7ECF
 
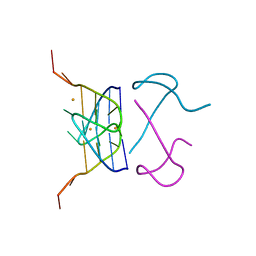 | | Crystal Structure of d(G4C2)2-Ba in C2221 space group | | Descriptor: | BARIUM ION, DNA (5'-D(*GP*GP*GP*GP*CP*CP*GP*GP*GP*GP*CP*C)-3') | | Authors: | Geng, Y, Liu, C, Cai, Q, Luo, Z, Zhu, G. | | Deposit date: | 2021-03-12 | | Release date: | 2021-06-09 | | Last modified: | 2024-03-27 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Crystal structure of parallel G-quadruplex formed by the two-repeat ALS- and FTD-related GGGGCC sequence.
Nucleic Acids Res., 49, 2021
|
|
6AA7
 
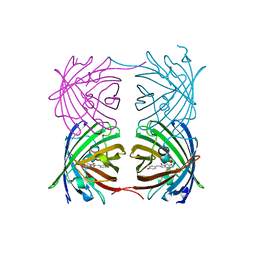 | |
2RH4
 
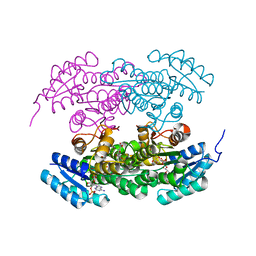 | | Actinorhodin ketoreductase, actKR, with NADPH and Inhibitor Emodin | | Descriptor: | 3-METHYL-1,6,8-TRIHYDROXYANTHRAQUINONE, Actinorhodin Polyketide Ketoreductase, NADPH DIHYDRO-NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE | | Authors: | Korman, T.P, Tsai, S.-C. | | Deposit date: | 2007-10-05 | | Release date: | 2008-08-19 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Inhibition kinetics and emodin cocrystal structure of a type II polyketide ketoreductase
Biochemistry, 47, 2008
|
|
2RHC
 
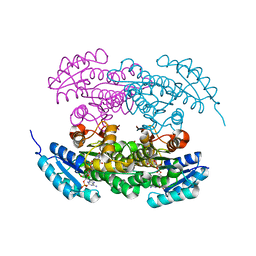 | | Actinorhodin ketordeuctase, actKR, with NADP+ and Inhibitor Emodin | | Descriptor: | 3-METHYL-1,6,8-TRIHYDROXYANTHRAQUINONE, Actinorhodin Polyketide Ketoreductase, NADP NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE | | Authors: | Korman, T.P, Tsai, S.-C. | | Deposit date: | 2007-10-08 | | Release date: | 2008-08-19 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Inhibition kinetics and emodin cocrystal structure of a type II polyketide ketoreductase
Biochemistry, 47, 2008
|
|
3EIV
 
 | |
3RRO
 
 | | Crystal structure of 3-ketoacyl-(acyl-carrier-protein) reductase (FabG) from Vibrio cholerae | | Descriptor: | 1,2-ETHANEDIOL, 3-ketoacyl-(acyl-carrier-protein) reductase, CHLORIDE ION, ... | | Authors: | Hou, J, Chruszcz, M, Cooper, D.R, Grabowski, M, Zheng, H, Osinski, T, Shumilin, I, Anderson, W, Minor, W, Center for Structural Genomics of Infectious Diseases (CSGID) | | Deposit date: | 2011-04-29 | | Release date: | 2011-05-18 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Dissecting the Structural Elements for the Activation of beta-Ketoacyl-(Acyl Carrier Protein) Reductase from Vibrio cholerae.
J.Bacteriol., 198, 2015
|
|
4UR1
 
 | | Crystal structure of the PCE reductive dehalogenase from S. multivorans in complex with dibromoethene | | Descriptor: | BENZAMIDINE, CIS-DIBROMOETHENE, GLYCEROL, ... | | Authors: | Bommer, M, Kunze, C, Fesseler, J, Schubert, T, Diekert, G, Dobbek, H. | | Deposit date: | 2014-06-25 | | Release date: | 2014-10-15 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (1.649 Å) | | Cite: | Structural Basis for Organohalide Respiration.
Science, 346, 2014
|
|
4UR3
 
 | | Crystal structure of the PCE reductive dehalogenase from S. multivorans P2(1) crystal form | | Descriptor: | IRON/SULFUR CLUSTER, NORPSEUDO-B12, TETRACHLOROETHENE REDUCTIVE DEHALOGENASE CATALYTIC SUBUNIT | | Authors: | Bommer, M, Kunze, C, Fesseler, J, Schubert, T, Diekert, G, Dobbek, H. | | Deposit date: | 2014-06-25 | | Release date: | 2014-10-15 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2.235 Å) | | Cite: | Structural Basis for Organohalide Respiration.
Science, 346, 2014
|
|
4UR2
 
