MUP11, MUP15, MUP18 & MUP19 | | Descriptor: | Major urinary protein 6 | | Authors: | Phelan, M.M, Mclean, L, Beynon, R.J, Hurst, J.L, Lian, L. | | Deposit date: | 2011-03-23 | | Release date: | 2012-03-28 | | Last modified: | 2023-12-06 | | Method: | SOLUTION NMR | | Cite: | Structural insights into the specificity of darcin, an atypical major urinary protein.
To be Published
|
|
3RSW
 
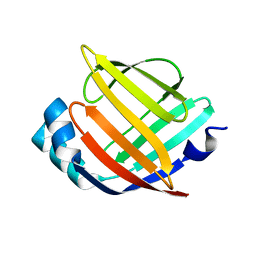 | | Crystal Structure of Heart Fatty Acid Binding Protein (FABP3) | | Descriptor: | Fatty acid-binding protein, heart | | Authors: | Carney, D.F, Shankaran, B, Zwart, P.H, Stebbins, J.W, Prasad, G.S. | | Deposit date: | 2011-05-02 | | Release date: | 2012-05-16 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.599 Å) | | Cite: | Crystal Structure of Heart Fatty Acid Binding Protein (FABP3)
To be Published
|
|
2LFO
 
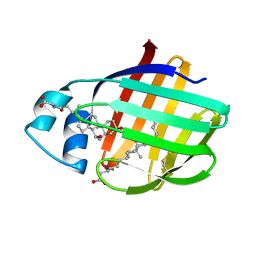 | | NMR structure of cl-BABP/SS complexed with glycochenodeoxycholic and glycocholic acids | | Descriptor: | Fatty acid-binding protein, liver, GLYCOCHENODEOXYCHOLIC ACID, ... | | Authors: | Tomaselli, S, Cogliati, C, Pagano, K, Zetta, L, Zanzoni, S, Assfalg, M, Molinari, H, Ragona, L. | | Deposit date: | 2011-07-07 | | Release date: | 2012-07-11 | | Last modified: | 2024-10-30 | | Method: | SOLUTION NMR | | Cite: | A disulfide bridge allows for site-selective binding in liver bile acid binding protein thereby stabilising the orientation of key amino acid side chains.
Chemistry, 18, 2012
|
|
4ALO
 
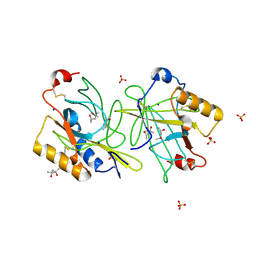 | | STRUCTURE AND PROPERTIES OF H1 CRUSTACYANIN FROM LOBSTER HOMARUS AMERICANUS | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, H1 APOCRUSTACYANIN, SODIUM ION, ... | | Authors: | Ferrari, M, Folli, C, Pincolini, E, Mcclintock, T.S, Roessle, M, Berni, R, Cianci, M. | | Deposit date: | 2012-03-05 | | Release date: | 2012-08-08 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.37 Å) | | Cite: | Structural Characterization of Recombinant Crustacyanin Subunits from the Lobster Homarus Americanus.
Acta Crystallogr.,Sect.F, 68, 2012
|
|
4A1H
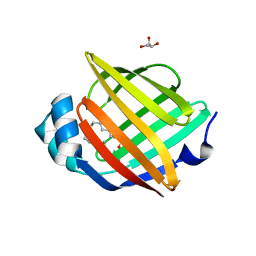 
 | | Human myelin P2 protein, K45S mutant | | Descriptor: | CHLORIDE ION, GLYCEROL, MYELIN P2 PROTEIN, ... | | Authors: | Lehtimaki, M, Kursula, P. | | Deposit date: | 2011-09-15 | | Release date: | 2012-09-26 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.201 Å) | | Cite: | Structure-Function Relationships in the Myelin Peripheral Membrane Protein P2
To be Published
|
|
4A1Y
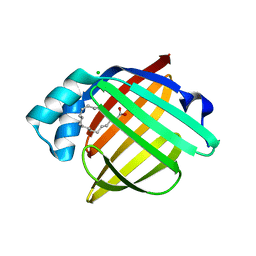 
 | | Human myelin P2 protein, K65Q mutant | | Descriptor: | CHLORIDE ION, MYELIN P2 PROTEIN, PALMITIC ACID | | Authors: | Lehtimaki, M, Kursula, P. | | Deposit date: | 2011-09-20 | | Release date: | 2012-10-10 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.2 Å) | | Cite: | Structure-Function Relationships in the Myelin Peripheral Membrane Protein P2
To be Published
|
|
4A8Z
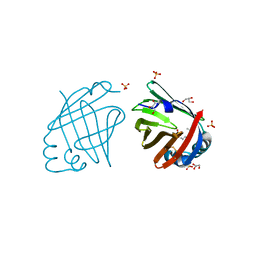 
 | |
4GKC
 | |
4GH7
 
