1I87
 
 | | SOLUTION STRUCTURE OF THE WATER-SOLUBLE FRAGMENT OF RAT HEPATIC APOCYTOCHROME B5 | | Descriptor: | CYTOCHROME B5 | | Authors: | Falzone, C.J, Wang, Y, Vu, B.C, Scott, N.L, Bhattacharya, S, Lecomte, J.T. | | Deposit date: | 2001-03-12 | | Release date: | 2001-05-16 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Structural and dynamic perturbations induced by heme binding in cytochrome b5.
Biochemistry, 40, 2001
|
|
4EKB
 
 | | Initial Thaumatin Structure for Radiation Damage Experiment at 100 K | | Descriptor: | L(+)-TARTARIC ACID, Thaumatin-1 | | Authors: | Warkentin, M, Badeau, R, Hopkins, J.B, Thorne, R.E. | | Deposit date: | 2012-04-09 | | Release date: | 2012-08-29 | | Last modified: | 2013-01-23 | | Method: | X-RAY DIFFRACTION (1.52 Å) | | Cite: | Spatial distribution of radiation damage to crystalline proteins at 25-300 K.
Acta Crystallogr.,Sect.D, 68, 2012
|
|
1IXT
 | | Structure of a Novel P-Superfamily Spasmodic Conotoxin Reveals an Inhibitory Cystine Knot Motif | | Descriptor: | spasmodic protein tx9a-like protein | | Authors: | Miles, L.A, Dy, C.Y, Nielsen, J, Barnham, K.J, Hinds, M.G, Olivera, B.M, Bulaj, G, Norton, R.S. | | Deposit date: | 2002-07-04 | | Release date: | 2003-01-28 | | Last modified: | 2023-12-27 | | Method: | SOLUTION NMR | | Cite: | Structure of a Novel P-Superfamily Spasmodic Conotoxin Reveals an Inhibitory Cystine Knot Motif
J.Biol.Chem., 277, 2002
|
|
4EL2
 
 | | Initial Thaumatin Structure for Radiation Damage Experiment at 240 K | | Descriptor: | L(+)-TARTARIC ACID, Thaumatin-1 | | Authors: | Warkentin, M, Badeau, R, Hopkins, J.B, Thorne, R.E. | | Deposit date: | 2012-04-10 | | Release date: | 2012-08-29 | | Last modified: | 2024-10-09 | | Method: | X-RAY DIFFRACTION (1.52 Å) | | Cite: | Spatial distribution of radiation damage to crystalline proteins at 25-300 K.
Acta Crystallogr.,Sect.D, 68, 2012
|
|
2L28
 
 | | Solution structure of lactobacillus casei dihydrofolate reductase apo-form, 25 conformers | | Descriptor: | Dihydrofolate reductase | | Authors: | Polshakov, V.I, Birdsall, B, Feeney, J. | | Deposit date: | 2010-08-13 | | Release date: | 2011-04-06 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | NMR Structures of Apo L. casei Dihydrofolate Reductase and Its Complexes with Trimethoprim and NADPH: Contributions to Positive Cooperative Binding from Ligand-Induced Refolding, Conformational Changes, and Interligand Hydrophobic Interactions.
Biochemistry, 50, 2011
|
|
4EKO
 
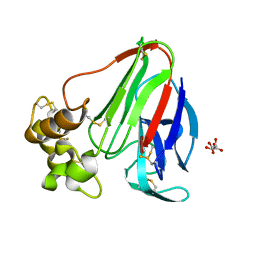 | | Initial Thaumatin Structure for Radiation Damage Experiment at 180 K | | Descriptor: | L(+)-TARTARIC ACID, Thaumatin-1 | | Authors: | Warkentin, M, Badeau, R, Hopkins, J.B, Thorne, R.E. | | Deposit date: | 2012-04-09 | | Release date: | 2012-08-29 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (1.52 Å) | | Cite: | Spatial distribution of radiation damage to crystalline proteins at 25-300 K.
Acta Crystallogr.,Sect.D, 68, 2012
|
|
4EPB
 
