6NM7
 
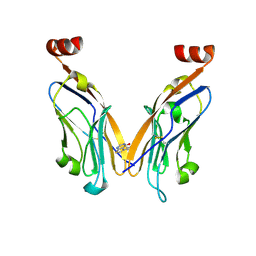 | | PD-L1 IgV domain bound to fragment | | Descriptor: | 5-phenylthieno[2,3-d]pyrimidin-4(3H)-one, Programmed cell death 1 ligand 1 | | Authors: | Perry, E, Zhao, B. | | Deposit date: | 2019-01-10 | | Release date: | 2019-02-20 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (2.426 Å) | | Cite: | Fragment-based screening of programmed death ligand 1 (PD-L1).
Bioorg. Med. Chem. Lett., 29, 2019
|
|
4KKN
 
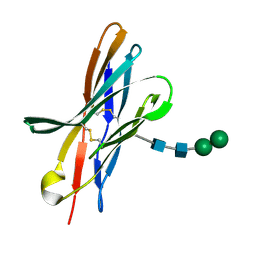 | | Crystal structure of bovine CTLA-4, PSI-NYSGRC-012704 | | Descriptor: | Cytotoxic T-lymphocyte associated protein 4, alpha-D-mannopyranose-(1-3)-beta-D-mannopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose | | Authors: | Kumar, P.R, Ahmed, M, Banu, R, Bhosle, R, Bonanno, J, Calarese, D.A, Celikgil, A, Chamala, S, Chan, M.K, Chowdhury, S, Fiser, A, Garforth, S.J, Scott Glenn, A, Hammonds, J, Hillerich, B, Khafizov, K, Lafleur, J, Attonito, J, Love, J.D, Patel, H, Patel, R, Seidel, R.D, Smith, B, Stead, M, Toro, R, Casadevall, A, Almo, S.C, New York Structural Genomics Research Consortium (NYSGRC), Atoms-to-Animals: The Immune Function Network (IFN) | | Deposit date: | 2013-05-06 | | Release date: | 2013-06-12 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.253 Å) | | Cite: | Crystal structure of bovine CTLA-4, PSI-NYSGRC-012704
to be published
|
|
4K55
 | |
4L1H
 
 | |
4M62
 
 | |
4LRN
 
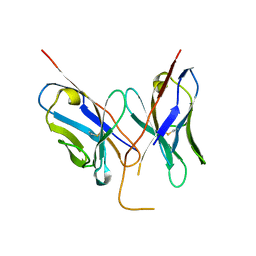 | |
1I9E
 
 | | TCR DOMAIN | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, CYTOTOXIC TCELL VALPHA DOMAIN | | Authors: | Rudolph, M.G, Huang, M, Teyton, L, Wilson, I.A. | | Deposit date: | 2001-03-19 | | Release date: | 2001-12-05 | | Last modified: | 2023-08-09 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Crystal structure of an isolated V(alpha) domain of the 2C T-cell receptor.
J.Mol.Biol., 314, 2001
|
|
1I8I
 
 | | CRYSTAL STRUCTURE OF DSFV MR1 IN COMPLEX WITH THE PEPTIDE ANTIGEN OF THE MUTANT EPIDERMAL GROWTH FACTOR RECEPTOR, EGFRVIII, AT ROOM TEMPERATURE | | Descriptor: | EPIDERMAL GROWTH FACTOR RECEPTOR ANTIBODY MR1SCFV HEAVY CHAIN, EPIDERMAL GROWTH FACTOR RECEPTOR ANTIBODY MR1SCFV LIGHT CHAIN, EPIDERMAL GROWTH FACTOR RECEPTOR, ... | | Authors: | Landry, R.C, Klimowicz, A.C, Lavictoire, S.J, Borisova, S, Kottachchi, D.T, Lorimer, I.A, Evans, S.V. | | Deposit date: | 2001-03-14 | | Release date: | 2002-03-14 | | Last modified: | 2021-10-27 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Antibody recognition of a conformational epitope in a peptide antigen: Fv-peptide complex of an antibody fragment specific for the mutant EGF receptor, EGFRvIII.
J.Mol.Biol., 308, 2001
|
|
1JV5
 
