1BC9
 
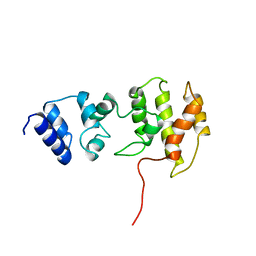 | | CYTOHESIN-1/B2-1 SEC7 DOMAIN, NMR, MINIMIZED AVERAGE STRUCTURE | | Descriptor: | CYTOHESIN-1 | | Authors: | Betz, S.F, Schnuchel, A, Wang, H, Olejniczak, E.T, Meadows, R.P, Fesik, S.W. | | Deposit date: | 1998-05-06 | | Release date: | 1999-05-11 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Solution structure of the cytohesin-1 (B2-1) Sec7 domain and its interaction with the GTPase ADP ribosylation factor 1.
Proc.Natl.Acad.Sci.USA, 95, 1998
|
|
1BCI
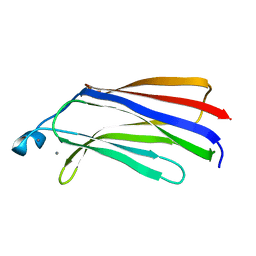 
 | | C2 DOMAIN OF CYTOSOLIC PHOSPHOLIPASE A2, NMR, MINIMIZED AVERAGE STRUCTURE | | Descriptor: | CALCIUM ION, CYTOSOLIC PHOSPHOLIPASE A2 | | Authors: | Xu, G.Y, Mcdonagh, T, Yu, H.A, Nalefski, E.A, Clark, J.D, Cumming, D.A. | | Deposit date: | 1998-04-30 | | Release date: | 1998-11-25 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Solution structure and membrane interactions of the C2 domain of cytosolic phospholipase A2.
J.Mol.Biol., 280, 1998
|
|
1AW3
 
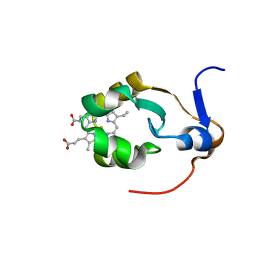 | | THE SOLUTION NMR STRUCTURE OF OXIDIZED RAT MICROSOMAL CYTOCHROME B5, MINIMIZED AVERAGE STRUCTURE | | Descriptor: | CYTOCHROME B5, PROTOPORPHYRIN IX CONTAINING FE | | Authors: | Arnesano, F, Banci, L, Bertini, I, Felli, I.C. | | Deposit date: | 1997-10-09 | | Release date: | 1998-02-04 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | The solution structure of oxidized rat microsomal cytochrome b5.
Biochemistry, 37, 1998
|
|
6U6G
 
 | |
6V5T
 
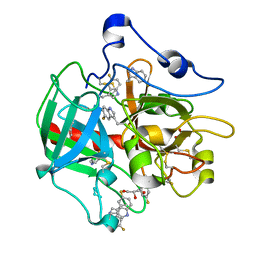 | |
6V64
 
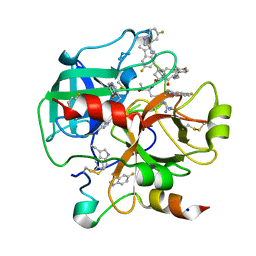 | | Crystal structure of human thrombin bound to ppack with tryptophans replaced by 5-F-tryptophan | | Descriptor: | D-phenylalanyl-N-[(2S,3S)-6-{[amino(iminio)methyl]amino}-1-chloro-2-hydroxyhexan-3-yl]-L-prolinamide, SODIUM ION, Thrombin heavy chain, ... | | Authors: | Ruben, E.A, Chen, Z, Di Cera, E. | | Deposit date: | 2019-12-04 | | Release date: | 2020-05-13 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (2.29 Å) | | Cite: | 19F NMR reveals the conformational properties of free thrombin and its zymogen precursor prethrombin-2.
J.Biol.Chem., 295, 2020
|
|
1RSB
 