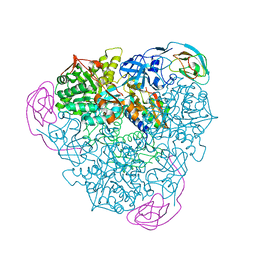 | | Final Urease Structure for Radiation Damage Experiment at 100 K | | Descriptor: | NICKEL (II) ION, Urease subunit alpha, Urease subunit beta, ... | | Authors: | Warkentin, M, Badeau, R, Hopkins, J.B, Thorne, R.E. | | Deposit date: | 2012-04-17 | | Release date: | 2012-08-29 | | Last modified: | 2013-01-23 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Spatial distribution of radiation damage to crystalline proteins at 25-300 K.
Acta Crystallogr.,Sect.D, 68, 2012
|
|
1WT1
 
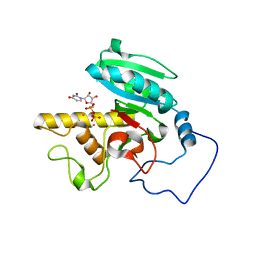 | | Mutant ABO(H) blood group glycosyltransferase with bound UDP and acceptor | | Descriptor: | Histo-blood group ABO system transferase, MERCURY (II) ION, URIDINE-5'-DIPHOSPHATE, ... | | Authors: | Lee, H.J, Barry, C.H, Borisova, S.N, Seto, N.O.L, Zheng, R.B, Blancher, A, Evans, S.V, Palcic, M.M. | | Deposit date: | 2004-11-12 | | Release date: | 2004-12-07 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (1.55 Å) | | Cite: | Structural basis for the inactivity of human blood group o2 glycosyltransferase
J.Biol.Chem., 280, 2005
|
|
1X41
 
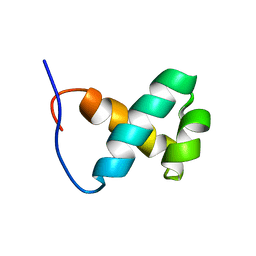 | | Solution structure of the Myb-like DNA binding domain of human Transcriptional adaptor 2-like, isoform B | | Descriptor: | Transcriptional adaptor 2-like, isoform b | | Authors: | Sasagawa, A, Sato, M, Koshiba, S, Inoue, M, Kigawa, T, Yokoyama, S, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 2005-05-12 | | Release date: | 2005-11-12 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Solution structure of the Myb-like DNA binding domain of human Transcriptional adaptor 2-like, isoform B
To be Published
|
|
1WT2
 
 | | Mutant human ABO(H) blood group glycosyltransferase A with bound UDP and inhibitor | | Descriptor: | Histo-blood group ABO system transferase, MERCURY (II) ION, URIDINE-5'-DIPHOSPHATE, ... | | Authors: | Lee, H.J, Barry, C.H, Borisova, S.N, Seto, N.O.L, Zheng, R.B, Blancher, A, Evans, S.V, Palcic, M.M. | | Deposit date: | 2004-11-12 | | Release date: | 2004-12-07 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Structural basis for the inactivity of human blood group o2 glycosyltransferase
J.Biol.Chem., 280, 2005
|
|
1NTR
 
 | | SOLUTION STRUCTURE OF THE N-TERMINAL RECEIVER DOMAIN OF NTRC | | Descriptor: | NTRC RECEIVER DOMAIN | | Authors: | Volkman, B.F, Nohaile, M.J, Amy, N.K, Kustu, S, Wemmer, D.E. | | Deposit date: | 1994-09-16 | | Release date: | 1995-01-26 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Three-dimensional solution structure of the N-terminal receiver domain of NTRC.
Biochemistry, 34, 1995
|
|
4G1H
 | |
3P2T
 
 | | Crystal Structure of Leukocyte Ig-like Receptor LILRB4 (ILT3/LIR-5/CD85k) | | Descriptor: | Leukocyte immunoglobulin-like receptor subfamily B member 4, SULFATE ION | | Authors: | Chen, Y, Nam, G, Cheng, H, Zhang, J.H, Willcox, B.E, Gao, G.F. | | Deposit date: | 2010-10-04 | | Release date: | 2011-03-30 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (1.699 Å) | | Cite: | Crystal structure of leukocyte Ig-like receptor LILRB4 (ILT3/LIR-5/CD85k): a myeloid inhibitory receptor involved in immune tolerance
J.Biol.Chem., 286, 2011
|
|
1WSZ
 
 | | Mutant human ABO(H) blood group transferase A | | Descriptor: | Histo-blood group ABO system transferase, MERCURY (II) ION | | Authors: | Lee, H.J, Barry, C.H, Borisova, S.N, Seto, N.O.L, Zheng, R.B, Blancher, A, Evans, S.V, Palcic, M.M. | | Deposit date: | 2004-11-12 | | Release date: | 2004-12-07 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (1.59 Å) | | Cite: | Structural basis for the inactivity of human blood group o2 glycosyltransferase
J.Biol.Chem., 280, 2005
|
|
1WT3
 