 | | Anti-blood group A Fv | | Descriptor: | Ig chain heavy chain precursor V region, Ig kappa chain precursor V region | | Authors: | Thomas, R, Patenaude, S.I, MacKenzie, C.R, To, R, Hirama, T, Young, N.M, Evans, S.V. | | Deposit date: | 2001-08-28 | | Release date: | 2002-01-09 | | Last modified: | 2018-04-18 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Structure of an anti-blood group A Fv and improvement of its binding affinity without loss of specificity.
J.Biol.Chem., 277, 2002
|
|
6B8O
 
 | | WT Ig-like V Domain with Phosphatidylserine | | Descriptor: | 1,2-DICAPROYL-SN-PHOSPHATIDYL-L-SERINE, 1,2-ETHANEDIOL, 2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Sudom, A, Wang, Z. | | Deposit date: | 2017-10-09 | | Release date: | 2018-06-27 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Molecular basis for the loss-of-function effects of the Alzheimer's disease-associated R47H variant of the immune receptor TREM2.
J. Biol. Chem., 293, 2018
|
|
6AW0
 
 | | Crystal structure of CEACAM3 L44Q | | Descriptor: | CHLORIDE ION, Carcinoembryonic antigen-related cell adhesion molecule 3, GLYCEROL, ... | | Authors: | Bonsor, D.A, Sundberg, E.J. | | Deposit date: | 2017-09-05 | | Release date: | 2018-05-16 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (2.43 Å) | | Cite: | TheHelicobacter pyloriadhesin protein HopQ exploits the dimer interface of human CEACAMs to facilitate translocation of the oncoprotein CagA.
EMBO J., 37, 2018
|
|
1HKF
 
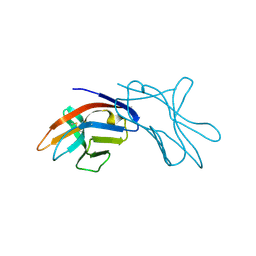 | | The three dimensional structure of NK cell receptor Nkp44, a triggering partner in natural cytotoxicity | | Descriptor: | NK CELL ACTIVATING RECEPTOR | | Authors: | Ponassi, M, Cantoni, C, Biassoni, R, Conte, R, Spallarossa, A, Moretta, A, Moretta, L, Bolognesi, M, Bordo, D. | | Deposit date: | 2003-03-10 | | Release date: | 2003-06-11 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | The Three-Dimensional Structure of the Human Nk Cell Receptor Nkp44, a Triggering Partner in Natural Cytotoxicity
Structure, 11, 2003
|
|
4LVE
 
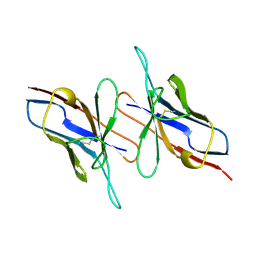 | |
4L1D
 | | Voltage-gated sodium channel beta3 subunit Ig domain | | Descriptor: | Sodium channel subunit beta-3 | | Authors: | Namadurai, S, Weimhofer, M, Rajappa, R, Stott, K, Klingauf, J, Chirgadze, D.Y, Jackson, A.P. | | Deposit date: | 2013-06-03 | | Release date: | 2014-03-05 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Crystal Structure and Molecular Imaging of the Nav Channel beta 3 Subunit Indicates a Trimeric Assembly.
J.Biol.Chem., 289, 2014
|
|
1I8K
 