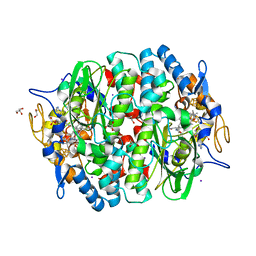 | | Crystal structure of the PCE reductive dehalogenase from S. multivorans in complex with iodide | | Descriptor: | GLYCEROL, IODIDE ION, IRON/SULFUR CLUSTER, ... | | Authors: | Bommer, M, Kunze, C, Fesseler, J, Schubert, T, Diekert, G, Dobbek, H. | | Deposit date: | 2014-06-25 | | Release date: | 2014-10-15 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2.096 Å) | | Cite: | Structural Basis for Organohalide Respiration.
Science, 346, 2014
|
|
4UR0
 
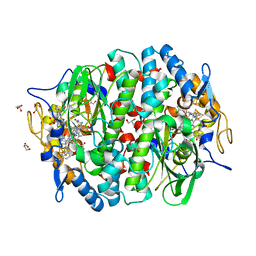 | | Crystal structure of the PCE reductive dehalogenase from S. multivorans in complex with trichloroethene | | Descriptor: | 1,1,2-trichloroethene, BENZAMIDINE, GLYCEROL, ... | | Authors: | Bommer, M, Kunze, C, Fesseler, J, Schubert, T, Diekert, G, Dobbek, H. | | Deposit date: | 2014-06-25 | | Release date: | 2014-10-15 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (1.798 Å) | | Cite: | Structural Basis for Organohalide Respiration.
Science, 346, 2014
|
|
4I4J
 
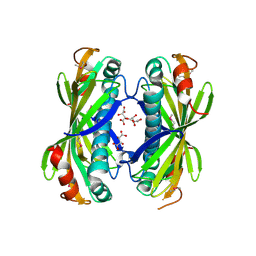 | | The structure of SgcE10, the ACP-polyene thioesterase involved in C-1027 biosynthesis | | Descriptor: | 1,2-ETHANEDIOL, ACP-polyene thioesterase, D(-)-TARTARIC ACID, ... | | Authors: | Kim, Y, Bigelow, L, Bearden, J, Babnigg, J, Bingman, C.A, Yennamalli, R, Lohman, J, Ma, M, Shen, B, Phillips Jr, G.N, Joachimiak, A, Midwest Center for Structural Genomics (MCSG), Enzyme Discovery for Natural Product Biosynthesis (NatPro) | | Deposit date: | 2012-11-27 | | Release date: | 2012-12-12 | | Last modified: | 2022-05-04 | | Method: | X-RAY DIFFRACTION (2.784 Å) | | Cite: | Crystal Structure of Thioesterase SgcE10 Supporting Common Polyene Intermediates in 9- and 10-Membered Enediyne Core Biosynthesis.
Acs Omega, 2, 2017
|
|
4L8Q
 
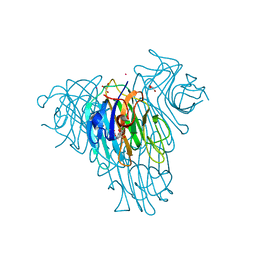 | | Crystal structure of Canavalia grandiflora seed lectin complexed with X-Man. | | Descriptor: | 5-bromo-4-chloro-1H-indol-3-yl alpha-D-mannopyranoside, CADMIUM ION, CALCIUM ION, ... | | Authors: | Barroso-Neto, I.L, Rocha, B.A.M, Simoes, R.C, Bezerra, M.J.B, Pereira-Junior, F.N, Osterne, V.J.S, Nascimento, K.S, Nagano, C.S, Delatorre, P, Sampaio, A.H, Cavada, B.S. | | Deposit date: | 2013-06-17 | | Release date: | 2014-05-21 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Vasorelaxant activity of Canavalia grandiflora seed lectin: A structural analysis.
Arch.Biochem.Biophys., 543, 2014
|
|
4CR8
 
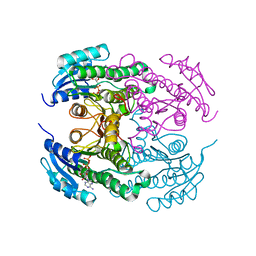 | | Crystal structure of the N-acetyl-D-mannosamine dehydrogenase with NAD | | Descriptor: | N-ACYLMANNOSAMINE 1-DEHYDROGENASE, NICOTINAMIDE-ADENINE-DINUCLEOTIDE | | Authors: | Gil-Ortiz, F, Sola-Carvajal, A, Garcia-Carmona, F, Sanchez-Ferrer, A, Rubio, V. | | Deposit date: | 2014-02-25 | | Release date: | 2014-07-09 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Crystal Structures and Functional Studies Clarify Substrate Selectivity and Catalytic Residues for the Unique Orphan Enzyme N-Acetyl-D-Mannosamine Dehydrogenase.
Biochem.J., 462, 2014
|
|
6SW5
 