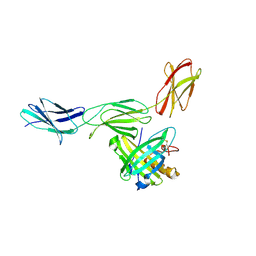 | |
4EDE
 | |
4EEJ
 | |
4EFG
 | |
4EXZ
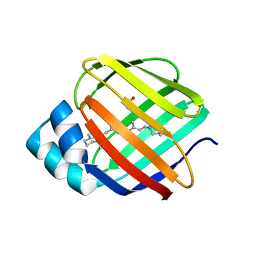 
 | |
4GNY
 
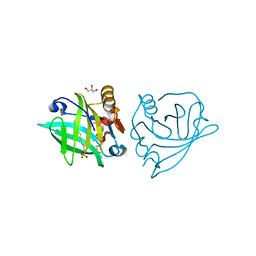 | | Bovine beta-lactoglobulin complex with dodecyl sulfate | | Descriptor: | Beta-lactoglobulin, DODECYL SULFATE, GLYCEROL | | Authors: | Gutierrez-Magdaleno, G, Torres-Rivera, A, Garcia-Hernandez, E, Rodriguez-Romero, A. | | Deposit date: | 2012-08-17 | | Release date: | 2013-02-13 | | Last modified: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (1.6367 Å) | | Cite: | Ligand-binding and self-association cooperativity of beta-lactoglobulin
J.Mol.Recognit., 26, 2013
|
|
4IMN
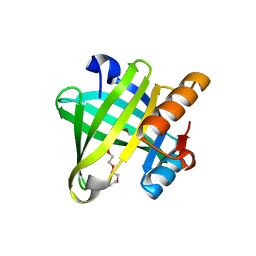 
 | | Crystal structure of wild type human Lipocalin PGDS bound with PEG MME 2000 | | Descriptor: | 2-(2-{2-[2-(2-METHOXY-ETHOXY)-ETHOXY]-ETHOXY}-ETHOXY)-ETHANOL, Lipocalin-type prostaglandin-D synthase | | Authors: | Lim, S.M, Chen, D, Teo, H, Roos, A, Nyman, T, Tresaugues, L, Pervushin, K, Nordlund, P. | | Deposit date: | 2013-01-03 | | Release date: | 2013-03-20 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.09 Å) | | Cite: | Structural and dynamic insights into substrate binding and catalysis of human lipocalin prostaglandin D synthase.
J.Lipid Res., 54, 2013
|
|
4IMO
 
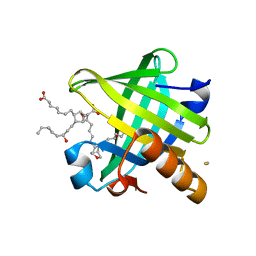 | | Crystal structure of wild type human Lipocalin PGDS in complex with substrate analog U44069 | | Descriptor: | (5E)-7-{(1R,4S,5S,6R)-5-[(1E,3S)-3-hydroxyoct-1-en-1-yl]-2-oxabicyclo[2.2.1]hept-6-yl}hept-5-enoic acid, Lipocalin-type prostaglandin-D synthase, THIOCYANATE ION | | Authors: | Lim, S.M, Chen, D, Teo, H, Roos, A, Nyman, T, Tresaugues, L, Pervushin, K, Nordlund, P. | | Deposit date: | 2013-01-03 | | Release date: | 2013-03-20 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.88 Å) | | Cite: | Structural and dynamic insights into substrate binding and catalysis of human lipocalin prostaglandin D synthase.
J.Lipid Res., 54, 2013
|
|
4ES7
 
 | |
3U9P
 
 | | Crystal Structure of Murine Siderocalin in Complex with an Fab Fragment | | Descriptor: | Monoclonal Fab Fragment Heavy Chain, Monoclonal Fab Fragment Light Chain, Neutrophil gelatinase-associated lipocalin | | Authors: | Correnti, C, Strong, R.K. | | Deposit date: | 2011-10-19 | | Release date: | 2013-05-01 | | Last modified: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Siderocalin/Lcn2/NGAL/24p3 does not drive apoptosis through gentisic acid mediated iron withdrawal in hematopoietic cell lines.
Plos One, 7, 2012
|
|
4I3C
 