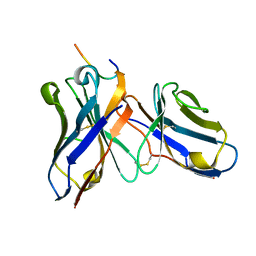 | | CRYSTAL STRUCTURE OF DSFV MR1 IN COMPLEX WITH THE PEPTIDE ANTIGEN OF THE MUTANT EPIDERMAL GROWTH FACTOR RECEPTOR, EGFRVIII, AT LIQUID NITROGEN TEMPERATURE | | Descriptor: | EPIDERMAL GROWTH FACTOR RECEPTOR ANTIBODY MR1SCFV HEAVY CHAIN, EPIDERMAL GROWTH FACTOR RECEPTOR ANTIBODY MR1SCFV LIGHT CHAIN, EPIDERMAL GROWTH FACTOR RECEPTOR, ... | | Authors: | Landry, R.C, Klimowicz, A.C, Lavictoire, S.J, Borisova, S, Kottachchi, D.T, Lorimer, I.A, Evans, S.V. | | Deposit date: | 2001-03-14 | | Release date: | 2002-03-14 | | Last modified: | 2021-10-27 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Antibody recognition of a conformational epitope in a peptide antigen: Fv-peptide complex of an antibody fragment specific for the mutant EGF receptor, EGFRvIII.
J.Mol.Biol., 308, 2001
|
|
6C74
 
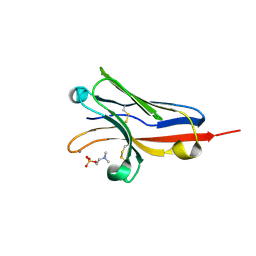 | |
6BIT
 
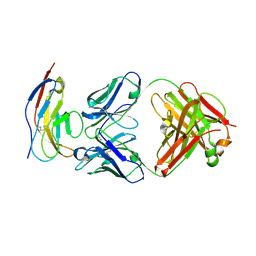 | | SIRPalpha antibody complex | | Descriptor: | KWAR23 Fab heavy chain, KWAR23 Fab light chain, Tyrosine-protein phosphatase non-receptor type substrate 1 | | Authors: | Ring, N.G, Herndler-Brandstetter, D, Weiskopf, K, Shan, L, Volkmer, J.P, George, B.M, Lietzenmayer, M, McKenna, K.M, Naik, T.J, McCarty, A, Zheng, Y, Ring, A.M, Flavell, R.A, Weissman, I.L. | | Deposit date: | 2017-11-03 | | Release date: | 2017-12-06 | | Last modified: | 2019-11-20 | | Method: | X-RAY DIFFRACTION (2.191 Å) | | Cite: | Anti-SIRP alpha antibody immunotherapy enhances neutrophil and macrophage antitumor activity.
Proc. Natl. Acad. Sci. U.S.A., 114, 2017
|
|
4K07
 
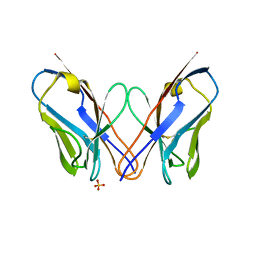 | | Crystal structure of the amyloid-forming immunoglobulin AL-103 cis-proline 95 mutant | | Descriptor: | Amyloidogenic immunoglobulin light chain protein AL-103, SULFATE ION | | Authors: | Thompson, J.R, Berkholz, D.S, Mahlum, E.W, Ramirez-Alvarado, M. | | Deposit date: | 2013-04-03 | | Release date: | 2013-10-30 | | Last modified: | 2014-01-22 | | Method: | X-RAY DIFFRACTION (2.83 Å) | | Cite: | Kinetic control in protein folding for light chain amyloidosis and the differential effects of somatic mutations.
J.Mol.Biol., 426, 2014
|
|
1J05
 