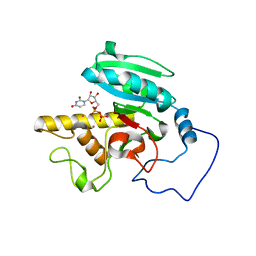 | | Mutant human ABO(H) blood group glycosyltransferase with bound UDP and acceptor | | Descriptor: | Histo-blood group ABO system transferase, MERCURY (II) ION, URIDINE-5'-DIPHOSPHATE, ... | | Authors: | Lee, H.J, Barry, C.H, Borisova, S.N, Seto, N.O.L, Zheng, R.B, Blancher, A, Evans, S.V, Palcic, M.M. | | Deposit date: | 2004-11-12 | | Release date: | 2004-12-03 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Structural basis for the inactivity of human blood group o2 glycosyltransferase
J.Biol.Chem., 280, 2005
|
|
1OZO
 
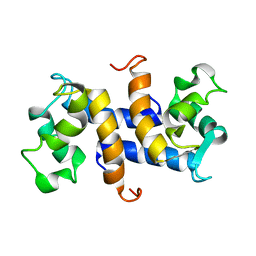 | | Three-dimensional solution structure of apo-S100P protein determined by NMR spectroscopy | | Descriptor: | S-100P protein | | Authors: | Lee, Y.-C, Volk, D.E, Thiviyanathan, V, Kleerekoper, Q, Gribenko, A.V, Zhang, S, Gorenstein, D.G, Makhatadze, G.I, Luxon, B.A. | | Deposit date: | 2003-04-09 | | Release date: | 2004-04-20 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | NMR structure of the Apo-S100P protein.
J.Biomol.Nmr, 29, 2004
|
|
1JLI
 
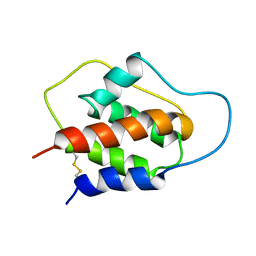 | |
3QAC
 
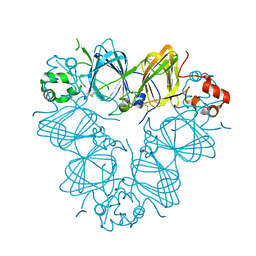 | | Structure of amaranth 11S proglobulin seed storage protein from Amaranthus hypochondriacus L. | | Descriptor: | 11S globulin seed storage protein | | Authors: | Tandang-Silvas, M.R, Carrazco-Pena, L, Barba de la Rosa, A.P, Osuna-Castro, J.A, Utsumi, S, Mikami, B, Maruyama, N. | | Deposit date: | 2011-01-10 | | Release date: | 2012-01-11 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.275 Å) | | Cite: | Structure of amaranth 11S proglobulin, a major seed storage protein from Amaranthus hypochondriacus L.
To be Published
|
|
3P0Q
 
 | | Human Tankyrase 2 - Catalytic PARP domain in complex with an inhibitor | | Descriptor: | N-[2-(4-chlorophenyl)ethyl]-6-methyl[1,2,4]triazolo[4,3-b]pyridazin-8-amine, SODIUM ION, SULFATE ION, ... | | Authors: | Karlberg, T, Siponen, M.I, Arrowsmith, C.H, Berglund, H, Bountra, C, Collins, R, Edwards, A.M, Flodin, S, Flores, A, Graslund, S, Hammarstrom, M, Johansson, I, Kotenyova, T, Kouznetsova, E, Moche, M, Nordlund, P, Nyman, T, Persson, C, Schutz, P, Sehic, A, Thorsell, A.G, Tresaugues, L, Van Den Berg, S, Wahlberg, E, Weigelt, J, Welin, M, Schuler, H, Structural Genomics Consortium (SGC) | | Deposit date: | 2010-09-29 | | Release date: | 2010-10-20 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Family-wide chemical profiling and structural analysis of PARP and tankyrase inhibitors
Nat.Biotechnol., 30, 2012
|
|
2LZX
 