 | |
1P0O
 | | HP (2-20) substitution of Trp for Gln and Asp at position 17 and 19 MODIFICATION IN SDS-D25 MICELLES | | Descriptor: | 19-mer peptide from 50S ribosomal protein L1 | | Authors: | Lee, K.H, Lee, D.G, Park, Y.K, Harm, K.S, Kim, Y.M. | | Deposit date: | 2003-04-10 | | Release date: | 2003-05-20 | | Last modified: | 2021-11-10 | | Method: | SOLUTION NMR | | Cite: | Interactions between antimicrobial peptide, HP(2-20) derived from helicobacter pylori, and membrain studied by nmr spectroscopy
To be published
|
|
4BKT
 
 | | von Hippel Lindau protein:ElonginB:ElonginC complex, in complex with (2S,4R)-N-methyl-1-[2-(3-methyl-1,2-oxazol-5-yl)ethanoyl]-4-oxidanyl-pyrrolidine-2-carboxamide | | Descriptor: | (2S,4R)-N-methyl-1-[2-(3-methyl-1,2-oxazol-5-yl)ethanoyl]-4-oxidanyl-pyrrolidine-2-carboxamide, ARGININE, GLUTAMIC ACID, ... | | Authors: | Van Molle, I, Dias, D.M, Baud, M, Galdeano, C, Geraldes, C.F.G.C, Ciulli, A. | | Deposit date: | 2013-04-29 | | Release date: | 2013-11-27 | | Last modified: | 2017-07-12 | | Method: | X-RAY DIFFRACTION (2.35 Å) | | Cite: | Is NMR Fragment Screening Fine-Tuned to Assess Druggability of Protein-Protein Interactions?
Acs Med.Chem.Lett., 5, 2014
|
|
4BKS
 
 | | von Hippel Lindau protein:ElonginB:ElonginC complex, in complex with (2S,4R)-1-ethanoyl-N-[[4-(1,3-oxazol-5-yl)phenyl]methyl]-4-oxidanyl-pyrrolidine-2-carboxamide | | Descriptor: | (2S,4R)-1-ethanoyl-N-[[4-(1,3-oxazol-5-yl)phenyl]methyl]-4-oxidanyl-pyrrolidine-2-carboxamide, ACETATE ION, TRANSCRIPTION ELONGATION FACTOR B POLYPEPTIDE 1, ... | | Authors: | Van Molle, I, Dias, D.M, Baud, M, Galdeano, C, Geraldes, C.F.G.C, Ciulli, A. | | Deposit date: | 2013-04-29 | | Release date: | 2013-11-27 | | Last modified: | 2014-01-29 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Is NMR Fragment Screening Fine-Tuned to Assess Druggability of Protein-Protein Interactions?
Acs Med.Chem.Lett., 5, 2014
|
|
1M6T
 
 | | CRYSTAL STRUCTURE OF B562RIL, A REDESIGNED FOUR HELIX BUNDLE | | Descriptor: | SULFATE ION, Soluble cytochrome b562 | | Authors: | Chu, R, Takei, J, Knowlton, J.R, Andrykovitch, M, Pei, W, Kajava, A.V, Steinbach, P.J, Ji, X, Bai, Y. | | Deposit date: | 2002-07-17 | | Release date: | 2002-11-06 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.81 Å) | | Cite: | Redesign of a Four-Helix Bundle Protein by Phage Display Coupled with Proteolysis
and Structural Characterization by NMR and X-ray Crystallography
J.Mol.Biol., 323, 2002
|
|
1P0L
 | | HP (2-20) Substitution GLN To TRP Modification In SDS-D25 MICELLES | | Descriptor: | 19-mer peptide from 50S ribosomal protein L1 | | Authors: | Lee, K.H, Lee, D.G, Park, Y.K, Harm, K.S, Kim, Y.M. | | Deposit date: | 2003-04-10 | | Release date: | 2003-05-20 | | Last modified: | 2021-11-10 | | Method: | SOLUTION NMR | | Cite: | Interactions between antimicrobial peptide, HP(2-20) derived from helicobacter pylori, and membrain studied by nmr spectroscopy
To be published
|
|
1P5L
 
 | | HP (2-20) Substitution PHE5 to SER modification in sds-d25 micelles | | Descriptor: | 19-mer peptide from 50S ribosomal protein L1 | | Authors: | Lee, K.H, Lee, D.G, Park, Y.K, Harm, K.S, Kim, Y.M. | | Deposit date: | 2003-04-27 | | Release date: | 2003-06-03 | | Last modified: | 2021-11-10 | | Method: | SOLUTION NMR | | Cite: | Interactions between antimicrobial peptide, HP(2-20) derived from helicobacter pylori, and membrain studied by nmr spectroscopy
To be published
|
|
1P0J
 