 | | Crystal structure of fluorescent protein UnaG N57Q mutant | | Descriptor: | 3-[5-[(Z)-(4-ethenyl-3-methyl-5-oxidanylidene-pyrrol-2-ylidene)methyl]-2-[[5-[(Z)-(3-ethenyl-4-methyl-5-oxidanylidene-pyrrol-2-ylidene)methyl]-3-(3-hydroxy-3-oxopropyl)-4-methyl-1H-pyrrol-2-yl]methyl]-4-methyl-1H-pyrrol-3-yl]propanoic acid, Bilirubin-inducible fluorescent protein UnaG | | Authors: | Kumagai, A, Ando, R, Miyatake, H, Miyawaki, A. | | Deposit date: | 2012-11-26 | | Release date: | 2013-06-19 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.001 Å) | | Cite: | A bilirubin-inducible fluorescent protein from eel muscle
Cell(Cambridge,Mass.), 153, 2013
|
|
4I3B
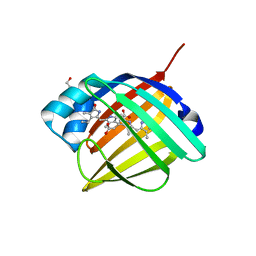 
 | | Crystal structure of fluorescent protein UnaG wild type | | Descriptor: | 3-[5-[(Z)-(4-ethenyl-3-methyl-5-oxidanylidene-pyrrol-2-ylidene)methyl]-2-[[5-[(Z)-(3-ethenyl-4-methyl-5-oxidanylidene-pyrrol-2-ylidene)methyl]-3-(3-hydroxy-3-oxopropyl)-4-methyl-1H-pyrrol-2-yl]methyl]-4-methyl-1H-pyrrol-3-yl]propanoic acid, Bilirubin-inducible fluorescent protein UnaG, DI(HYDROXYETHYL)ETHER | | Authors: | Kumagai, A, Ando, R, Miyatake, H, Miyawaki, A. | | Deposit date: | 2012-11-26 | | Release date: | 2013-06-19 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.199 Å) | | Cite: | A bilirubin-inducible fluorescent protein from eel muscle
Cell(Cambridge,Mass.), 153, 2013
|
|
4IAX
 
 | | Engineered human lipocalin 2 (CL31) in complex with Y-DTPA | | Descriptor: | N-{(1S,2S)-2-[bis(carboxymethyl)amino]cyclohexyl}-N-{(2R)-2-[bis(carboxymethyl)amino]-3-[4-({[2-hydroxy-1,1-bis(hydroxymethyl)ethyl]carbamothioyl}amino)phenyl]propyl}glycine, Neutrophil gelatinase-associated lipocalin, YTTRIUM (III) ION | | Authors: | Eichinger, A, Skerra, A. | | Deposit date: | 2012-12-07 | | Release date: | 2013-06-19 | | Last modified: | 2024-10-09 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Structure-guided engineering of Anticalins with improved binding behavior and biochemical characteristics for application in radio-immuno imaging and/or therapy
J.Struct.Biol., 185, 2014
|
|
4I3D
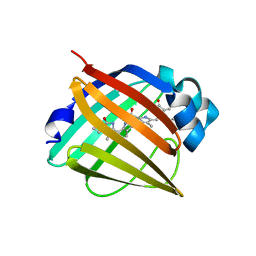 
 | | Crystal structure of fluorescent protein UnaG N57A mutant | | Descriptor: | 3-[5-[(Z)-(4-ethenyl-3-methyl-5-oxidanylidene-pyrrol-2-ylidene)methyl]-2-[[5-[(Z)-(3-ethenyl-4-methyl-5-oxidanylidene-pyrrol-2-ylidene)methyl]-3-(3-hydroxy-3-oxopropyl)-4-methyl-1H-pyrrol-2-yl]methyl]-4-methyl-1H-pyrrol-3-yl]propanoic acid, Bilirubin-inducible fluorescent protein UnaG | | Authors: | Kumagai, A, Ando, R, Miyatake, H, Miyawaki, A. | | Deposit date: | 2012-11-26 | | Release date: | 2013-06-19 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.298 Å) | | Cite: | A bilirubin-inducible fluorescent protein from eel muscle
Cell(Cambridge,Mass.), 153, 2013
|
|
4IAW
 
 | | Engineered human lipocalin 2 (C26) in complex with Y-DTPA | | Descriptor: | N-{(1S,2S)-2-[bis(carboxymethyl)amino]cyclohexyl}-N-{(2R)-2-[bis(carboxymethyl)amino]-3-[4-({[2-hydroxy-1,1-bis(hydroxymethyl)ethyl]carbamothioyl}amino)phenyl]propyl}glycine, Neutrophil gelatinase-associated lipocalin, YTTRIUM (III) ION | | Authors: | Eichinger, A, Skerra, A. | | Deposit date: | 2012-12-07 | | Release date: | 2013-06-19 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Structure-guided engineering of Anticalins with improved binding behavior and biochemical characteristics for application in radio-immuno imaging and/or therapy
J.Struct.Biol., 185, 2014
|
|