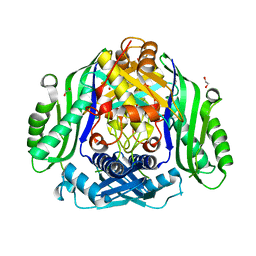 | | Crystal structure of the human S-adenosylmethionine synthetase 1 (ligand-free form) | | Descriptor: | 1,2-ETHANEDIOL, DI(HYDROXYETHYL)ETHER, S-adenosylmethionine synthase isoform type-1 | | Authors: | Panmanee, J, Antoyuk, S.V, Hasnain, S.S. | | Deposit date: | 2019-09-19 | | Release date: | 2020-06-17 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (2.35 Å) | | Cite: | Structural basis of the dominant inheritance of hypermethioninemia associated with the Arg264His mutation in the MAT1A gene.
Acta Crystallogr D Struct Biol, 76, 2020
|
|
4CAZ
 
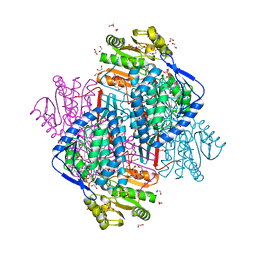 | | CRYSTAL STRUCTURE OF BETAINE ALDEHYDE DEHYDROGENASE FROM Pseudomonas aeruginosa IN COMPLEX WITH NADH | | Descriptor: | 1,2-ETHANEDIOL, 2-{2-[2-(2-{2-[2-(2-ETHOXY-ETHOXY)-ETHOXY]-ETHOXY}-ETHOXY)-ETHOXY]-ETHOXY}-ETHANOL, BETAINE ALDEHYDE DEHYDROGENASE, ... | | Authors: | Gonzalez-Segura, L, Diaz-Sanchez, A.G, Rodriguez-Sotres, R, Mujica-Jimenez, C, Munoz-Clares, R.A. | | Deposit date: | 2013-10-09 | | Release date: | 2014-10-29 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.55 Å) | | Cite: | The Structural Bases of the Dual Coenzyme Specificity of Betaine Aldehyde Dehydrogenase from Pseudomonas Aeruginosa
To be Published
|
|
2W7E
 
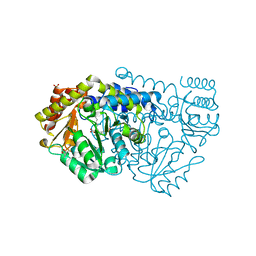 | | Crystal structure of Y51FbsSHMT obtained in the presence of Glycine | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, GLYCINE, PHOSPHATE ION, ... | | Authors: | Rajaram, V, Bhavani, B.S, Bisht, S, Kaul, P, Prakash, V, Appaji Rao, N, Savithri, H.S, Murthy, M.R.N. | | Deposit date: | 2008-12-22 | | Release date: | 2010-08-18 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.69 Å) | | Cite: | Importance of Tyrosine Residues of Bacillus Stearothermophilus Serine Hydroxymethyltransferase in Cofactor Binding and L-Allo-Thr Cleavage.
FEBS J., 275, 2008
|
|
5C8A
 
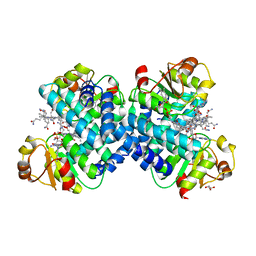 | |
4CR6
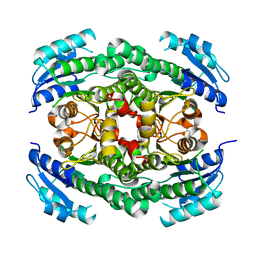 
 | | Crystal structure of the N-acetyl-D-mannosamine dehydrogenase without substrates | | Descriptor: | N-ACYLMANNOSAMINE 1-DEHYDROGENASE, alpha-D-mannopyranose | | Authors: | Gil-Ortiz, F, Sola-Carvajal, A, Garcia-Carmona, F, Sanchez-Ferrer, A, Rubio, V. | | Deposit date: | 2014-02-25 | | Release date: | 2014-07-09 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Crystal Structures and Functional Studies Clarify Substrate Selectivity and Catalytic Residues for the Unique Orphan Enzyme N-Acetyl-D-Mannosamine Dehydrogenase.
Biochem.J., 462, 2014
|
|
6T62
 