 | | HP (2-20) Substitution ASP To TRP Modification In SDS-D25 Micelles | | Descriptor: | 19-mer peptide from 50S ribosomal protein L1 | | Authors: | Lee, K.H, Lee, D.G, Park, Y.K, Harm, K.S, Kim, Y.M. | | Deposit date: | 2003-04-10 | | Release date: | 2003-05-20 | | Last modified: | 2021-11-10 | | Method: | SOLUTION NMR | | Cite: | Interactions between antimicrobial peptide, HP(2-20) derived from helicobacter pylori, and membrain studied by nmr spectroscopy
To be published
|
|
1P5K
 
 | | HP (2-20) Substitution SER to LEU11 modification in sds-d25 micelles | | Descriptor: | 19-mer peptide from 50S ribosomal protein L1 | | Authors: | Lee, K.H, Lee, D.G, Park, Y.K, Harm, K.S, Kim, Y.M. | | Deposit date: | 2003-04-27 | | Release date: | 2003-06-03 | | Last modified: | 2021-11-10 | | Method: | SOLUTION NMR | | Cite: | Interactions between antimicrobial peptide, HP(2-20) derived from helicobacter pylori, and membrain studied by nmr spectroscopy
To be published
|
|
1TVG
 | | X-ray structure of human PP25 gene product, HSPC034. Northeast Structural Genomics Target HR1958. | | Descriptor: | CALCIUM ION, LOC51668 protein, SAMARIUM (III) ION | | Authors: | Kuzin, A.P, Vorobiev, S.M, Lee, I, Acton, T.B, Montelione, G.T, Tong, L, Hunt, J.F, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2004-06-29 | | Release date: | 2004-11-09 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Improving NMR protein structure quality by Rosetta refinement: a molecular replacement study.
Proteins, 75, 2009
|
|
1KRW
 
 | | SOLUTION STRUCTURE AND BACKBONE DYNAMICS OF BERYLLOFLUORIDE-ACTIVATED NTRC RECEIVER DOMAIN | | Descriptor: | NITROGEN REGULATION PROTEIN NR(I) | | Authors: | Hastings, C.A, Lee, S.-Y, Cho, H.S, Yan, D, Kustu, S, Wemmer, D.E. | | Deposit date: | 2002-01-10 | | Release date: | 2003-08-19 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | High-Resolution Solution Structure of the Beryllofluoride-Activated NtrC Receiver Domain
Biochemistry, 42, 2003
|
|
1A1U
 
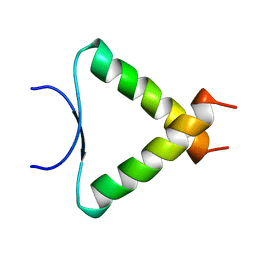 | | SOLUTION STRUCTURE DETERMINATION OF A P53 MUTANT DIMERIZATION DOMAIN, NMR, MINIMIZED AVERAGE STRUCTURE | | Descriptor: | P53 | | Authors: | Mccoy, M.A, Stavridi, E.S, Waterman, J.L.F, Wieczorek, A, Opella, S.J, Halezonetis, T.D. | | Deposit date: | 1997-12-16 | | Release date: | 1998-04-08 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Hydrophobic side-chain size is a determinant of the three-dimensional structure of the p53 oligomerization domain.
EMBO J., 16, 1997
|
|
1A6X
 
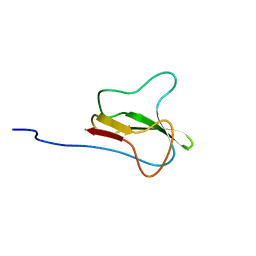 | | STRUCTURE OF THE APO-BIOTIN CARBOXYL CARRIER PROTEIN (APO-BCCP87) OF ESCHERICHIA COLI ACETYL-COA CARBOXYLASE, NMR, 49 STRUCTURES | | Descriptor: | APO-BIOTIN CARBOXYL CARRIER PROTEIN OF ACETYL-COA CARBOXYLASE | | Authors: | Yao, X, Wei, D, Soden Junior, C, Summers, M.F, Beckett, D. | | Deposit date: | 1998-03-04 | | Release date: | 1998-10-14 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Structure of the carboxy-terminal fragment of the apo-biotin carboxyl carrier subunit of Escherichia coli acetyl-CoA carboxylase.
Biochemistry, 36, 1997
|
|
1A56
 