 | |
1DWU
 
 | | Ribosomal protein L1 | | Descriptor: | RIBOSOMAL PROTEIN L1 | | Authors: | Tishchenko, S.V, Nevskaya, N.A, Pavelyev, M.N, Nikonov, S.V, Garber, M.B, Piendl, W. | | Deposit date: | 1999-12-13 | | Release date: | 2000-12-07 | | Last modified: | 2023-12-06 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Structure of Ribosomal Protein L1 from Methanococcus Thermolithotrophicus. Functionally Important Structural Invariants on the L1 Surface
Acta Crystallogr.,Sect.D, 58, 2002
|
|
1VPP
 
 | | COMPLEX BETWEEN VEGF AND A RECEPTOR BLOCKING PEPTIDE | | Descriptor: | PROTEIN (PEPTIDE V108), PROTEIN (VASCULAR ENDOTHELIAL GROWTH FACTOR) | | Authors: | Wiesmann, C, Christinger, H.W, Cochran, A.G, Cunningham, B.C, Fairbrother, W.J, Keenan, C.J, Meng, G, de Vos, A.M. | | Deposit date: | 1998-10-09 | | Release date: | 1999-02-23 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Crystal structure of the complex between VEGF and a receptor-blocking peptide.
Biochemistry, 37, 1998
|
|
4DAM
 
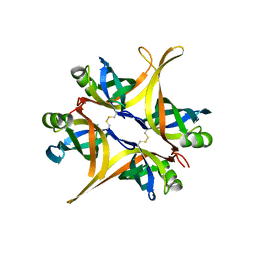 | | Crystal structure of small single-stranded DNA-binding protein from Streptomyces coelicolor | | Descriptor: | Single-stranded DNA-binding protein 1 | | Authors: | Filic, Z, Herron, P, Ivic, N, Luic, M, Manjasetty, B.A, Paradzik, T, Vujaklija, D. | | Deposit date: | 2012-01-13 | | Release date: | 2013-01-16 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Structure-function relationships of two paralogous single-stranded DNA-binding proteins from Streptomyces coelicolor: implication of SsbB in chromosome segregation during sporulation.
Nucleic Acids Res., 41, 2013
|
|
4DEG
 
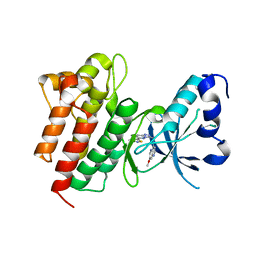 | | Crystal structure of c-Met in complex with triazolopyridazine inhibitor 2 | | Descriptor: | 7-methoxy-N-{[6-(3-methyl-1,2-thiazol-5-yl)[1,2,4]triazolo[4,3-b]pyridazin-3-yl]methyl}-1,5-naphthyridin-4-amine, Hepatocyte growth factor receptor | | Authors: | Whittington, D.A, Bellon, S.F, Long, A.M. | | Deposit date: | 2012-01-20 | | Release date: | 2012-05-30 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Discovery and optimization of a potent and selective triazolopyridinone series of c-Met inhibitors.
Bioorg.Med.Chem.Lett., 22, 2012
|
|
2O5B
 
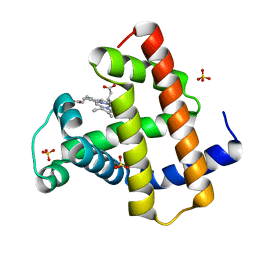 | | Manganese horse heart myoglobin, reduced | | Descriptor: | MANGANESE PROTOPORPHYRIN IX, Myoglobin, SULFATE ION | | Authors: | Richter-Addo, G.B, Zahran, Z.N, Chooback, L, Copeland, D.M, West, A.H. | | Deposit date: | 2006-12-05 | | Release date: | 2007-10-16 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Crystal structures of manganese- and cobalt-substituted myoglobin in complex with NO and nitrite reveal unusual ligand conformations.
J.Inorg.Biochem., 102, 2008
|
|