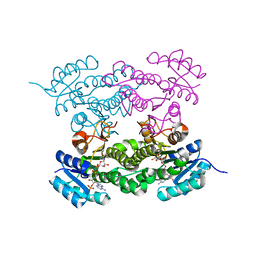 | |
5F2V
 
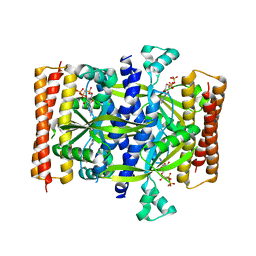 | | Crystal structure of the small alarmone synthethase 1 from Bacillus subtilis bound to AMPCPP | | Descriptor: | DIPHOSPHOMETHYLPHOSPHONIC ACID ADENOSYL ESTER, GTP pyrophosphokinase YjbM, MAGNESIUM ION | | Authors: | Steinchen, W, Schuhmacher, J.S, Altegoer, F, Bange, G. | | Deposit date: | 2015-12-02 | | Release date: | 2015-12-16 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Catalytic mechanism and allosteric regulation of an oligomeric (p)ppGpp synthetase by an alarmone.
Proc.Natl.Acad.Sci.USA, 112, 2015
|
|
2W7D
 
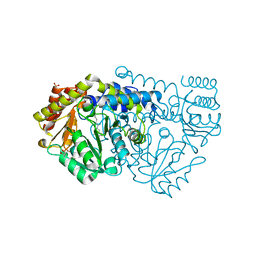 | | Crystal structure of Y51FbsSHMT internal aldimine | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, PHOSPHATE ION, PYRIDOXAL-5'-PHOSPHATE, ... | | Authors: | Rajaram, V, Bhavani, B.S, Bisht, S, Kaul, P, Prakash, V, Appaji Rao, N, Savithri, H.S, Murthy, M.R.N. | | Deposit date: | 2008-12-22 | | Release date: | 2010-08-18 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Importance of Tyrosine Residues of Bacillus Stearothermophilus Serine Hydroxymethyltransferase in Cofactor Binding and L-Allo-Thr Cleavage.
FEBS J., 275, 2008
|
|
6T6P
 
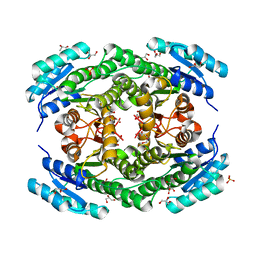 | | Crystal structure of Klebsiella pneumoniae FabG2(NADH-dependent) at 1.57 A resolution | | Descriptor: | 3-oxoacyl-[acyl-carrier protein] reductase, GLYCEROL, PHOSPHATE ION | | Authors: | Vella, P, Schnell, R, Lindqvist, Y, Schneider, G. | | Deposit date: | 2019-10-18 | | Release date: | 2020-11-18 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (1.57 Å) | | Cite: | A FabG inhibitor targeting an allosteric binding site inhibits several orthologs from Gram-negative ESKAPE pathogens.
Bioorg.Med.Chem., 30, 2021
|
|
3U34
 
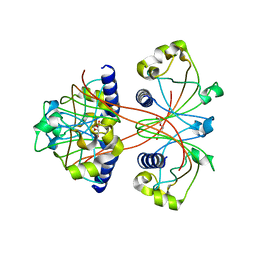 | |
6JVB
 
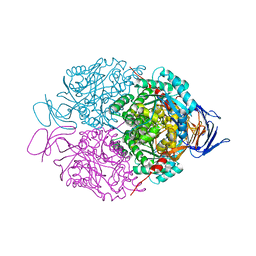 | |
6TJN
 
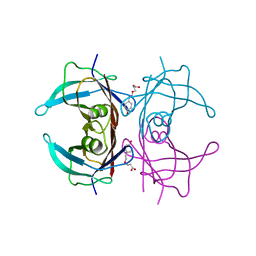 | | Human transthyretin (TTR) complexed with (E)-3-(((4-hydroxybenzylidene)amino)oxy)propanoic acid | | Descriptor: | 3-[(~{E})-(4-hydroxyphenyl)methylideneamino]oxypropanoic acid, Transthyretin | | Authors: | Ciccone, L, Shepard, W, Nencetti, S, Orlandini, E, Rossello, A. | | Deposit date: | 2019-11-26 | | Release date: | 2020-12-16 | | Last modified: | 2024-01-24 | | Method: | X-RAY DIFFRACTION (1.702 Å) | | Cite: | Monoaryl derivatives as transthyretin fibril formation inhibitors: Design, synthesis, biological evaluation and structural analysis.
Bioorg.Med.Chem., 28, 2020
|
|