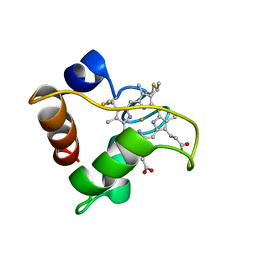 | | PRIMARY SEQUENCE AND SOLUTION CONFORMATION OF FERRICYTOCHROME C-552 FROM NITROSOMONAS EUROPAEA, NMR, MEAN STRUCTURE REFINED WITH EXPLICIT HYDROGEN BOND CONSTRAINTS | | Descriptor: | FERRICYTOCHROME C-552, HEME C | | Authors: | Timkovich, R, Bergmann, D, Arciero, D.M, Hooper, A.B. | | Deposit date: | 1998-02-20 | | Release date: | 1998-10-21 | | Last modified: | 2020-12-16 | | Method: | SOLUTION NMR | | Cite: | Primary sequence and solution conformation of ferrocytochrome c-552 from Nitrosomonas europaea.
Biophys.J., 75, 1998
|
|
6CKV
 | |
6CMY
 
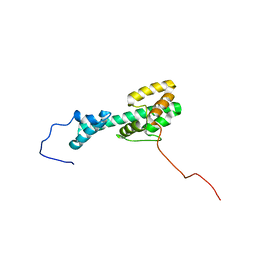 | | Solution NMR Structure Determination of Mouse Melanoregulin | | Descriptor: | Melanoregulin | | Authors: | Rout, A.K, Wu, X, Strub, M.P, Starich, M.R, Hammer III, J.A, Tjandra, N. | | Deposit date: | 2018-03-06 | | Release date: | 2018-09-19 | | Last modified: | 2024-05-15 | | Method: | SOLUTION NMR | | Cite: | The Structure of Melanoregulin Reveals a Role for Cholesterol Recognition in the Protein's Ability to Promote Dynein Function.
Structure, 26, 2018
|
|
1AM0
 
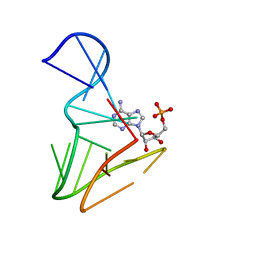 | | AMP RNA APTAMER COMPLEX, NMR, 8 STRUCTURES | | Descriptor: | ADENOSINE MONOPHOSPHATE, RNA APTAMER | | Authors: | Jiang, F, Kumar, R.A, Jones, R.A, Patel, D.J. | | Deposit date: | 1997-06-19 | | Release date: | 1997-07-23 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Structural Basis of RNA Folding and Recognition in an AMP-RNA Aptamer Complex
Nature, 382, 1996
|
|
1A8N
 
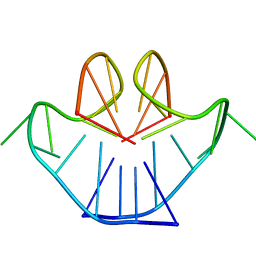 | | SOLUTION STRUCTURE OF A NA+ CATION STABILIZED DNA QUADRUPLEX CONTAINING G.G.G.G AND G.C.G.C TETRADS FORMED BY G-G-G-C REPEATS OBSERVED IN AAV AND HUMAN CHROMOSOME 19, NMR, 8 STRUCTURES | | Descriptor: | DNA QUADRUPLEX CONTAINING G.G.G.G AND G.C.G.C TETRADS | | Authors: | Kettani, A, Bouaziz, S, Gorin, A, Zhao, H, Jones, R, Patel, D.J. | | Deposit date: | 1998-03-27 | | Release date: | 1998-10-14 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Solution structure of a Na cation stabilized DNA quadruplex containing G.G.G.G and G.C.G.C tetrads formed by G-G-G-C repeats observed in adeno-associated viral DNA.
J.Mol.Biol., 282, 1998
|
|
1AJF
 
 | |