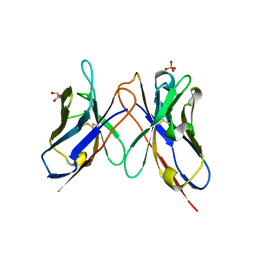 | | The crystal structure of anti-carcinoembryonic antigen monoclonal antibody T84.66 Fv fragment | | Descriptor: | GLYCEROL, PHOSPHATE ION, anti-CEA mAb T84.66, ... | | Authors: | Kondo, H, Nishimura, Y, Shiroishi, M, Asano, R, Noro, N, Tsumoto, K, Kumagai, I. | | Deposit date: | 2002-11-01 | | Release date: | 2003-12-16 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | The crystal structure of anti-carcinoembryonic antigen monoclonal antibody T84.66 Fv fragment
To be Published
|
|
1JFQ
 
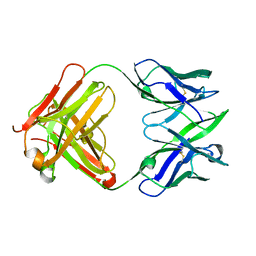 | | ANTIGEN-BINDING FRAGMENT OF THE MURINE ANTI-PHENYLARSONATE ANTIBODY 36-71, "FAB 36-71" | | Descriptor: | ANTIGEN-BINDING FRAGMENT OF ANTI-PHENYLARSONATE ANTIBODY | | Authors: | Parhami-Seren, B, Viswanathan, M, Strong, R.K, Margolies, M.N. | | Deposit date: | 2001-06-21 | | Release date: | 2002-02-27 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Structural analysis of mutants of high-affinity and low-affinity
p-azophenylarsonate-specific antibodies generated by alanine
scanning of heavy chain
complementarity-determining region 2.
J.Immunol., 167, 2001
|
|
6AW1
 | | Crystal structure of CEACAM3 | | Descriptor: | CHLORIDE ION, Carcinoembryonic antigen-related cell adhesion molecule 3, DI(HYDROXYETHYL)ETHER, ... | | Authors: | Bonsor, D.A, Sundberg, E.J. | | Deposit date: | 2017-09-05 | | Release date: | 2018-05-16 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | TheHelicobacter pyloriadhesin protein HopQ exploits the dimer interface of human CEACAMs to facilitate translocation of the oncoprotein CagA.
EMBO J., 37, 2018
|
|
4M8Q
 | |
1I85
 
 | | CRYSTAL STRUCTURE OF THE CTLA-4/B7-2 COMPLEX | | Descriptor: | CYTOTOXIC T-LYMPHOCYTE-ASSOCIATED PROTEIN 4, T LYMPHOCYTE ACTIVATION ANTIGEN CD86 | | Authors: | Schwartz, J.-C.D, Zhang, X, Fedorov, A.A, Nathenson, S.G, Almo, S.C. | | Deposit date: | 2001-03-12 | | Release date: | 2001-04-04 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (3.2 Å) | | Cite: | Structural basis for co-stimulation by the human CTLA-4/B7-2 complex.
Nature, 410, 2001
|
|
4NOF
 
 | | Crystal structure of the second Ig domain from mouse Polymeric Immunoglobulin receptor [PSI-NYSGRC-006220] | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, GLYCEROL, Polymeric immunoglobulin receptor | | Authors: | Sampathkumar, P, Kumar, P.R, Ahmed, M, Banu, R, Bhosle, R, Calarese, D.A, Celikgil, A, Chamala, S, Chan, M.K, Chowdhury, S, Fiser, A, Garforth, S.J, Glenn, A.S, Hillerich, B, Khafizov, K, Attonito, J, Love, J.D, Patel, H, Patel, R, Seidel, R.D, Smith, B, Stead, M, Casadevall, A, Almo, S.C, New York Structural Genomics Research Consortium (NYSGRC), Atoms-to-Animals: The Immune Function Network (IFN) | | Deposit date: | 2013-11-19 | | Release date: | 2013-12-04 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | Crystal structure of the second Ig domain from mouse Polymeric Immunoglobulin receptor
to be published
|
|
4MZ3
 
 | |
